Abstract
Lipid nanoparticles have transformed the drug delivery field enhancing the therapeutic drug performance of small molecules and biologics with several approved drug products. However, in industry, these more complex drug delivery systems such as liposomes require more material and time to develop. Here, we report a liposome and lipodisk decision tree with model compounds of diverse physicochemical properties to understand how to resourcefully optimize encapsulation efficiency (EE) for these lipid-based drug delivery systems. We have identified trends with physicochemical properties such as Log P, where higher Log P compounds such as curcumin were able to efficiently load into the lipid bilayer resulting in high EE with altering the drug/lipid (D/L) ratio. Moderate Log P compounds such as cyclosporine A and dexamethasone had significantly higher encapsulation in lipodisks, which contain higher amounts of PEG lipid compared to liposomes. The EE of negative Log P compounds, like acyclovir, remained low regardless of altering the D/L ratio and PEG concentrations. In this study, microfluidic techniques were employed to fabricate liposomes and lipodisks formulations allowing for a reproducible strategy for formulation development. Both liposome and lipodisk of curcumin demonstrated enhanced in vivo performance compared with a conventional formulation in the rat pharmacokinetic study. This combination of approaches with multiple model compounds and lipid-based drug delivery systems provides a systematic guidance to effective strategies to generate higher EE with minimal drug waste and expedite the process for preclinical development when applied to industry compounds.
Similar content being viewed by others
INTRODUCTION
As a revolutionizing carrier introduced by Bangham et al. in 1965, liposomes have exhibited favorable aspects in drug delivery by demonstrating their biocompatible features, enhanced pharmacokinetic profiles, and reduced off-target toxicity (1,2,3,4,5,6,7). Since then, liposomes and lipid nanoparticle formulations have led to 23 approved products in the USA and Europe over the past 30 years (8). In the late 1990s, most liposome formulations were dedicated to cancer therapy and administered intravenously (9, 10). Recently, the delivery approach of liposome or lipid nanoparticle formulation has shifted its paradigm from small molecule delivery to other therapeutic modalities, such as oligonucleotides (11, 12), CRISPR (13,14,15), and DNA/mRNA antigens (16,17,18,19), the most prominent of which were used to develop and deliver COVID-19 vaccines (20, 21).
The self-assembly characteristics of a liposome results in a closed structure consisting of phospholipid bilayer membranes composed of an amphiphilic structure of polar head groups and hydrophobic alkyl tails surrounding an internal aqueous core. These unique bilayer structures expand the versatility of liposomes as potential delivery carriers that can encapsulate hydrophilic compounds in the aqueous core and hydrophobic compounds in the lipid bilayers (5, 22). Multiple factors impact the liposome design, and lipid composition is one of the critical aspects that govern the quality of liposome formulations (23, 24). In general, the formulation optimization of liposomes is based on the structure of each target molecule and is mostly performed on a case-by-case basis. The lack of a general decision tree or platform formulation approach can lead to a lengthy development cycle with significant resource commitments. One of the main reasons most preclinical formulations shy away from novel formulations instead of the conventional formulations is to reduce drug material requirements and a prolonged developmental timeline. However, drug discovery is becoming increasingly more challenging, and if novel formulations are not partnered in the preclinical stage, promising drug candidates may be missed due to sub-optimal delivery of drug candidates. Therefore, it is critical to create an efficient and general approach to guide early liposome formulation development based on the physicochemical properties of drug compounds. This decision tree will enable liposome formulations to be more accessible during early discovery support by significantly reducing the resources during the formulation optimization.
This study evaluated five model compounds with varying physicochemical properties, including Log P, solubility, pKa, etc., to identify critical properties that influence liposome performance. This design would provide a systematic discovery formulation scheme to facilitate the process of formulation development. Microfluidic technology has been employed to produce numerous nanoparticle systems including liposomes (25, 26). The rapid and tunable system provides precise control of nanoparticle formation. As a result, the microfluidic technique was selected as the primary formulation method to prepare the lipid-based nanoparticles in this study. Compared with the traditional techniques for liposome preparation, such as thin-film hydration (27, 28), membrane extrusion, and reverse-phase evaporation, the microfluidic method offers a more efficient and reproducible strategy of liposome formulations in a well-controlled manner at nanoliter to liters scale (27,28,29,30). Microfluidics has revolutionized the development and continuous manufacturing of liposomes enabling clinical scale production (31). Early clinical liposome formulations such as Doxil were prepared in a lengthy multistep extrusion process. The difficulties in scalability of the system lead to the development of microfluidic technology that can overcome limitations in traditional liposome preparation techniques (26).
Among all the final critical quality attributes, encapsulation efficiency (EE) and particle size distribution of liposomes were selected to assess the formulation process. The particle size of liposomes prepared by microfluidic methods was homogeneous with sizes between 115 and 144 nm regardless of the model compounds tested. Therefore, the main effort was to focus on the improvement of EE. Several formulation approaches were adapted to improve the EE, including altering drug to lipid (D/L) ratio (23, 32), leveraging remote loading (33, 34), and increasing polyethylene glycol (PEG) ratios, which promotes the morphology change from liposomes to lipodisks. Lipodisks consist of a disk-like structure where the phospholipids form a flat lipid surface, and the PEG-lipids are located at the highly curved rim (35,36,37). Many published results suggest that lipodisks are promising nanocarriers, and the rich contents of PEG may promote improved EE (38,39,40,41). This is the first report adopting microfluidic methods to prepare lipodisk formulations to the best of our knowledge. Two microfluidic platforms, NanoAssemblr® Benchtop and NxGen Blaze™, were used to prepare liposome and lipodisk formulations in this study. The formulation parameters and quality attributes were translatable between these two platforms at various batch sizes. After optimizing the formulations, our work also provides the first example to have the head-to-head in vivo performance comparisons of conventional, liposome, and lipodisk formulations in the rat intravenous pharmacokinetic study.
