Abstract
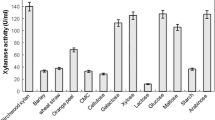
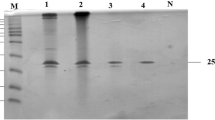
Xylanases constitute an important industrial enzyme, which hydrolyzes the polysaccharide xylan. In this work, a novel Streptomyces strain producing cellulase-free xylanase was isolated from the soil samples collected from the mangrove forest of Kadalundi, Kerala, India. The strain produced unique enzyme, which exhibited optimal activity at pH 9.0 and tolerance up to pH 12.0. Media engineering was carried out to improve the enzyme production, which showed best enzyme production at 30°C, medium pH 9.0 and incubation time of 48 h. Enzyme was highly thermo-tolerant up to 70°C and alkaline tolerant. Partial gene amplification as well as partial purification of enzyme was carried out to characterize the enzyme. The unique features of the enzyme make it an ideal candidate for industrial application for paper and pulp industry.
Similar content being viewed by others
References
Abdelwahed N.A.M., El-Naggar N.E. & Saber W.I.A. 2011. Factors and correlations controlling cellulase-free xylanase production by Streptomyces halstedii NRRL B-1238 in submerged culture. Aust. J. Basic Appl. Sci. 5: 45–53.
Altschul S.F., Gish W., Miller W., Myers E.W. & Lipman D.J. 1990. Basic local alignment search tool. J. Mol. Biol. 215: 403–410.
Bailey M.J., Beily P.& Poutanen K. 1992. Interlaboratory testing and methods for assay of xylanase activity. J. Biotechnol. 23: 257–270.
Beg Q.K., Bharat B., Mukesh K. & Hoondal G.S. 2000. Enhanced production of a thermostable xylanase from Streptomyces sp. QG-11-3 and its application in biobleaching of eucalyptus kraft pulp. Enzyme Microb. Technol. 27: 459–466.
Beg Q.K., Kapoor M., Mahajan L. & Hoondal G.S. 2001. Microbial xylanases and their industrial applications: a review. Appl. Microbiol. Biotechnol. 56: 326–338.
Benson D.A., Cavanaugh M., Clark K., Karsch-Mizrachi I., Lipman D.J., Ostell J. & Sayers E.W. 2013. GenBank. Nucleic Acids Res. 41(Database issue): D36–D42.
Collins T., Gerday C. & Feller G. 2005. Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol. Rev. 29: 3–23.
Corpet F. 1988. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 16: 10881–10890.
Dhanasekaran D., Sivamani P., Arunagrinathan N., Paneerselvum A. & Thajuddin N. 2005. Screening and identification of antibiotic producing strains of marine Streptomyces. J. Microbial World 7: 62–66.
Dhiman S.S., Sharma J. & Battan B. 2008. Industrial applications and future prospects of microbial xylanases: a review. BioResources 3: 1377–1402.
Ghose T.K. 1987. Measurement of cellulase activities. Pure Appl. Chem. 59: 257–268.
Gupta B.N., Mishra S. & Basak U.C. 2007. Occurrence of Streptomyces aurantiacus in mangroves of Bhitarkanika. Malaysian J. Microbiol. 3: 7–14.
Johnson L.I. & Curl E.A. 1972. Methods for Research on Ecologyof Soil Born Pathogens. Burgess Pub. Co., Minneapolis, 247 pp.
Kumar A., Gupta P., Shrivastava B., Khosa Y.P. & Kuhad R.C. 2012. Xylanase production from an alkalophilic actinomycete isolate Streptomyces sp. RCK 2010, its characterization and application in saccharification of second generation biomass. J. Mol. Catal. B Enzymatic 74: 170–177.
Laemmli U.K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Larkin M.A., Blackshields G., Brown N.P., Chenna R., McGettigan P.A., McWilliam H., Valentin F., Wallace I.M., Wilm A., Lopez R., Thompson J.D., Gibson T.J. & Higgins D.G. 2007. Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947–2948.
Lescic I., Zehl M., Muller R., Vukelic B., Abramic M., Pigac J., Allmaier G. & Kojicprodic B. 2004. Structural characterization of extracellular lipase from Streptomyces rimosus: assignment of disulphide bridge pattern by mass spectrometry. Biol. Chem. 385: 1147–1156.
Maheswari M.U. & Chandra T.S. 2000. Production and potential applications of a xylanase from a new strain of Streptomyces cuspidosporus. World J. Microbiol. Biotechnol. 16: 257–263.
Miller G.L. 1959. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 31: 426–428.
Ninawe S., Lal R. & Kuhad R.C. 2006. Isolation of three xylanase producing strains of actinomycetes and their identification using molecular methods. Curr. Microbiol. 53: 178–182.
Peixoto-Nogueira S., Michelin M., Betini J.H.A., Jorge J.A., Terenzi H.F. & Polizeli M.L.T.M. 2009. Production of xylanase by Aspergilli using alternative carbon sources: application of the crude extract on cellulose pulp biobleaching. J. Ind. Microbiol. Biotechnol. 36: 149–155.
Ratanakhanokchai K., Kyu K.L. & Tanticharoen M. 1999. Purification and properties of a xylan-binding endoxylanase from alkalophilic Bacillus sp. strain K-1. Appl. Environ. Microbiol. 65: 694–697.
Saratale G.D., Sartale R.G. & Koh S.E. 2012. Production and characterization of multiple cellulolytic enzymes by isolated Streptomyces sp. MDS. Biomass Bioenergy 47: 302–315.
Sharma P. & Bajaj B.K. 2006. Production and partial characterization of alkali tolerant xylanases from an alkalophilic Streptomyces sp. CD3. J. Sci. Ind. Res. 64: 688–697.
Singh R., Kapoor V. & Kumar V. 2012. Utilization of agroindustrial waste for the simultaneous production of amylase and xylanases by thermophilic actinomycetes. Braz. J. Microbiol. 43: 1545–1552.
Thomas L. Arumugam M. & Pandey A. 2013. Production, purification, characterization and over-expression of xylanases from actinomycetes. Ind. J. Expt. Biol.
Uyar F. & Baysal Z. 2004. Production and optimization of process parameters for alkaline protease production by a newly isolated Bacillus sp. under solid state fermentation. Process Biochem. 39: 1893–1898.
Author information
Authors and Affiliations
Corresponding author
Additional information
Based on a contribution presented at the International Conference on Industrial Biotechnology (ICIB-2012), November 21–23, 2012, Punjabi University, Patiala (Pb.), India
Rights and permissions
About this article
Cite this article
Thomas, L., Sindhu, R. & Pandey, A. Identification and characterization of a highly alkaline and thermotolerant novel xylanase from Streptomyces sp.. Biologia 68, 1022–1027 (2013). https://doi.org/10.2478/s11756-013-0248-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-013-0248-5