Abstract
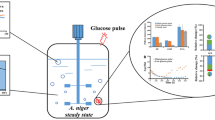
We conducted an integrated study of cell growth parameters, product formation, and the dynamics of intracellular metabolite concentrations using Escherichia coli with genes knocked out in the glycolytic and oxidative pentose phosphate pathway (PPP) for glucose catabolism. We investigated the same characteristics in the wild-type strain, using acetate or pyruvate as the sole carbon source. Dramatic effects on growth parameters and extracellular and intracellular metabolite concentrations were observed after blocking either glycolytic breakdown of glucose by inactivation of phosphoglucose isomerase (disruption of pgi gene) or pentose phosphate breakdown of glucose by inactivation of glucose-6-phosphate dehydrogenase (disruption of zwf gene). Reducing power (NADPH) was mainly produced through PPP when the pgi gene was knocked out, while NADPH was produced through the tricarboxylic acid (TCA) cycle by isocitrate dehydrogenase or NADP-linked malic enzyme when the zwf gene was knocked out. As expected, when the pgi gene was knocked out, intracellular concentrations of PPP metabolites were high and glycolytic and concentrations of TCA cycle pathway metabolites were low. In the zwf gene knockout, concentrations of PPP metabolites were low and concentrations of intracellular glycolytic and TCA cycle metabolites were high.
Similar content being viewed by others
Abbreviations
- CER:
-
CO2 evolution rate
- DCW:
-
dry cell weight
- ED:
-
Entner-Doudoroff
- EMP:
-
Embden-Mayerhof-Paranas
- MEZ:
-
malic enzyme
- OD:
-
optical density
- OUR:
-
oxygen uptake rate
- PGI:
-
phosphoglucose isomerase
- PPP:
-
pentose phosphate pathway
- TCA:
-
tricarboxylic acid
- Yx/s :
-
cell mass yield
- 6PGDH:
-
6-phosphogluconate dehydrogenase
- Eda:
-
Entner-Douderoff aldolase
- Edd:
-
Entner-Douderoff dehydralase
- Eno:
-
enolase
- Fba:
-
fructose-1,6-bisphosphate aldolase
- G6PDH:
-
glucose-6-phosphate dehydrogenase
- GAPDH:
-
glyceraldehyde-3-phosphate dehydrogenase
- GDH:
-
glutamate dehydrogenase
- Hxk:
-
hexokinase
- ICDH:
-
isocitrate dehydrogenase
- LDH:
-
lactate dehydrogenase
- MDH:
-
malate dehydrogenase
- Mk:
-
myokinase
- Pck:
-
PEP carboxykinase
- Pgi:
-
phosphoglucose isomerase
- Ppc:
-
phosphoenolpyruvate carboxylase
- Pta:
-
phosphotransacetylase
- PTS:
-
phosphotransferase system
- Pyk:
-
pyruvate kinase
- Rpe:
-
ribose-phosphate epimerase
- Rpi:
-
ribose-phosphate isomerase
- Tkt:
-
transketolase
- Tpi:
-
triosephosphate isomerase
- Tal:
-
transaldolase
- 2PG:
-
2-phosphoglycerate
- 6PG:
-
6-phosphogluconate
- AcCoA:
-
acetyl-coenzyme A
- ADP:
-
adenosine diphosphate
- AKG:
-
α-ketoglutarate
- AMP:
-
adenosine monophosphate
- ATP:
-
adenosine triphosphate
- DHAP:
-
dihydroxyacetone phosphate
- E4P:
-
erythrose-4-phosphate
- F6P:
-
fructose-6-phosphate
- FBP:
-
fructose-1,6-bisphosphate
- G6P:
-
glucose-6-phosphate
- GAP:
-
glyceraldehyde-3-phosphate
- ICT:
-
isocitrate
- NAD:
-
diphosphopyridindinucleotide, oxidized
- NADH:
-
diphosphopyridindinucleotide, reduced
- NADP:
-
diphosphopyridindinucleotide-phosphate, oxidized
- NADPH:
-
diphosphopyridindinucleotide-phosphate, reduced
- OAA:
-
oxaloacetate
- PEP:
-
phosphoenolpyruvate
- PYR:
-
pyruvate
- R5P:
-
ribose-5-phosphate
- RU5P:
-
ribulose-5-phosphate
- SUC:
-
succinate
References
Bailey J.E. 1991. Toward a science of metabolic engineering. Science 252: 1668–1675.
Bergmeyer H.U. 1984a. Methods of Enzymatic Analysis, 3rd Ed., Vol. 6. Verlag Chemie, Weinheim, Germany.
Bergmeyer H.U. 1984b. Methods of Enzymatic Analysis, 3rd Ed., Vol. 7. Verlag Chemie, Weinheim, Germany.
Berry A. 1996. Improving production of aromatic compounds in Escherchia coli by metabolic engineering. Trends Biotechnol. 14: 250–256.
Buchholz A., Jurlebaus J., Christian W. & Takors R. 2002. Metabolomics: quantification of intracellular metabolite dynamics. Biomol. Eng. 19: 5–15.