MATERIALS AND METHODS
Materials
The lipids including 1,2-distearoyl-sn-glycero-3-phosphocholine (DSPC), 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] (DSPE-PEG2000), and cholesterol were purchased from Avanti (AL, USA). Ethanol and DMSO were purchased from JT. Baker (PA, USA). PEG-400 was purchased from Spectrum (CA, USA). The model compounds cyclosporine A was purchased from Sigma (MO, USA), dexamethasone was purchased from TRC (SC, USA), acyclovir was purchased from Spectrum (CA, USA), and curcumin was purchased from Alfa Aesar (MA, USA). The dialysis cassettes were purchased from Fisher (NH, USA). The hollow fiber TFF cartridges (D02-E300-05-N) were purchased from Repligen (MA, USA).
Liposome and Lipodisk Preparation
Liposomes with a diameter of ~ 120 nm were prepared through microfluidics with the NanoAssemblr® Benchtop (Precision NanoSystems, BC, Canada) as previously described (29, 42, 43). Briefly DSPC, DSPE-PEG2000, and cholesterol (61.56%:18.8%:19.56% (w/w)) were dissolved at 30 mg/mL in ethanol. The hydrophobic compounds, dexamethasone, cyclosporine A, doxorubicin, and curcumin, were dissolved in the lipid ethanol solution. For hydrophobic model compounds, the D/L mixtures were prepared by varying model compound concentrations to achieve various D/L ratios, 0.01, 0.03, 0.1, and 0.2 (w/w). To achieve a D/L 0.1 for doxorubicin, 6 mg/mL total lipids was prepared with 0.6 mg/mL of doxorubicin due to the lower solubility of doxorubicin in ethanol. Aqueous soluble compounds, acyclovir, were added to the aqueous phase at 1 mg/mL. For aqueous soluble compounds, the D/L mixtures were prepared by varying total lipid concentrations (D/L 0.03, 0.1, 0.2 (w/w)). The organic ethanol solution was mixed with the aqueous phase at a 3:1 (aqu:org) flow rate ratio (FRR) through the NanoAssemblr at a total flow rate (TFR) of 10 mL/min.
Lipodisks were prepared with the same technique as described above with the exception of a lipid composition modification to DSPC:DSPE-PEG:cholesterol (18.8%:61.56%:19.56% (w/w)). Liposomes and lipodisks were purified by centrifugation (Beckman Coulter Allegra X-12 Centrifuge, CA, USA) at 2000 g for 10 min to remove precipitated hydrophobic drug, then dialysis (20-kDa MWCO Slide-A-Lyzer) was performed over 7 h with 3 PBS buffer exchanges to remove the soluble unencapsulated drug, extra lipids, and organic solvent.
Doxorubicin remote loading was performed as previously described (44). Briefly, blank liposomes were prepared with 300 mM citrate buffer pH 4 as the aqueous solvent. Buffer exchange was performed through dialysis with 10 mM [4-(2-hydroxyethyl)-piperazino]-ethanesulfonic acid (HEPES) buffer 140 mM NaCl pH 7.4 to replace the extra-liposomal solution. Doxorubicin HCl was then added to the liposomes. Purification was performed as described above with centrifugation and dialysis.
Characterization of Model Compound-Loaded Liposomes and Lipodisks
Particle size distribution and polydispersity index (Pdi) were characterized by dynamic light scattering (DLS) using a Wyatt DynaPro Plate Reader III (Wyatt Technology, CA, USA). Liposomes and lipodisks were diluted 5–tenfold depending on the starting concentration. Then 30 µL was placed in a 384 well black wall, clear bottom polystyrene microplate (Corning, NY, USA) and analyzed for particle diameter and polydispersity index. Each particle size measurement consisted of 15 acquisitions with a 3 s acquisition time at 25 °C.
Curcumin liposome and lipodisk structures were analyzed by cryo-transmission electron microscope (cryo-TEM). Briefly, an EMGP2 automatic plunge freezer (Leica Microsystems, IL, USA) system was used to prepare the samples. Samples were placed on a lacey carbon copper EM grid overlaid with graphene oxide (Electron Microscopy Sciences GOL200CU100, PA, USA). Excess liquid was blotted and grids were transferred to a Glacios 200-keV electron microscope (Thermo Fisher Scientific, OR, USA) for imaging.
