-
PDF
- Split View
-
Views
-
Cite
Cite
Paul E. Griffiths, Eva M. Neumann-Held, The many faces of the gene, BioScience, Volume 49, Issue 8, August 1999, Pages 656–662, https://doi.org/10.2307/1313441
Close - Share Icon Share
Despite the centrality of the term “gene” to modern biology, its definition remains a matter of controversy. A univocal definition may not be necessary or even desirable; different “gene concepts” may be useful in different areas of biology. However, it is necessary to clearly distinguish these different concepts and to use each in its proper domain.
In this article, we argue for a strict differentiation between two gene concepts: the “molecular gene” and the “evolutionary gene.” The definition of both is highly problematic. The molecular gene is, roughly speaking, a DNA sequence that codes for a polypeptide. But, as we will discuss, this general conception raises several problems: Does a “gene” include its introns? Does it include its regulator and promoter sequences? In cases in which the spliced mRNA transcript undergoes further editing, does the gene include the machinery that dictates the sequence of the final transcript? We believe that these questions point to a tension between two theoretical goals. The first goal is to identify genes with particular DNA segments on chromosomes, thus maintaining one of the theoretical commitments of classical genetics; the second is to make genes a core part of the developmental explanation of phenotypic traits—especially, although not exclusively, the heritable trait differences studied by classical genetics. As we will discuss, the first goal is simply not achievable in light of current knowledge about how final gene products are constructed. Abandoning the first goal in favor of the second leads to what we call the “molecular process” concept of the gene.
A very different gene definition is the “evolutionary gene concept,” which was introduced by George C. Williams (1966). An evolutionary gene is any stretch of DNA that can be thought of as being in competition with other stretches of DNA for representation in future generations. As Williams and others have noted, the theoretical utility of this gene concept is independent of changing ideas about molecular genes. However, advocates of the evolutionary gene concept have generally failed to recognize the price of this independence: Assumptions about evolutionary genes cannot be defended by appeal to the validity of these assumptions for molecular genes. The evolutionary gene is a genuinely separate theoretical entity. Therefore, claims about evolutionary genes—such as the claim that they are inherited relatively independently of one another—must be defended in the context of evolutionary, rather than molecular, biology. Before addressing these issues, we briefly review the history of the concept of the gene.
Opening the black box
The population genetic theories that are central to contemporary neo-Darwinism were formulated before any direct examination of the genetic material was possible. Consequently, in population genetics and, more generally, in “modern synthesis” biology, the gene was and is treated as a “black box.”
The input to the genetic black box is a pair of parental factors; the output, via some simple rules of dominance, is the phenotype of the offspring. If the black box fails to yield the expected output in accordance with these rules, the failure is ascribed to “incomplete penetrance”—an ad hoc explanation that leads to no new testable predictions or theoretical inquiries. What is omitted in this approach is the contents of the black box: the process of development. It is development that connects the genetic units, whose obedience to Mendel's laws is explained by meiosis, to the phenotypes that are inherited in Mendelian fashion. In classical genetics, therefore, the effects of cellular and environmental influences on the phenotypic expression of these genetic units are not available for study because they occur inside the black box.
The “black boxing” of development allowed evolutionary biology to enrich itself with the findings of classical genetics without having to confront the complexities of development. This strategy has been so successful that there is a certain reluctance in some quarters to look inside the box now that molecular biology has provided the tools with which to do so.
The first step toward opening the box is to recognize that the metaphorical ideas of DNA containing the “information for,” “codingfor,” or “programming for” phenotypic traits cannot be the final stopping points in an explanation of development (Oyama 1985). These phrases are simply another way to express the existence of the black box. Indeed, the idea that developmental biology could be replaced by going straight to the genes and reading the instructions for development has been rightly mocked as “neo-preformationism.” The homunculus, the tiny man of early embryology who solved the “problem of generation” by already possessing all the complexity of the adult, has been replaced by a little blueprint that solves the problem in the same unsatisfactory way. References to programs and blueprints are promissory notes redeemable against developmental biology.