Buchholz A., Takors R. & Christian W. 2001. Quantification of intracellular metabolites in Escherichia coli using liquid chromatographic-electrospray ionization tandem mass spectrometric techniques. Anal. Biochem. 295: 129–137.
Burgard A.P. & Maranas C.D. 2001. Probing the performance limits of the Escherichia coli metabolic network subject to additions or deletions. Biotechnol. Bioeng. 74: 364–375.
Canonaco F., Hess T.A., Heri S., Wang T., Szyperski T. & Sauer U. 2001. Metabolic flux response to phosphoglucose isomerase knock-out in Escherichia coli and impact of overexpression of the soluble transhydrogenase UdhA. FEMS Microbiol. Lett. 204: 247–252.
Choi I.Y., Sup K.I., Kim H.J. & Park J.W. 2003. Thermosensitive phenotype of Escherichia coli mutant lacking (NADP(+)-dependent isocitrate dehydrogenase. Redox Rep. 8: 51–56.
Datsenko K.A. & Wanner B.L. 2000. One-step inactivation of chromosomal genes in Escherichia coli K12 using PCR products. Proc. Natl. Acad. Sci. USA 97: 6640–6645.
Dauner M., Storni T. & Sauer U. 2001. Bacillus subtilis metabolism and energetic in carbon-limited and excesscarbon chemostat culture. J. Bacteriol. 183: 7308–7317.
De Koning W. & Van Dam K. 1992. A method for the determination of changes of glycolytic metabolites in yeast on a subsecond time scale using extraction at neutral pH. Anal. Biochem. 204: 118–123.
Emmerling M., Dauner M., Ponti A., Fiaux J., Hochuli M., Szyperski T., Wuthrich K., Bailey J.E. & Sauer U. 2002. Metabolic flux responses to pyruvate kinase knockout in Escherichia coli. J. Bacteriol. 184: 152–164.
Choi G.G., Bae M.S., Ahn C.Y. & Oh H.M. 2008. Enhanced biomass and γ-linolenic acid production of mutant strain Arthrospira platensis. J. Microbiol. Biotechnol. 18: 539–544.
Goel A.J., Ferrance J., Jeong J. & Attai A. 1993. Analysis of metabolic fluxes in batch and continuous cultures of Bacillus subtilis. Biotechnol. Bioeng. 42: 686–696.
Junke H., Krems B., Kotter P. & Entian K.D. 1996. Mutants that show increased sensitivity to hydrogen peroxide reveal an important role for the pentose phosphate pathway in protection of yeast against oxidative stress. Mol. Gen. Genet. 252: 456–464.
Hoque M.A., Siddiquee K.A.Z. & Shimizu K. 2004. Metabolic control analysis of gene-knockout Escherichia coli based on the inverse flux analysis with experimental verification. Biochem. Eng. J. 19: 53–59.
Hoque M.A., Ushiyama H., Tomita M. & Shimizu K. 2005. Dynamic responses of the intracellular metabolite concentrations of the wild type and pykA Escherichia coli against pulse addition of glucose or NH3 under those limiting continuous cultures. Biochem. Eng. J. 26: 38–49.
Hua Q., Yang C., Baba T., Mori H. & Shimizu K. 2003. Responses of the central metabolism in Escherichia coli to phosphoglucose isomerase and glucose-6-phosphate dehydrogenase knock-out. J. Bacteriol. 185: 7053–7067.
Hua Q., Yang C., Oshima T., Mori H. & Shimizu K. 2004. Analysis of gene expression in Escherichia coli in response to changes of growth-limiting nutrient in chemostat cultures. Appl. Environ. Microbiol. 70: 2354–2366.
Hurlebaus J., Buchholz A., Alt W., Wiechert W. & Takors R. 2002. MMT-A pathway modeling tool for data from rapid sampling experiments. In Silico Biology 2: 467–484.
Ishii N., Nakahigashi K., Baba T., Robert M., Soga T., Kanai A., Hirasawa T., Naba M., Hirai K., Hoque A., Ho P.Y., Kakazu Y., Sugawara K., Igarashi S., Harada S., Masuda T., Sugiyama N., Togashi T., Hasegawa M., Takai Y., Yugi K., Arakawa K., Iwata N., Toya Y., Nakayama Y., Nishioka T., Shimizu K., Mori H. & Tomita M. 2007. Multiple highthroughput analyses monitor the tesponse of E. coli to perturbations. Science 316: 593–597.
Larsson C.U., von Stokar U., Marison I. & Gustafsson L. 1993. Growth and metabolism of Saccharomyce cerevisiae in chemostat cultures under carbon, nitrogen, or carbon- and nitrogen-limiting conditions. J. Bacteriol. 175: 4809–4816.
Lim S.J., Jung Y.M., Shin H.D. & Lee Y.H. 2002. Application of the NADPH-related genes zwf and gnd for the oddball biosynthesis of PHB in an E. coli transformant harboring a cloned phbCAB operon. J. Biosci. Bioeng. 93: 543–549.