Encapsulation efficiency and in vivo formulation dose check was determined for curcumin, dexamethasone, doxorubicin, and acyclovir by a reverse-phase high-performance liquid chromatography (RP-HPLC) system (Agilent Technologies, Santa Clara, CA USA) with Waters RP18 XTerra column (4.6 × 50 mm, 3.5 µm particle size) (MA, USA) at 254 nm detection wavelength. A gradient elution method was used with mobile phase A of 0.05% v/v TFA in water and mobile phase B of 0.05% v/v TFA in ACN at a constant 1.25 mL/min flow rate. From 0 to 0.5 min, 98% mobile phase A and 2% mobile phase B equilibrated in the column. From 0.5 min to 3 min, mobile phase A was reduced to 35% and mobile phase B was increased to 65% and held for 1 min. Mobile phase A was then increased to 98% and mobile phase B decreased to 2% over 0.1 min, then this was held for 2.4 min. For CSA, a Waters C18 XBridge column (3.0 × 100 mm, 3.5 µm particle size) was used at 50 °C at 210 nm detection wavelength. An isocratic method was run with 20% of 0.05% TFA in water and 80% of 0.05% TFA in methanol for 5 min at a flow rate of 1.25 mL/min. Total drug was measured prior to purification. After purification by centrifugation and dialysis, the encapsulated drug was measured. Encapsulation efficiency was based on the weight concentrations and calculated using the equation below:
In Vivo Formulation Preparation
Three groups of formulations for in vivo studies were prepared for this study, including liposome, lipodisk, and solution formulations containing 1 mg/mL curcumin concentrations. Liposomes and lipodisks were prepared under the microfluidic methods as described above at a D/L 0.01. The NxGen Blaze™ (Precision NanoSystems, BC, Canada) was employed to supply large-scale formulations and the formulation parameters were similar to the settings used in the NanoAssemblr. Purification was performed by centrifugation at 2000 g for 10 min followed by tangential flow filtration (TFF) (KrosFlo KR2i, Repligen, MA, USA) to remove unencapsulated drug and organic solvent. For TFF, the liposome or lipodisks were loaded onto the system and passed through a 300 kDa mPES Hollow Fiber Membrane for 8 diafiltration volumes (DVs) with PBS then concentrated to the target concentration. A solution formulation of curcumin was prepared in a 10% DMSO, 35% PEG400, and 55% water vehicle. The final dose check performed by RP-LC confirmed that the concentration of three dosing arms was all within the target concentration range.
In Vivo Intravenous Pharmacokinetic Study in Rats
Animal studies were approved by the Institutional Animal Care and Use Committee (IACUC) of Genentech and in accordance with federal guidelines. A total of 9 Sprague Dawley rats weighing ~ 250 to 310 g were acquired from Charles River Labs (Wilmington, MA, USA) and divided evenly into 3 treatment groups (2 females, 1 male per group). Each group was intravenously administered 5 mg/kg of curcumin in either liposome, lipodisk, or solution form by IV bolus through a jugular vein cannula. Blood collection was performed at the following time points: predose, 2 min (0.033 h), 5 min (0.083 h), 15 min (0.25 h), 30 min (0.5 h), and 1, 2, 4, 8, 24 and 48-h post-dose. All samples were collected through a femoral artery cannula by a Culex automated blood sampling machine (West Lafayette, IN, USA) and placed in tubes containing K2 EDTA. Blood samples were centrifuged for 5 min at 3700 rpm (2235 G’s) for plasma collection.
Drug plasma concentrations were quantified via LC/MS/MS. For this analysis, plasma concentrations were measured by a liquid chromatographic-tandem mass spectrometry (LC/MS/MS) assay method with an internal standard.
Statistical Analysis
One-way analysis of variance (ANOVA) with a Dunnett multiple post-test comparison or t-test was used to determine significance between each group using GraphPad Prism 8.0. All plots express mean ± standard deviation (SD) where n.s. indicates not significant, * indicates p < 0.05, ** indicated p < 0.01, and *** indicated p < 0.001. Pharmacokinetic parameters from non-compartmental analysis were calculated using the trapezoidal rule and relevant pharmacokinetic equations.
RESULTS AND DISCUSSION
The Selection of Tool Compounds
Model compounds were chosen with various structures and physicochemical properties (Fig. 1 and Table I). Compounds were selected with a wide range of parameters to understand how the EE is influenced by properties such as molecular weight, water solubility, and Log P. The molecular weights range over threefold, the water solubility ranges over 3,000-fold, and the Log P values range over fourfold.
Due to complexity in synthesis as well as the investigation of many different types of chemical matter, early compounds in industry generally are made on small scales potentially limiting extensive formulation development. When developing liposomes, low encapsulation efficacy can lead to significant waste of compounds. Investigating and optimizing encapsulation efficiency with model compounds can help inform strategies for encapsulating compounds in industry, limiting material requirements. Additionally, model compounds can help expedite the process for development of promising liposome encapsulated compounds. Utilizing various physicochemical properties will allow for application across a wide variety of chemical matter.
Liposome Formulation and Characterization
Model compounds were encapsulated in liposomes through microfluidics. Microfluidic mixing is a scalable and reproducible technology that can be applied to liposome manufacturing (45). TFR and FRR were selected based on previous studies to achieve the desired 100–150 nm size (29, 45). The size and polydispersity were measured through DLS (Table II). The blank liposomes (117 nm) and liposomes encapsulated with the model compounds at a 0.1 D/L (w/w) ratio remained consistent around 120 nm, indicating that the model compounds do not significantly influence the size of the liposomes.
Next, EE was measured by RP-LC to investigate the model compound encapsulation with various molecular properties. EE is an important parameter to understand the interaction of the model compounds and the liposomes. Optimizing encapsulation improves the efficiency of the process along with minimizing compound waste during the liposome formation process. To compare EE for the model compounds, the D/L ratio was constant at 0.1, which is in the range for literature values of drug encapsulation in liposomes (46, 47). EE was the highest for cyclosporine A (CSA) (21.96%) and decreased to 0.7% for acyclovir (Table II). Log P trended with EE where the higher Log P compounds curcumin (11.41%) and CSA (21.96%) lead to higher EE compared to lower Log P values, doxorubicin (5.80%), dexamethasone (1.49%), and acyclovir (0.7%) (Fig. 2). The findings with Log P and encapsulations are in line with what has been seen in literature. Log P has been shown to influence encapsulation of drugs in liposomes where the higher Log P compounds lead to higher encapsulation efficiency (48). Generally, the higher Log P compounds entrap almost completely in the lipid bilayer while intermediate partition coefficient can partition between the lipid and aqueous phase leading to poor encapsulation at the D/L ratio 0.1 (49). However, the EE of all 5 tool compounds was still low, and further optimization was needed.