The conventional defense of such informal “information talk” in genetics is that it is meant “more or less in the spirit of information theory” (quote from John Maynard-Smith; Fifth International Congress of Systematic & Evolutionary Biology, Budapest, 17–24 August 1996). But the possibility of translating the information metaphor into substantive theory is an illusion. Repeated attempts to bridge the gap between information talk and information theory in molecular biology have been unsuccessful (Sarkar 1996, Janich 1998). In fact, an adequate reduction of information talk is simply the same thing as an understanding of developmental biology. In the cases of inheritance that classical genetics was able to study, the relationship between alleles and phenotypic traits was manageably simple because the other resources that help to produce the trait could be held constant. These other resources are the “channel conditions” under which the life cycle of the organism carries information about which DNA sequences are present (Griffiths and Gray 1994). That is to say, if these other factors are held constant, then the phenotype will vary predictably with the DNA sequences that are present and thus carry information about the presence of those sequences. It is this information that was recovered in the breeding experiments of classical genetics. However, understanding the channel along which this information flows means elucidating the developmental channel conditions.
The evolution of the molecular gene concept
A distinguished lineage of research strategies postulates a one-to-one correspondence between a gene and some meaningful developmental unit. Most historians of biology place the beginning of this tradition with the establishment of Thomas Hunt Morgan's research group at Columbia University. Beginning in 1917, when Morgan published his classic paper, “The Theory of the Gene,” this group accumulated statistical data from hybridization, breeding, and mutation experiments to defend the claim that genes—as independent hereditary elements—must exist in the germ plasm and must serve as the units of heredity. Although this claim was frequently challenged, it was extremely influential.
By the 1940s, molecular and biochemical investigation of the nature of the gene and its physiological functioning had begun. In their famous paper, Beadle and Tatum (1941) argued that one gene corresponds to one “primary” character and one enzyme. The demonstration that DNA rather than protein is the basis of Mendelian heredity soon followed (Avery et al. 1944). Finally, in the 1960s, the breaking of the genetic code led to the conceptualization of the gene that still predominates today. According to this “classical molecular gene concept” (Neumann-Held 1999), a gene is a stretch of DNA, initially thought to be uninterrupted, that codes for a single polypeptide chain.
The history of the molecular gene concept reflects the continuing conviction on the part of geneticists that a theory of inheritance should be a theory of the inheritance of units of development, be these phenotypic traits, enzymes, or polypeptides. An initial difficulty with the classical molecular concept of the gene is that the actual activity of the gene, and hence its developmental effect, depends on elements outside the open reading frame. This complexity has led to definitions of the gene that include the promoter and regulatory sequences that affect whether the gene will be transcribed.
In some cases, such as that of the famous lac operon of Escherichia coli, these regulatory regions are immediately upstream of the site at which transcription is initiated, which makes it easy to regard them as part of the gene. In eukaryotes, however, regulatory regions can be distant from the rest of the gene and can be involved in the regulation of more than one gene. The discovery of introns—portions of the gene discarded during post-transcriptional processing—in eukaryote genes (Jeffreys and Flavell 1977) created further difficulties for the classical molecular gene concept. Several gene products can be made from one gene by cutting and splicing the primary transcript in alternative ways. This discovery led to further proposals in the conceptual lineage we have outlined. For instance, it has been proposed that a gene is the stretch of DNA corresponding to one primary mRNA transcript (sometimes including, sometimes excluding promoter and regulatory sequences).
In the face of this proliferation of meanings, Kenneth C. Waters made his well-known proposal that all of these different definitions of the term “gene” can be reduced to the same underlying concept of the gene (Waters 1994, Neumann-Held 1999): The “fundamental concept…is that of a gene for a linear sequence in a product at some stage of genetic expression” (Waters 1994). Waters' concept allows for either inclusion or exclusion of introns. Whether introns are part of the gene depends on which particular “linear sequence in a product at some stage of genetic expression” is being referred to (Waters 1994). An intron, for example, is part of the gene when the focus is on the process of transcription at the stage of the primary mRNA. It is not part of the gene when the focus is on the polypeptide chain.
