Abstract
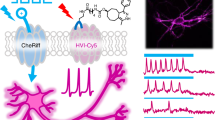
The precise co-localization and stoichiometric expression of two different light-gated membrane proteins can vastly improve the physiological usefulness of optogenetics for the modulation of cell excitability with light. Here we present a gene-fusion strategy for the stable 1:1 expression of any two microbial rhodopsins in a single polypeptide chain. By joining the excitatory channelrhodopsin-2 with the inhibitory ion pumps halorhodopsin or bacteriorhodopsin, we demonstrate light-regulated quantitative bi-directional control of the membrane potential in HEK293 cells and neurons in vitro. We also present synergistic rhodopsin combinations of channelrhodopsin-2 with Volvox carteri channelrhodopsin-1 or slow channelrhodopsin-2 mutants, to achieve enhanced spectral or kinetic properties, respectively. Finally, we demonstrate the utility of our fusion strategy to determine ion-turnovers of as yet uncharacterized rhodopsins, exemplified for archaerhodopsin and CatCh, or to correct pump cycles, exemplified for halorhodopsin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Nagel, G. et al. Light activation of channelrhodopsin-2 in excitable cells of Caenorhabditis elegans triggers rapid behavioral responses. Curr. Biol. 15, 2279–2284 (2005).
Han, X. & Boyden, E. Multiple-color optical activation, silencing, and desynchronization of neural activity, with single-spike temporal resolution. PLoS ONE 2, e299 (2007).
Zhang, F. et al. Multimodal fast optical interrogation of neural circuitry. Nature 446, 633–639 (2007).
Douin, V. et al. Use and comparison of different internal ribosomal entry sites (IRES) in tricistronic retroviral vectors. BMC Biotechnol. 4, 16 (2004).
Han, X., Qian, X., Stern, P., Chuong, A. & Boyden, E. Informational lesions: optical perturbation of spike timing and neural synchrony via microbial opsin gene fusions. Front. Mol. Neurosci. 2, 12 (2009).
Tang, W. et al. Faithful expression of multiple proteins via 2A-peptide self-processing: a versatile and reliable method for manipulating brain circuits. J. Neurosci. 29, 8621–8629 (2009).
de Felipe, P. et al. E unum pluribus: multiple proteins from a self-processing polyprotein. Trends Biotechnol. 24, 68–75 (2006).
Gradinaru, V. et al. Molecular and cellular approaches for diversifying and extending optogenetics. Cell 141, 154–165 (2009).
Geibel, S. et al. The voltage-dependent proton pumping in bacteriorhodopsin is characterized by optoelectic behavior. Biophys. J. 81, 2059–2068 (2001).
Seki, A. et al. Heterologous expression of Pharaonis halorhodopsin in Xenopus laevis oocytes and electrophysiological characterization of its light-driven Cl− pump activity. Biophys. J. 92, 2559–2569 (2007).
Nagel, G., Möckel, B., Büldt, G. & Bamberg, E. Functional expression of bacteriorhodopsin in oocytes allows direct measurement of voltage dependence of light induced H+ pumping. FEBS Lett. 377, 263–266 (1995).
Shcherbo, D. et al. Bright far-red fluorescent protein for whole-body imaging. Nat. Methods 4, 741–746 (2007).
Feldbauer, K. et al. Channelrhodopsin-2 is a leaky proton pump. Proc. Natl. Acad. Sci. USA 106, 12317–12322 (2009).
Zufferey, R. et al. Self-inactivating lentivirus vector for safe and efficient in vivo gene delivery. J. Virol. 72, 9873–9880 (1998).
Bamann, C., Gueta, R., Kleinlogel, S., Nagel, G. & Bamberg, E. Structural guidance of the photocycle of channelrhodopsin-2 by an interhelical hydrogen bond. Biochemistry 49, 267–278 (2010).
Berndt, A., Yizhar, O., Gunaydin, L., Hegemann, P. & Deisseroth, K. Bi-stable neural state switches. Nat. Neurosci. 12, 229–234 (2009).
Zhang, F. et al. Red-shifted optogenetic excitation: a tool for fast neural control derived from Volvox carteri. Nat. Neurosci. 11, 631–633 (2008).
Chizhov, I. & Engelhard, M. Temperature and halide dependence of the photocycle of halorhodopsin from Natronobacterium pharaonis. Biophys. J. 81, 1600–1612 (2001).
Lanyi, J. Halorhodopsin: A light-driven chloride ion pump. Annu. Rev. Biophys. Biophys. Chem. 15, 11–28 (1986).
Dencher, N. & Wilms, M. Flash photometric experiments on the photochemical cycle of bacteriorhodopsin. Biophys. Struct. Mech. 1, 259–271 (1975).