a Model compound’s Log P correlates to compound’s encapsulation efficiency with D/L ratio of 0.1. Altering drug/lipid ratio impacts encapsulation efficiency. b Curcumin. c Cyclosporine A. d Dexamethasone. e Acyclovir. Data represents mean ± s.d., N = 3. One-way ANOVA, * indicates p < 0.05, ** p < 0.01, *** p < 0.001 w.r.t. D/L ratio of 0.1
Formulation Approaches to Improve Encapsulation Efficiency
We next investigated strategies to improve EE, first altering the D/L ratio. Lipid amounts were kept constant while the drug amount was altered for the hydrophobic compounds (Log P > 1.89). In general, a nearly twofold increase of EE corresponded with decreasing D/L ratio for all 4 tool compounds (Fig. 2). Curcumin with the highest Log P in the model compounds has shown the most remarkable improvement in EE with altering D/L ratios. With altering the D/L ratio of curcumin, the EE significantly increased where the encapsulation increased from 11.41% at a D/L ratio of 0.1 to 82.4% with a D/L ratio of 0.01 (Fig. 2). The low EE at the higher D/L ratios indicates that the amount of curcumin has saturated the lipid bilayer leading to the increased amount of free drug. The drug cargo can affect the stability and release kinetics in liposome systems. More hydrophobic drugs have been shown in vitro to result in faster release kinetics (50). In vivo release through lipid exchange would lead to faster drug release with the hydrophobic drugs saturated in the lipid bilayer compared to hydrophilic drugs in core (51). For compounds with moderate or negative Log P, altering the D/L ratio improved the overall EE, but the impact was still limited. Since CSA, dexamethasone, and acyclovir only had modest changes in EE with altering D/L ratio, we investigated another strategy by altering the liposome structure to understand the effects of EE.
Liposome vs. Lipodisk
By increasing the PEG ratios from 18.8 to 62%, the lipid nanoparticle structures transform from liposomes to lipodisks. Lipodisks consist of a lipid bilayer in a discoidal shape that can encapsulate hydrophobic compounds within the lipid bilayer along with the potential to interact with compounds on the surface. Compared to liposomes which are ~ 150 nm in size, lipodisks are smaller at ~ 50 nm (Fig. 3a). Lipodisks have mainly been utilized as biomimetic membranes; however, there has been promising evidence for application in the drug delivery field. Lipodisks have been shown to be biocompatible, increase drug circulation half-life, and increase tumor accumulation (36, 40, 52). CSA, dexamethasone, curcumin, and acyclovir were encapsulated in lipodisks with the microfluidic system, the NanoAssemblr, utilizing a 0.03 D/L ratio for CSA and acyclovir and a 0.01 D/L ratio for dexamethasone and curcumin. For consistency, the TFR and FRR parameters in the microfluidic method were maintained when making the lipodisks and liposomes. The difference in EE between liposomes and lipodisks was measured. The EE for CSA significantly increased to 80% when encapsulated in lipodisks compared to liposomes at 39.8% EE (Fig. 3b). Lipodisks increased EE of dexamethasone from 2.3 to 9.7% with a D/L ratio 0.03 (Fig. S1). With decreasing the D/L ratio to 0.01, dexamethasone EE increased to 27.46% in lipodisks (Fig. 3c). The combination of the lipodisk strategy and altering D/L ratio led to a synergistic effect to optimize and increase EE. Curcumin lipodisk and liposome EE remained constant while the acyclovir EE did not significantly increase in lipodisks (Fig. 3d,e). CSA and dexamethasone have moderate Log P values generally resulting in poor EE. Here, with lipodisks, we have shown promise to increase EE of these model compounds with moderate Log P values. Lipodisks did not improve the EE of acyclovir, an aqueous soluble compound with a low Log P (− 1.56). As the lipodisks lack an aqueous core, the minimal EE of acyclovir was expected. Since the EE was similar for the curcumin loaded in the liposomes and lipodisks, we further investigated the effects of these two delivery systems in vivo.
Altering DSPE-PEG (2000) amount in liposome composition. Increasing DSPE-PEG (2000) percent a decreases liposome size, b increases encapsulation of cyclosporine A with a D/L ratio 0.03, c increases dexamethasone encapsulation with a D/L ratio 0.01, d no significant change with curcumin at a D/L ratio 0.01, e no significant change with acyclovir with a D/L ratio 0.03. Data represents mean ± s.d., N = 3. One way ANOVA, n.s. indicates not significant, *** indicates p < 0.001 w.r.t. 18.8% DSPE-PEG (2000)
Remote Loading
Generally, the alteration of the D/L ratio improves the EE for most conditions and a similar trend was also observed from doxorubicin. EE increased from 5.8 to 39% when altering the D/L from 0.1 to 0.03 (Fig. 4). To further increase EE, another strategy, remote loading, was investigated. Remote loading utilizes a pH gradient to efficiently encapsulate drugs into the core of preformed liposomes (53). As a technique for active encapsulation, the remote loading approach can result in high EEs and minimal waste of drugs (54). Doxorubicin is a weakly acidic compound as an ideal tool compound to demonstrate the concept of remote loading and the EE increased with remote loading from 39 to 95.9% at a 0.03 D/L ratio (Fig. 4). A synergistic effect is seen with altering D/L ratio and remote loading where EE can reach over 90%. For remote loading the external buffer of blank liposomes in citrate buffer at pH 4 was exchanged with HEPES buffer pH 7.4 to create a pH gradient across the liposome. Doxorubicin added to the external HEPES buffer in a nonionized state resulting in permeation across the lipid membrane. Once internalized, doxorubicin is ionized in the low pH buffer limiting doxorubicin from crossing the lipid membrane resulting in an accumulation of doxorubicin in the core of the liposome. These findings with high EE for doxorubicin are in line with literature where EE is close to 100% utilizing the remote loading strategy (44, 53). However, this technique generally works best for amphiphilic weak acids and bases, limiting the application across different chemical compounds (55).