Although Waters' proposed definition certainly reflects the current usage of the term “gene,” we doubt whether it can help clarify the conceptual issues raised by the growing understanding of the complexity of gene expression. According to Waters' proposed definition, various “genes” come into being at different stages of the expression process. On the one hand, the entire sequence of the DNA from which a primary mRNA is transcribed can be called a gene. On the other hand, the term “gene” might refer only to those parts of that sequence that correspond to the mature mRNA. In the case of differential mRNA processing, several “genes” would stem from one “gene.”
However, it seems strange, to say the least, to assert that single genes on the DNA correspond to a variety of genes on the mRNA. There seems to be no conceptual gain from Waters' proposed definition and no clarification in language use. On the contrary, in molecular biology, the empirical data can be described much more precisely in terms that are already available, such as noncoding region, primary mRNA, mature mRNA, intron, exon, and coding region. The use of more differentiated terms such as these is particularly important when trying to understand developmental processes on the molecular level.
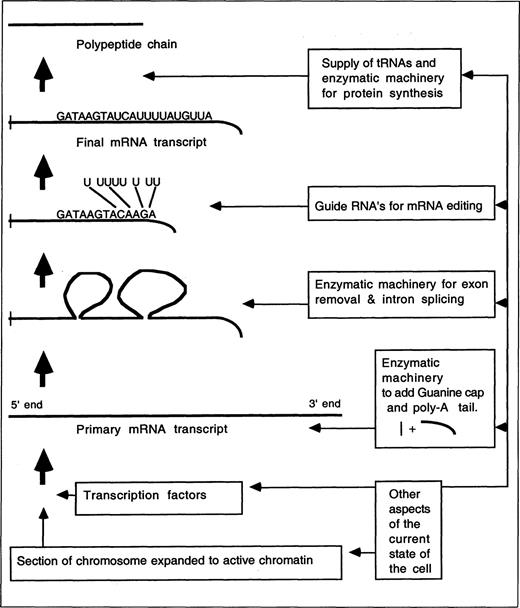
Whereas in Waters' approach introns are included or excluded in the gene concept, depending on the context, Christopher Epp has recently suggested that introns should be definitively excluded (Epp 1997). He suggests that a gene is “the nucleotide sequence that stores the information which specifies the order of monomers in a final functional polypeptide or RNA molecule, or set of closely related isoforms” (Epp 1997). Allowing for “closely related isoforms” enables Epp to cope with the discovery of alternative mRNA splicing. However, the discovery of mRNA editing (Hanson 1996) renders Epp's proposal problematic. With mRNA editing, the order of nucleotides in the DNA need not determine the order of monomers in the gene product. Individual bases may be added or deleted from the mRNA during processing (see Figure 1).
Schematic representation of some transcription, editing, and translation processes, highlighting the context dependency of the expression of gene products.
The molecular process gene concept
It has clearly become difficult, as more has been learned about the expression of gene products, to preserve the idea that a single gene corresponds to a single product at some chosen stage of post-transcriptional processing while maintaining a clear and univocal meaning for the term “gene.” One option is to retain the idea that a gene is a linear DNA sequence but to abandon the idea that a gene has a single developmental role. A gene might, for instance, be defined as a DNA sequence corresponding to a single “norm of reaction” of gene products across varying cellular conditions.
Because this approach represents the most conservative response to the empirical problems facing the classical molecular gene concept, we might call it the “contemporary molecular gene concept.” However, the underlying rationale of the lineage of gene concepts described above—making genes developmentally meaningful units—suggests an alternative way to look at the problem. Rather than allowing the unit of development corresponding to each gene to become a disjunction of possible consequences under different epigenetic conditions, why not build the epigenetic conditions into the gene, just as some earlier definitions built regulator and promoter sequences into the gene?