Chow, B. et al. High-performance genetically targetable optical neural silencing by light-driven proton pumps. Nature 463, 98–102 (2010).
Kleinlogel, S. et al. Ultra light-sensitive and fast neuronal activation with the Ca2+-permeable channelrhodopsin CatCh. Nat. Neurosci. 14, 513–518 (2011).
Yang, F., Moss, L. & Phillips, G. The molecular structure of green fluorescent protein. Nat. Biotechnol. 14, 1246–1251 (1996).
Patterson, G. & Lippincott-Schwartz, J. A photoactivatable GFP for selective photolabeling of proteins and cells. Science 297, 1873–1877 (2002).
Nagel, G. et al. Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proc. Natl. Acad. Sci. USA 100, 13940–13945 (2003).
Hackmann, C. et al. Static and time-resolved step-scan Fourier transform infrared investigations of the photoreaction of halorhodopsin from Natronobacterium pharaonis: consequences for models of the anion translocation mechanism. Biophys. J. 81, 394–406 (2001).
Hegemann, P., Oesterhelt, D. & Bamberg, E. The transport activity of the light-driven chloride pump halorhodopsin is regulated by green and blue light. Biochim. Biophys. Acta 819, 195–205 (1985).
Pletnev, S. et al. A crystallographic study of bright far-red fluorescent protein mKate reveals pH-induced cis-trans isomerization of the chromophore. J. Biol. Chem. 283, 28980–28987 (2008).
Allocca, M. et al. Serotype-dependent packaging of large genes in adeno-associated viral vectors results in effecive gene delivery in mice. J. Clin. Invest. 118, 1955–1964 (2008).
Büning, H. et al. Receptor targeting of adeno-associated virus vectors. Gene Ther. 10, 1142–1151 (2003).
Allocca, M. et al. Novel adeno-associated virus serotypes efficiently transduce murine photoreceptors. J. Virol. 81, 11372–11380 (2007).
Arenkiel, B. et al. In vivo light-induced activation of neural circuitry in transgenic mice expressing channelrhodopsin-2. Neuron 54, 205 (2007).
Thyagarajan, S. et al. Visual function in mice with photoreceptor degeneration and transgenic expression of channelrhodopsin 2 in ganglion cells. J. Neurosci. 30, 8745 (2010).
Zimmermann, D. et al. Effects on capacitance by overexpression of membrane proteins. Biochem. Biophys. Res. Commun. 369, 1022–1026 (2008).
Acknowledgements
We thank V. Busskamp and B. Roska (Friedrich Miescher Institute for Biomedical Research, Basel) for the pAAV-LTR-EGFP plasmid and help with the rAAV construction, I. Bartnik for preparation of hippocampal neuron cultures, and H. Biehl and V. Eulenburg (Max Planck Institute for Brain Research, Frankfurt) for help with the antibody screening. The transgenic mouse expressing ChR2-EYFP was a gift from H. Wässle (Max Planck Institute for Brain Research, Frankfurt). The pAAV2-Rho-EGFP vector was a gift from A. Auricchio (Telethon Institute of Genetics and Medicine, Naples). The work was supported by the Deutsche Forschungsgemeinschaft Sonderforschungsbereich 807, Centre of Excellence Frankfurt Macromolecular Complexes, the German Federal Ministry of Education and Research (01GQ0815) and by the EU FP7 OptoNeuro Project (249867) to E.B. and by the Max Planck Society.
Author information
Authors and Affiliations
Contributions
E.B., S.K. and P.G.W. conceived the molecular and electrophysiological experiments, and S.K., B.L. and U.T. carried them out. D.G. and C.B. performed the antibody screening. S.K., U.T. and C.B. designed and carried out the data analysis. S.K. and E.B. wrote the paper. E.S.B. supplied the Arch plasmid and contributed to writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Notes 1–5 (PDF 2135 kb)
Rights and permissions
About this article
Cite this article
Kleinlogel, S., Terpitz, U., Legrum, B. et al. A gene-fusion strategy for stoichiometric and co-localized expression of light-gated membrane proteins. Nat Methods 8, 1083–1088 (2011). https://doi.org/10.1038/nmeth.1766
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1766
This article is cited by
-
PACmn for improved optogenetic control of intracellular cAMP
BMC Biology (2021)
-
BiPOLES is an optogenetic tool developed for bidirectional dual-color control of neurons
Nature Communications (2021)
-
Potassium channel-based optogenetic silencing
Nature Communications (2018)
-
Overexpression of Different Types of Microbial Rhodopsins with a Highly Expressible Bacteriorhodopsin from Haloarcula marismortui as a Single Protein in E. coli
Scientific Reports (2018)
-
Dynamic all-optical drug screening on cardiac voltage-gated ion channels
Scientific Reports (2018)