In Vivo IV Pharmacokinetics of Liposome and Lipodisk of Curcumin in Rats
For larger scale studies, the Blaze instrument was utilized to make liposomes and lipodisks loaded with curcumin to understand the in vivo properties. Ethanol was removed and the liposomes and lipodisks were concentrated with TFF. Blaze and TFF allow for efficient scaling to generate reproducibility and precise liposomes and lipodisks for further applications. Cryo-EM was used to visualize the structural difference between the liposomes and lipodisks (Fig. 5). The particle size distribution of liposome and lipodisk formulations fabricated by NanoAssemblr and Blaze is comparable, and the formulation parameters between the two methods were also translatable (Fig. S2). Liposome consisted of a lipid bilayer with an aqueous core. With the increase of DSPE-PEG, the cryo-EM depicted a dispersion of nanosized disk particles, lipodisks. The size of the liposomes and lipodisks remained stable over 8 days at 4 °C (Fig. 5).
To understand the influence of liposome and lipodisk encapsulation on the plasma circulation of drugs, a pharmacokinetic study was performed. Sprague Dawley rats were divided into 3 cohorts and administered an equivalent dose of curcumin (5 mg/kg) in either solution, liposome, or lipodisk formulations. Curcumin was chosen from the 5 test compounds based on the known rapid clearance in rats (56, 57) and encapsulation efficiency (~ 82.4%) in both liposomes and lipodisks (Fig. 3d). The plasma concentration time profiles for each cohort are presented in Fig. 6.
Plasma concentration versus time profile of curcumin (5 mg/kg) administered IV in solution, liposome, or lipodisk formulation. From 0 to 8 h, a trend is observed with higher concentrations of curcumin in plasma associated with liposome (dashed line, triangle ▲) and lipodisk formulations (dashed line, square ■) compared to the solution formulation (solid line, circle ●). Data represents mean ± s.d., N = 3
If curcumin were embedded in the lipid bilayers of liposome or lipodisks, then it would likely be inaccessible to the endogenous clearance mechanism of freely soluble drug. The plasma concentration time profile of the liposome and lipodisk formulations (compared to the solution control) should then be enhanced due to the protection of curcumin from in vivo metabolism. From our in vivo results, we observe a trend of higher plasma concentrations in the lipodisk and liposome cohorts. The average measured plasma concentrations of curcumin were 1.5 to 2.5 fold greater than the free solution over the time course when given the same dose (Fig. 6). We next analyzed the plasma drug concentration time courses by non-compartmental analysis to estimate the pharmacokinetic parameters of each cohort. These parameters are presented in Table III. Similar to the plasma concentration time profile, both liposomes and lipodisks formulations showed higher AUCs than the solution control (1.56-fold and 1.38-fold, respectively). Additional pharmacokinetic parameters such as clearance and Vss change proportionally to the AUC for each formulation but do not reach statistical significance (p > 0.05). Given the high encapsulation efficiency of curcumin in the liposomes and lipodisks, a greater change in clearance was expected with the assumption that encapsulation would shield curcumin from metabolism. However, it is possible that encapsulation of curcumin can trigger alternative clearance mechanisms. For example, nanoparticles can accumulate in the liver through endocytosis by Kupffer cells (58). The subsequent lysosomal degradation of liposomes or lipodisks could represent a distinct clearance mechanism not present in the solution cohort (59). Additional experiments to identify the distinct clearance mechanisms of liposomes or lipodisks are in consideration but fall outside the scope of the current study. The average Vss of curcumin is much higher in the solution cohort compared to liposomes (4.1-fold) or lipodisks (2.8-fold), although significant variability is observed in the parameter estimation for the solution cohort. The reduction of curcumin Vss in liposomes or lipodisks aligns with our initial expectation as the encapsulated drug is restricted from passive distribution into peripheral compartments.
The focus of this report is to provide a systematic discovery scheme to facilitate formulation development. The current work provides a foundation for the optimization of critical quality attributes and demonstrates the potential effect of encapsulation on the in vivo pharmacokinetics of curcumin. From our 5 test compounds, curcumin is the most lipophilic and expected to have the highest affinity for the lipid bilayers of the liposomes or lipodisks. However, the in vivo release of a drug from a nanoparticle carrier is not only a function of the drug’s affinity for the carrier but also the various interactions that occur in the biological milieu. In previous reports, it has been shown that hydrophobic drugs may exhibit a greater release from lipid-based nanoparticle carriers when compared to hydrophilic drugs (60). Hydrophobic drugs that reside on the surface of lipid bilayers may interact with plasma proteins and lead to greater than expected in vivo release compared to in vitro measurements. In contrast, hydrophilic drugs residing in the aqueous core of liposomes or the pegylated borders of lipodisks may have greater retention in the nanocarrier after interaction with plasma proteins. In the case of curcumin, there is a known interaction between free curcumin and human serum albumin and fibrinogen which may support this hypothesis (61) but further study is required. Additionally, in future studies, it would be of interest to investigate how further optimization of these attributes will translate in vivo for all five chemically diverse compounds. The integration of these data can facilitate novel formulations in the preclinical stage to develop the best drug candidates.