To understand the rationale behind this new proposal, it is useful to return to Epp's discussion. He argued against the idea that a gene includes the regulatory sequences required for its expression, saying that “…genes do not have to be expressed to be present. The somatic cells of a multicellular organism all have the same genes, but particular cell types express only some of them” (Epp 1997, p. 537). He went on to say that “inclusion of regulatory sequences expands the term ‘gene’ from a specification of ‘what is’ to indicate also ‘how it is used.’ It may be useful to separate these two concepts.” (Epp 1997, p. 537). We agree that these are two separate concepts, but contra Epp we would argue that the term “gene” points to the latter and not to the former. As we have argued above, the concept of the gene has always been intimately linked to how genes are used in development. It is the DNA itself, not the gene, that just “is.” For example, in the case of so-called overlapping genes it is known that DNA regions inside an open reading frame can function as promoter sites for a downstream open reading frame. This situation suggests that whether a DNA region is part of a particular gene depends on what function it is currently performing. When the shared region is functioning as a promoter, the fact that under other circumstances it is part of an open reading sequence is irrelevant.
The fact that a DNA sequence can be used for different purposes at different times and in different tissues has further consequences. It is natural to think of a promoter sequence as a structural entity—the sequence of DNA bases typical of promoters. In the cases of overlapping genes, it still seems attractive to say that the promoter sequence is a promoter, even when it is not functioning as a promoter. But the independence of function and structure is not restricted to the case of overlapping genes. If one looks at an open reading frame that participates in the expression of a particular polypeptide, one can identify fixed borders of that coding region and exon/intron boundaries within the region. However, at another time or in another tissue, those exon/intron boundaries and the borders of the coding region can shift. Therefore, it is the role of a particular DNA sequence in a developmental system—tissue specific, organism specific, involving environmental inputs and developmental history—that influences whether the sequence is used (and can be described) as an intron or a coding region, as well as whether it is seen as a promoter or as part of an open reading frame.
The sequence of the DNA can thus be compared to a sequence of letters without spaces or punctuation marks. The state of the developmental system is then analogous to a scheme imposed on these letters—grouping letters into words, adding punctuation marks and editing notes. A different developmental state imposes a different scheme over the letters, that is, over the DNA sequence. It is therefore misleading to think of functional descriptions of DNA, such as “promoter region,” as explicable solely in terms of structural descriptions of DNA, such as sequence. The structural description is, at best, a necessary condition for the functional description to apply.
These considerations lead away from the classical molecular gene concept to what we have christened the “molecular process gene concept.” According to this concept, “gene” denotes the recurring process that leads to the temporally and spatially regulated expression of a particular polypeptide product (Neumann-Held 1999). This gene concept allows for alternative mRNA splicing as well as for mRNA editing by including the particular processes involved in either. There is a great deal of continuity between this proposal and the classical molecular conception of the gene: The gene still has the function of coding for a polypeptide, and it still includes specific segments of DNA. However, the gene is identified not with these DNA sequences alone but rather with the process in whose context these sequences take on a definite meaning.
Further support for the molecular process gene concept comes from the fact that when one speaks of the “gene for” a particular product, one is implicitly referring not only to DNA sequences but also to all the other influences that cause that sequence to give rise to this product. The molecular process gene concept stresses these connections and helps scientists bear in mind the easily overlooked fact that the production of this polypeptide product is the result, not of the presence of the DNA sequence alone, but of a whole range of resources affecting gene expression. If there is anything that is “for” a gene product, it is the molecular process that produces that product rather than a sequence of nucleotides, which, as Epp puts it, just “is.”
The evolutionary gene concept
A radically different approach to the gene is embodied in the so-called evolutionary gene concept introduced by Williams (1966) and elaborated by Richard Dawkins (Dawkins 1982, 1989), according to which a gene is any stretch of DNA that could be replaced by an alternative sequence in future generations. There is considerable confusion concerning the relationship between evolutionary and molecular genes (see box page 660). We discuss Dawkins' views here in some depth because his popular writing seems to have been as influential in the scientific community as with the wider public and has contributed to this confusion.