Lowenstein J.M. 1969. Methods in Enzymology, Vol. XIII, Citric Acid Cycle. Academic Press, New York.
Matsudo M.C., Bezerra R.P., Sato S., Perego P., Converti A. & Carvalho J.C.M. 2009. Repeated fed-batch cultivation of Arthrospira (Spirulina) platensis using urea as nitrogen source. Biochem. Eng. J. 43: 52–57.
Park S.J., Cotter P.A. & Gunsalus R.P. 1995. Regulation of malate dehydrogenase (mdh) gene expression in Echerichia coli in response to oxygen, carbon and heme availability. J. Bacteriol. 177: 6652–6656.
Ping H., Leighton T., Ishkhanova G. & Kustu S. 1999. Sensing of nitrogen limited by Bacillus subtilis: comparison to enteric bacteria. J. Bacteriol. 181: 5042–5050.
Piorreck M., Hinnerk K., Pohl B. & Pohl P. 1984. Biomass production, total protein chlorophylls, lipids and fatty acids of freshwater green and blue green algae under different nitrogen regimes. Phytochemistry 23: 207–216.
Rerenci T. 1999. Regulation by nutrient limitation. Curr. Opin. Microbiol. 2: 208–213.
Rizzi M., Baltes M., Theobald U. & Reuss M. 1997. In vivo analysis of metabolic dynamics in Saccharomces cerevisiae: II. Mathematical model. Biotechnol. Bioeng. 55: 592–608.
Sarkar D., Siddiquee K.A.Z., Arauzo-Bravo M.J., Oba T. & Shimizu K. 2008. Effect of cra gene knockout together with edd and iclR genes knockout on the metabolism in Escherichia coli. Arch. Microbiol. 190: 559–571.
Sauer U., Lasko D.R., Fiaux J., Hochuli M., Glaser R., Szyperski T., Wuthrich K. & Bailey E.J. 1999. Metabolic flux ratio analysis of genetic environmental modulations of Escherichia coli central carbon metabolism. J. Bacteriol. 181: 6679–6688.
Schaefer U., Boos W., Takors R. & Weuster-Botz D. 1999. Automated sampling device for monitoring intracellular metabolite dynamics. Anal. Biochem. 270: 88–96.
Senior P.J. 1975. Regulation of nitrogen metabolism in Escherichia coli and Klebsiella aerogenes: studies with the continuous-culture technique. J. Bacteriol. 123: 407–418.
Siddiquee K.A.Z., Arauzo-Bravo M.J. & Shimizu K. 2004. Metabolic flux analysis of pykF gene knockout Escherichia coli based on 13C-labeling experiments together with measurements of enzyme activities and intracellular metabolite concentrations. Appl. Microbiol. Biotechnol. 63: 407–417.
Stephanopoulos G., Nielsen J. & Aristidou A. 1998. Metabolic Engineering: Principles and Methodologies. Academic Press, London.
Tao H., Bausch C., Richmond C., Blatner R.F. & Conway T. 1999. Functional genomics: expression analysis of Escherichia coli growing on minimal and rich media. J. Bacteriol. 181: 6425–6440.
Theobald U., Milinger W., Baltes M., Rizzi M. & Reuss M., 1997. In vivo analysis of metabolic dynamics in Saccharomyces cerevisiae: I. Experimental observations. Biotechnol. Bioeng. 55: 305–316.
Vaseghi S., Baumeister A., Rizzi M. & Reuss M. 1999. In vivo dynamics of the pentose phosphate pathway in Saccharomyces cerevisiae. Metab. Eng. 1: 128–140.
Westerhoff H.V. 2001. The silicon cell, not dead but live! Metab. Eng. 3: 207–210.
Weuster-Botz D. & de Graff A.A. 1996. Reaction engineering methods to study intracellular metabolite concentrations. Adv. Biochem. Eng. Biotechnol. 54: 75–108.
Yang C., Hua Q., Baba T., Mori T. & Shimizu K., 2003. Analysis of Escherichia coli anaplerotic metabolism and its regulation mechanisms from the metabolic responses to altered dilution rates and phosphoenolpyruvate carboxykinase knockout. Biotechnol. Bioeng. 84: 129–144.
Zhao J., Baba T., Mori H. & Shimizu K. 2004a. Global metabolic response of Escherichia coli to gnd or zwf gene-knockout, based on 13C-labeling experiments and the measurement of enzyme activities. Appl. Microbiol. Biotechnol. 64: 91–98.
Zhao J., Baba T., Mori H. & Shimizu K. 2004b. Effect of zwf gene knock-out on the metabolism of Escherichia coli grown on glucose or acetate. Metab. Eng. 6: 164–174.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hoque, M.A., Fard, A.T., Rahman, M. et al. Comparison of dynamic responses of cellular metabolites in Escherichia coli to pulse addition of substrates. Biologia 66, 954–966 (2011). https://doi.org/10.2478/s11756-011-0136-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-011-0136-9