Decision Trees of Lipid Nanoparticle Formulations
In Fig. 7, we outline the decision trees to achieve the highest EE in lipid nanoparticle formulations for the model compounds. The workflow suggested by the decision tree represents the most effective approach for each compound category. For compounds with Log P greater than 3 (curcumin), either altering the D/L ratio or increasing the DSPE-PEG% significantly increased EE to > 80%. For compounds with moderate Log P (0 < Log P < 3), increasing the DSPE-PEG% to generate lipodisks is the most effective approach to improve the EE. Remote loading is the most competent approach for weak acid or weak based compounds with an ionizable pKa. The combination of altering the D/L ratio with other methods could also achieve the synergistic effect. For hydrophilic compounds with high aqueous solubility and negative Log P such as acyclovir, altering the D/L ratio and encapsulating in lipodisks lead to low EE. Further development would be necessary to understand the driving factors to increase EE for compounds that fall into this category. The selected D/L ratio for each tool compound can be found in Table SI. This decision tree offered a systematic approach to prepare liposome and lipodisk via the microfluidic method, which provides facile and reproducible formulations. Our formulation strategy is simply based on the Log P value to achieve the optimized EE and this approach could save time and resources when developing drug-loaded lipid nanoparticles at the preclinical stage.
CONCLUSION
We have developed a systematic process outlined in a decision tree to determine the optimal strategy to generate lipid-based nanoparticles with optimized encapsulation efficiency based on the compound’s physicochemical properties such as Log P and ionization. Furthermore, we have identified a novel and promising strategy via the microfluidic preparation method to increase the EE for moderate Log P compounds through encapsulation with lipodisks. The in vivo PK studies exemplify the scalability and translatability of these promising lipodisks and liposomes delivery systems to increase drug circulation and retention. The optimization studies forming the decision tree will save significant material and time in the future development of these lipid-based nanoparticle formulations.
References
Bangham AD, Standish MM, Watkins JC. Diffusion of univalent ions across the lamellae of swollen phospholipids. J Mol Biol. 1965;13(1):238–52.
Yeh M-K, Chang H-I, Cheng M-Y. Clinical development of liposome-based drugs: formulation, characterization, and therapeutic efficacy. Int J Nanomedicine. 2012;7:49–60.
Zylberberg C, Matosevic S. Pharmaceutical liposomal drug delivery: a review of new delivery systems and a look at the regulatory landscape. Drug Deliv. 2016;23(9):1–11.
Immordino M, Dosio F, Cattel L. Stealth liposomes: review of the basic science, rationale, and clinical applications, existing and potential. Int J Nanomedicine. 2006;1(3):297–315.
Allen TM, Cullis PR. Liposomal drug delivery systems: from concept to clinical applications. Adv Drug Deliv Rev. 2013;65(1):36–48.
Akbarzadeh A, Rezaei-Sadabady R, Davaran S, Woo Joo S, Zarghami N, Hanifehpour Y, et al. Liposome: classification, preparation, and applications. 2013;8:102.
Sharma A, Sharma US. Liposomes in drug delivery: progress and limitations. Int J Pharm. 1997;154(2):123–40.
Fan Y, Marioli M, Zhang K. Analytical characterization of liposomes and other lipid nanoparticles for drug delivery. J Pharm Biomed Anal. 2020;192:113642.
Barenholz Y. Doxil® - the first FDA-approved nano-drug: lessons learned. J Control Release. 2012;160(2):117–34.
Yingchoncharoen P, Kalinowski DS, Richardson DR. Lipid-based drug delivery systems in cancer therapy: what is available and what is yet to come. Pharmacol Rev. 2016;68(3):701–87.
Wang Y, Miao L, Satterlee A, Huang L. Delivery of oligonucleotides with lipid nanoparticles. Adv Drug Deliv Rev. 2015;87:68–80.
Smith CIE, Zain R. Therapeutic oligonucleotides: state of the art. Annu Rev Pharmacol Toxicol. 2018;59(1):1–26.
Finn JD, Smith AR, Patel MC, Shaw L, Youniss MR, van Heteren J, et al. A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing. Cell Rep. 2018;22(9):2227–35.
Rosenblum D, Gutkin A, Kedmi R, Ramishetti S, Veiga N, Jacobi AM, et al. CRISPR-Cas9 genome editing using targeted lipid nanoparticles for cancer therapy. Sci Adv. 2020;6(47):eabc9450.
Lino CA, Harper JC, Carney JP, Timlin JA. Delivering CRISPR: a review of the challenges and approaches. Drug Deliv. 2018;25(1):1234–57.
Arteta MY, Kjellman T, Bartesaghi S, Wallin S, Wu X, Kvist AJ, et al. Successful reprogramming of cellular protein production through mRNA delivered by functionalized lipid nanoparticles. Proc Natl Acad Sci. 2018;115(15):201720542.
Granot Y, Peer D. Delivering the right message: challenges and opportunities in lipid nanoparticles-mediated modified mRNA therapeutics—an innate immune system standpoint. Semin Immunol. 2017;34:68–77.
Kulkarni JA, Myhre JL, Chen S, Tam YYC, Danescu A, Richman JM, et al. Design of lipid nanoparticles for in vitro and in vivo delivery of plasmid DNA. Nanomed Nanotechnol Biol Med. 2017;13(4):1377–87.