The new evolutionary gene concept was vigorously attacked by the molecular biologist Gunther Stent for excluding from the definition of the gene much of the knowledge that molecular biology had by then generated about the stretches of chromosome corresponding to Mendelian alleles (Stent 1977, 1994). Dawkins has treated Stent's criticisms as a territorial dispute over the use of the word “gene” (Dawkins 1982), but this response is too simplistic. Certainly, the concept of an evolutionary gene should not be confused with the various molecular concepts described above. In contrast to Dawkins, however, we contend that evolutionary genes cannot in general be identified with specific stretches of DNA, which is the price to be paid for the conceptual independence of evolutionary genes from molecular genes.
One of Dawkins' best-known statements about the evolutionary gene is that “any stretch of DNA, beginning and ending at arbitrarily chosen points on the chromosome, can be considered to be competing with allelomorphic stretches for the region of chromosome concerned” (Dawkins 1982, p. 87). Taken by itself, however, this definition of the “selfish gene” can lead to extensive misunderstandings. On further reading, it becomes obvious that this definition imposes several other requirements for evolutionary genehood. What Dawkins calls “dead-end replicators”—the genes in somatic cells in animals with a segregated germ line—are excluded from his definition of an evolutionary gene because they do not have the potential to give rise to indefinitely many copies. Within the remaining “germline replicators” he distinguishes between “active” and “passive” replicators. Dawkins explains that “an active replicator is any replicator whose nature has some influence over its probability of being copied” (Dawkins 1982). This statement seems to imply that to be an active replicator, a sequence must compete for representation in a population with other, alternative sequences. Thus, to Dawkins a sequence is an active replicator and thereby an evolutionary gene only if substituting some other sequence of equal length would change the fitness of the organism that contains it or the capacity of this DNA sequence to compete directly with other sequences, as in meiotic drive.
So Dawkins defines evolutionary genes not in terms of the polypeptides that they produce, but rather in terms of phenotypic differences between members of a population—specifically, differences that cause differential replication rates. The sole connection to molecular biology is Dawkins' stipulation that DNA sequences are the only candidates for the factors responsible for these differences in reproductive success (except in cultural evolution, which he regards as a separate process).
One consequence of Dawkins' definition is that single nucleotides become good candidates for evolutionary genehood. In the case of sickle-cell anemia, for example, a single-nucleotide change causes a structural change in a hemoglobin polypeptide chain, which leads in turn to further significant changes in a number of phenotypic traits affecting fitness. The two nucleotides that compete to occupy this locus in human populations fulfill all the requirements for evolutionary genehood. However, Dawkins himself rejects the application of the evolutionary gene concept to single nucleotides, describing this idea as “an absurdly reductionistic reductio ad absurdum” (Dawkins 1982) that “…becomes downright misleading if it suggests…that adenine at one locus is, in some sense, allied with adenine at other loci, pulling together for an adenine team. If there is any sense in which purines and pyrimidines compete with each other for heterozygous loci, the struggle at each locus is insulated from the struggle at other loci…. If they are competitors at all, they are competitors for each locus separately. They are indifferent to the fate of their exact replicas at other loci” (Dawkins 1982, p. 91).
If this argument is correct, then it actually makes a case against the whole idea of selfish replicators and evolutionary genes because Dawkins' case against the selfish nucleotide is equally applicable to larger DNA segments. Dawkins is correct that there is no “alliance” between copies of the same single nucleotide, but neither is there any alliance between large, physically identical sequences of DNA at separate loci. Indeed, there is no such alliance even between identical classical molecular genes at separate loci. For example, after a gene duplication event it will often be “in the interest” of one of the resulting gene copies for the other to be replaced by a mutant form. In monkeys, the gene for the red opsin and for the green opsin are allelic to one another and are X-linked, so males can see red or green but not both. In apes (including humans), a gene duplication has occurred such that males have both red and green opsins and can see both red and green (Jacobs 1996). Immediately after the duplication of one of the opsin genes, it is “in the interest” of a duplicated opsin gene for the other copy of that gene to be replaced by a different allele at the other locus so that its descendants will find themselves in males with full color vision. Here, contrary to what Dawkins suggests, genes are no more “allied” with copies of themselves at other loci than are single nucleotides.
Classical Mendelian: A gene is a stretch of chromosome that is associated with a phenotypic difference.
Classical molecular: A gene is a DNA sequence from which a particular molecular product is expressed.
Contemporary molecular: A gene is a DNA sequence with a particular norm of reaction of molecular products across different cellular conditions.
Molecular process: A gene is a process that includes DNA sequences and components outside the DNA sequence that contribute to the time- and tissue-regulated expression of a particular polypeptide product.
Evolutionary: A gene is any sequence of DNA on a chromosome, or, in other words, “that which segregates and recombines with appreciable frequency” (Williams 1966). See text for later elaborations by other authors.
Dawkins presents a second argument against the “selfish nucleotide,” which is that the causal powers of nucleotides are too context dependent for them to count as active replicators (Dawkins 1982). In the example of sickle-cell anemia, the single-nucleotide change leads to anemia only if it occurs in a specific place in the genome, and only if the organism is homozygous at this locus. But these facts can hardly be used to argue that “selfish nucleotides” are not evolutionary genes. A long DNA sequence that is not a molecular gene is probably more, not less, context dependent than a single nucleotide. The effect of inserting a single base will be the same in many different places in the genome. For example, inserting guanine between uracil and adenine will consistently produce a stop codon in one reading frame. Whenever this insertion occurs in a molecular gene such that the new stop codon is in frame, the result will be a truncated protein. Inserting a string of junk, however, is unlikely to have consistent effects. Even the effect of inserting the sequence of a classical molecular gene will be strongly context dependent, as Dawkins himself has often stressed, although in this case there is likely to be some measure according to which the effect is less context dependent than that of a single nucleotide.
It is outside the scope of this article to speculate about why Dawkins rejects the concept of the “selfish nucleotide” (see Neumann-Held 1998, Sterelny and Griffiths 1999). We believe that the idea of the “selfish nucleotide” is in fact a straightforward, logical consequence of Dawkins' assumptions that evolutionary genes are to be defined independently of molecular genes and that they are, nevertheless, DNA sequences. It is this last assumption that we now question.
Setting the evolutionary gene free
The evolutionary gene, as interpreted by Dawkins, is any DNA segment that causes a difference in phenotypic adaptation between members of a population. But evolutionary genes so defined cannot account for most evolutionary change. First, the evolutionary role that Dawkins describes—a heritable difference maker—need not be filled by a DNA sequence at all. A variation in DNA methylation pattern or in any other epigenetic inheritance system can have the same effect (Jablonka and Szathmary 1995). To capture all evolutionary change, it would be necessary to say that an evolutionary gene is any heritable change in an input to development that causes a difference in phenotypic adaptation (Sterelny et al. 1996). Second, empirical and theoretical research does not support the idea that when a heritable phenotypic character spreads by selection, a specific DNA sequence that acts as a difference maker with respect to that trait need also be spread. Particularly in regard to quantitative traits, which are influenced by many genes, it can be shown that the effects of single genes may often be too small to predict their differential spreading on the basis of selection for the phenotypic trait (Wagner 1988, 1990). Although a certain phenotypic trait can “win out” over the alternatives in a population over time, the successful variant can often be realized by different combinations of genes. Consequently, the adaptation of a trait by natural selection does not require consistent selection at the level of the genes. Although there are replicators that participate in the selection processes, “none of them ‘wins a competition’ since none of them is singled out by selection” (Wagner 1990).