Olbrich C, Bakowsky U, Lehr C-M, Muller R, Kneuer C. Cationic solid-lipid nanoparticles can efficiently bind and transfect plasmid DNA. J Control Release. 2001;77(3):345–55.
McKay PF, Hu K, Blakney AK, Samnuan K, Brown JC, Penn R, et al. Self-amplifying RNA SARS-CoV-2 lipid nanoparticle vaccine candidate induces high neutralizing antibody titers in mice. Nat Commun. 2020;11(1):3523.
Schoenmaker L, Witzigmann D, Kulkarni JA, Verbeke R, Kersten G, Jiskoot W, et al. mRNA-lipid nanoparticle COVID-19 vaccines: structure and stability. Int J Pharm. 2021;601:120586.
Szoka F. Comparative properties and methods of preparation of lipid vesicles (liposomes). Ann Rev Biophys Bioeng. 1980;9.
Caracciolo G, Pozzi D, Capriotti AL, Cavaliere C, Piovesana S, Amenitsch H, et al. Lipid composition: a “key factor” for the rational manipulation of the liposome–protein corona by liposome design. RSC Adv. 2014;5(8):5967–75.
Yang K, Mesquita B, Horvatovich P, Salvati A. Tuning liposome composition to modulate corona formation in human serum and cellular uptake. Acta Biomater. 2020;106:314–27.
Tiboni M, Tiboni M, Pierro A, Del Papa M, Sparaventi S, Cespi M, et al. Microfluidics for nanomedicines manufacturing: an affordable and low-cost 3D printing approach. Int J Pharm. 2021;599(December 2020):120464.
Roces CB, Port EC, Daskalakis NN, Watts JA, Aylott JW, Halbert GW, et al. Rapid scale-up and production of active-loaded PEGylated liposomes. Int J Pharm. 2020;586(July):119566.
Al-Amin MD, Bellato F, Mastrotto F, Garofalo M, Malfanti A, Salmaso S, et al. Dexamethasone loaded liposomes by thin-film hydration and microfluidic procedures: formulation challenges. Int J Mol Sci. 2020;21(5):1611.
Zhigaltsev IV, Belliveau N, Hafez I, Leung AKK, Huft J, Hansen C, Cullis PR. Bottom-up design and synthesis of limit size lipid nanoparticle systems with aqueous and triglyceride cores using millisecond microfluidic mixing. Langmuir. 2012;28(7):3633–40. https://doi.org/10.1021/la204833h.
Forbes N, Hussain MT, Briuglia ML, Edwards DP, ter Horst JH, Szita N, et al. Rapid and scale-independent microfluidic manufacture of liposomes entrapping protein incorporating in-line purification and at-line size monitoring. Int J Pharm. 2019;10(556):68–81.
Hamano N, Böttger R, Lee SE, Yang Y, Kulkarni JA, Ip S, et al. Robust microfluidic technology and new lipid composition for fabrication of curcumin-loaded liposomes: effect on the anticancer activity and safety of cisplatin. Mol Pharm. 2019;16(9):3957–67.
Carugo D, Bottaro E, Owen J, Stride E, Nastruzzi C. Liposome production by microfluidics: potential and limiting factors. Sci Rep. 2016;6:1–15.
Mayer LD, Tai LC, Ko DS, Masin D, Ginsberg RS, Cullis PR, et al. Influence of vesicle size, lipid composition, and drug-to-lipid ratio on the biological activity of liposomal doxorubicin in mice. Cancer Res. 1989;49(21):5922–30.
Zucker D, Marcus D, Barenholz Y, Goldblum A. Liposome drugs’ loading efficiency: a working model based on loading conditions and drug’s physicochemical properties. J Control Release. 2009;139(1):73–80.
Sur S, Fries AC, Kinzler KW, Zhou S, Vogelstein B. Remote loading of preencapsulated drugs into stealth liposomes. Proc Natl Acad Sci. 2014;111(6):2283–8.
Ahlgren S, Fondell A, Gedda L, Edwards K. EGF-targeting lipodisks for specific delivery of poorly water-soluble anticancer agents to tumour cells. RSC Adv. 2017;7(36):22178–86.
Zetterberg MM, Ahlgren S, Hernández VA, Parveen N, Edwards K. Optimization of lipodisk properties by modification of the extent and density of the PEG corona. J Colloid Interface Sci. 2016;484:86–96.
Feng C, Zhang H, Chen J, Wang S, Xin Y, Qu Y, et al. Ratiometric co-encapsulation and co-delivery of doxorubicin and paclitaxel by tumor-targeted lipodisks for combination therapy of breast cancer. Int J Pharm. 2019;560:191–204.
Gao J, Xie C, Zhang M, Wei X, Yan Z, Ren Y, et al. RGD-modified lipid disks as drug carriers for tumor targeted drug delivery. Nanoscale. 2016;8(13):7209–16.
Lundsten S, Hernández VA, Gedda L, Sarén T, Brown CJ, Lane DP, et al. Tumor-targeted delivery of the p53-activating peptide VIP116 with PEG-stabilized lipodisks. Nanomaterials. 2020;10(4):783.
Ahlgren S, Reijmar K, Edwards K. Targeting lipodisks enable selective delivery of anticancer peptides to tumor cells. Nanomed Nanotechnol Biol Med. 2017;13(7):2325–8.