The same point has been nicely demonstrated by H. Frederick Nijhout and Susan Paulsen, who modeled selection for wing patterns in butterflies in a model epigenetic system (Nijhout and Paulsen 1997). Their model contains six developmental parameters controlling the distribution of a diffusible morphogen. The model assumes that each parameter is determined by a single locus and that the population contains two alleles at each locus, one associated with a high parameter value, the other with a low value. Even in this simple genetic system, the authors conclude that “whether a particular gene is perceived to be a major gene, a minor gene or even a neutral gene depends entirely on the genetic background in which it occurs, and this apparent attribute of a gene can change rapidly in the course of selection on the phenotype” (Nijhout and Paulsen 1997).
Because it cannot be assumed that each evolutionary gene corresponds to a particular DNA sequence, evolutionary genes are best conceived in an earlier sense of “gene,” that is, as units of particulate inheritance. Thus, an evolutionary gene is a heritable potential for a phenotypic (or extended phenotypic) trait. As such, an evolutionary gene is a theoretical entity in the same sense that an organism's “fitness” is a theoretical entity. The postulation of “fitnesses” is justified by the ability to estimate them and to use them in a powerful predictive theory—population genetics. The theoretical value of fitnesses is not impugned by the inability to identify them with specific structures at a lower level of analysis. An organism inherits its fitness from its fit parents, but not as a DNA sequence coding for that fitness. We suggest that evolutionary genes be treated in the same way. An evolutionary gene, such as the evolutionary gene for the favored wing pattern, need not correspond to any part of the organism at a lower level of analysis, such as Nijhout and Paulsen's (1997) theoretical molecular genes.
This way of viewing evolutionary genes may strike many people as odd. The idea that all genes, however conceptualized, are segments of DNA is deeply entrenched. But setting the evolutionary gene free has important payoffs—helping to explain, for example, why the evolutionary gene concept has been used so productively in evolutionary game theory, a discipline that dispenses almost entirely with the idea that “genes” are inherited in a Mendelian fashion.
Most important, however, this change makes it clear that the legitimacy of the evolutionary gene concept rests on the success of the gene selectionist approach to phenotypic evolution. In the past, evolutionary biologists have been too ready to assume that evolutionary gene selectionism must be the right theoretical perspective simply because molecular genes play several key roles in contemporary biology. In particular, the atomistic approach to pheno-types embodied in gene selectionism has been accepted because, linkage aside, molecular genes are inherited independently of one another. But this truth about the molecular gene may not extend to evolutionary genes if, as many developmental biologists believe, the phenotype consists of relatively large developmental modules that function as single units in evolution (Raff 1996, Wagner 1996). If this latter theoretical perspective is correct, then it would be a severe error to atomize these modules into separate “evolutionary genes” and to model their independent evolution. With regard to issues such as phenotypic atomism, the evolutionary gene concept has to be defended on its own ground, that is, in the context of evolutionary theory and model building. Whether such a defense is possible remains to be seen.
Conclusions
We have distinguished two different conceptions of the gene: the molecular gene and the evolutionary gene. In both cases, it is necessary to distinguish between genes and bare DNA. In the case of the molecular gene, we have proposed that the gene is the entire molecular process underlying the capacity to express a particular polypeptide product.
We have followed Dawkins in arguing that evolutionary genes need not be molecular genes but have argued against his view that they are simply more loosely defined segments of DNA. Instead, we suggest that an evolutionary gene is a theoretical entity with a role in a particular, atomistic approach to the selection of phenotypic and extended phenotypic traits. Evolutionary genes need not, and often do not, correspond to specific stretches of DNA.
Acknowledgments
In preparing this article we have benefitted from discussions with Robin D. Knight.
References cited
Author notes
1 (e-mail: p.griffiths@scifac.usyd.edu.au) is director of the Unit for History & Philosophy of Science at the University of Sydney, NSW 2006 Australia.
2 (e-mail: eva.neumann-held@dlr.de) is a member of the scientific staff of the European Academy for the Study of Consequences of Scientific and Technological Advance Ltd, Bad Neuenahr-Ahrweiler, Germany D-53474.