Duong-Thi M-D, Bergström M, Edwards K, Eriksson J, Ohlson S, Ying JTY, et al. Lipodisks integrated with weak affinity chromatography enable fragment screening of integral membrane proteins. Analyst. 2015;141(3):981–8.
Joshi S, Hussain MT, Roces CB, Anderluzzi G, Kastner E, Salmaso S, Kirby DJ, Perrie Y. Microfluidics based manufacture of liposomes simultaneously entrapping hydrophilic and lipophilic drugs. Int J Pharm. 2016;514(1):160–8. https://doi.org/10.1016/j.ijpharm.2016.09.027.
Kastner E, Verma V, Lowry D, Perrie Y. Microfluidic-controlled manufacture of liposomes for the solubilisation of a poorly water soluble drug. Int J Pharm. 2015;485(1–2):122–30.
Fritze A, Hens F, Kimpfler A, Schubert R, Peschka-Süss R. Remote loading of doxorubicin into liposomes driven by a transmembrane phosphate gradient. Biochim Biophys Acta - Biomembr. 2006;1758(10):1633–40.
Shah VM, Nguyen DX, Patel P, Cote B, Al-Fatease A, Pham Y, et al. Liposomes produced by microfluidics and extrusion: a comparison for scale-up purposes. Nanomed Nanotechnol Biol Med. 2019;18:146–56.
Johnston MJW, Semple SC, Klimuk SK, Edwards K, Eisenhardt ML, Leng EC, et al. Therapeutically optimized rates of drug release can be achieved by varying the drug-to-lipid ratio in liposomal vincristine formulations. Biochim Biophys Acta - Biomembr. 2006;1758(1):55–64.
Ong JCL, Sun F, Chan E. Development of stealth liposome coencapsulating doxorubicin and fluoxetine. J Liposome Res. 2011;21(4):261–71.
Nii T, Ishii F. Encapsulation efficiency of water-soluble and insoluble drugs in liposomes prepared by the microencapsulation vesicle method. Int J Pharm. 2005;298(1):198–205.
Gulati M, Grover M, Singh S, Singh M. Lipophilic drug derivatives in liposomes. Int J Pharm. 1998;165(2):129–68.
Khan D, Rezler E, Lauer-Fields J, Fields G. Effects of drug hydrophobicity on liposomal stability. Chem Biol Drug Des. 2007;71(1):3–7.
Marc Ostro, Cullis P. Use of liposomes as injectable-drug delivery systems. Am J Hosp Pharm. 1989;46(8):1576–88.
Zhang W, Sun J, Liu Y, Tao M, Ai X, Su X, et al. PEG-stabilized bilayer nanodisks as carriers for doxorubicin delivery. Mol Pharm. 2014;11(10):3279–90.
Alyane M, Barratt G, Lahouel M. Remote loading of doxorubicin into liposomes by transmembrane pH gradient to reduce toxicity toward H9c2 cells. Saudi Pharm J. 2016;24(2):165–75.
Vakili-Ghartavol R, Rezayat SM, Faridi-Majidi R, Sadri K, Jaafari MR. Optimization of docetaxel loading conditions in liposomes: proposing potential products for metastatic breast carcinoma chemotherapy. Sci Rep. 2020;10(1):1–14. Available from: https://doi.org/10.1038/s41598-020-62501-1.
Cern A, Golbraikh A, Sedykh A, Tropsha A, Barenholz Y, Goldblum A. Quantitative structure - property relationship modeling of remote liposome loading of drugs. J Control Release. 2012;160(2):147–57.
Ravindranath V, Chandrasekhara N. Metabolism of curcumn-studies with [3H]curcumin. Toxicology. 1981;22(4):337–44.
Wahlström B, Blennow G. A study on the fate of curcumin in the rat. Acta Pharmacol Toxicol (Copenh). 1978;43(2):86–92.
Zhang YN, Poon W, Tavares AJ, McGilvray ID, Chan WCW. Nanoparticle–liver interactions: cellular uptake and hepatobiliary elimination. J Control Release. 2016;240:332–48.
Zylberberg C, Gaskill K, Pasley S, Matosevic S. Engineering liposomal nanoparticles for targeted gene therapy. Gene Ther. 2017;24(8):441–52.
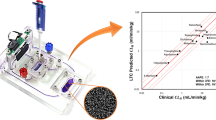
Perazzolo S, Shireman LM, McConnachie LA, Koehn J, Kinman L, Lee W, et al. Integration of computational and experimental approaches to elucidate mechanisms of first-pass lymphatic drug sequestration and long-acting pharmacokinetics of the injectable triple-HIV drug combination TLC-ART 101. J Pharm Sci. 2020;109(5):1789–801.
Leung MHM, Kee TW. Effective stabilization of curcumin by association to plasma proteins: human serum albumin and fibrinogen. Langmuir. 2009;25(10):5773–7.
Acknowledgements
This work was in part supported by summer internship program of Genentech Inc. The authors would also like to thank Genentech Small Molecule Pharmaceutics department for supporting this work.
Funding
This research was funded by Genentech Inc.
Author information
Authors and Affiliations
Contributions
ESL, CY, and DL designed the research. JY and JM designed and analyzed the PK studies. LL and ET performed the PK studies. AE performed and analyzed the imaging. ESL, CY, and JY wrote the manuscript. JY and DL revised the manuscript. All authors edited the final manuscript version.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Levy, E.S., Yu, J., Estevez, A. et al. A Systematic Approach for Liposome and Lipodisk Preclinical Formulation Development by Microfluidic Technology. AAPS J 23, 111 (2021). https://doi.org/10.1208/s12248-021-00651-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-021-00651-4