Abstract
Heme oxygenase (HO), the rate-limiting enzyme in heme degradation, plays a role in neonatal jaundice. Understanding the regulation of the developmental expression patterns of the two HO isozymes, HO-1 and HO-2, is essential for targeting HO to control pathologic jaundice, and uncovering the fundamental role that they play in mammalian development. Here we characterized the ontogeny of HO-1 and HO-2 expression in the developing mouse cortex by in vivo bioluminescence imaging, quantitative RT-PCR, and Western blot. HO-2, the predominant isoform in the adult cortex, was relatively stable throughout all ages. HO-1 was observed to be progressively down-regulated in an age-related manner. HO-1 expression in the adult cortex was also the lowest among the eight adult tissues analyzed. Because there is a 283-bp CpG island region in the HO-1 promoter, we hypothesized that methylation of the island is responsible for the age-related HO-1 down-regulation in the cortex. Methylation status was assessed using regular and quantitative methylation-specific PCR and the CpG island was found to be hypomethylated at all ages. Therefore, we conclude that HO-1 gene expression in the cortex is developmentally-regulated and that methylation of the HO-1 CpG island is not associated with the down-regulation of the gene.
Similar content being viewed by others
Main
Heme oxygenase (HO) is the rate-limiting enzyme for the degradation of heme derived from senescent red blood cells and other hemoproteins (1). In this pathway, HO cleaves the heme ring via oxidation at the α-methene bridge to yield equimolar quantities of biliverdin, carbon monoxide (CO), and free iron. Biliverdin is subsequently converted to bilirubin by biliverdin reductase. Among the many implications of HO in development, it is the controlling step in bilirubin production and has been identified as a target for prevention of pathologic jaundice.
HO has many physiologic functions (2,3) and plays a key role during development. It not only regulates cellular heme and hemoprotein levels (e.g. cytochrome P450), but also all of its heme degradation products are biologically active: the bile pigments biliverdin and bilirubin are potent antioxidants (4); CO generates cGMP and MAP kinase, and is shown to be a potential vasodilator and smooth muscle relaxant (5); and free iron regulates expression of various genes and hematopoiesis (6,7). To date, two primary isoforms of HO have been identified in the human and rodent: the inducible HO-1, also known as Hsp32, and the constitutive HO-2 (8,9). HO-3, a third isoform cloned only from rats, has been reported to be a processed pseudogene derived from HO-2 transcripts (10). HO-1 and HO-2 have some protein similarities, but are encoded from different genes.
Expression of HO-1 is transcriptionally regulated by a 15-kb regulatory region, consisting of a basal promoter and three enhancer regions (11). Several regulatory elements and transcriptional factor binding sites have been reported to play important roles in up-regulating HO-1, including metal response elements (MREs), stress response elements (StREs), AP-1, and NF-κB, etc. (12). However, only one transcription factor, Bach1, has been demonstrated to have a repressor effect on HO-1 expression (13). Modulation of HO-2 expression has only been observed in response to glucocorticoids (14). Recently, HO-2 has been demonstrated to be part of a calcium-sensitive potassium channel complex and functions as an oxygen sensor (15).
DNA methylation is the major epigenetic modification that can profoundly affect tissue-specific gene expression, X-chromosome inactivation, genomic imprinting, immobilization of mammalian transposons, and suppression of transcriptional noise (16,17). In mammals, this modification occurs mainly on cytosines in CpG dinucleotides located in CpG-rich DNA fragments, the so-called “CpG islands.” These islands primarily reside at the transcription start site of housekeeping genes and are also associated with tissue-specific genes. In general, it is believed that global demethylation occurs early in mammalian embryos after fertilization and until the blastocyst stage. DNA methylation is then restored quickly after implantation and maintained thereafter in somatic lineages (16). These tissue-dependent and differentially-methylated regions (T-DMR) are widely distributed and abundant in the normal mouse genome and believed to be involved in the establishment of cell- and tissue-specific gene expression (18,19). However, accumulating evidence refute the idea of the very restricted window of time when methylation occurs, showing that either methylation or demethylation occurs at many ages.
In this study, we first characterized the developmental pattern of HO-1 expression in the cortex of a luciferase reporter mouse (HO-1-luc) using in vivo bioluminescence imaging (BLI). We then systematically assessed the expression profiles of HO-1 and HO-2 in the developing cortex by determining mRNA and protein. We further focused on the mechanism of HO-1 down-regulation since HO-2 expression was found relatively stable throughout ages, suggesting that HO-2 was constitutively expressed. After identifying a CpG island region in the HO-1 (but not in the HO-2) promoter, we hypothesized that the methylation status of the CpG dinucleotides in this region was involved in the postnatal down-regulation of HO-1 expression in the mouse cortex. We also explored the role of Bach-1 in the regulation of HO-1 during cortical development.
MATERIALS AND METHODS
Animals.
The study was approved by the Institutional Animal Care and Use Committee at Stanford University. All transgenic HO-1-luc mice, with the transgene containing a 15-kb HO-1 promoter driving expression of the reporter gene luciferase, were obtained from the Stanford Transgenic Animal Facility (Stanford, CA). FVB mice were obtained from Charles River Laboratories (Wilmington, MA).
Reagents.
Luciferin (150 mg/mL, BioSynth, Naperville, IL) was dissolved in water and filter-sterilized. Nembutal (50 mg/mL) was diluted 1:9 (5 mg/mL) for adults or 1:19 (2.5 mg/mL) for newborns up to 10g, with normal saline.
In vivo bioluminescence imaging (BLI).
Adult mice were anesthetized using Nembutal (25–70 mg/kg BW) and imaged as previously described (20). Because mice <14d are virtually immobile, anesthesia was not required. Each newborn was placed into a well of a six-well cell culture plate. All animals were imaged using the In Vivo Imaging System (IVISTM, Xenogen Corp, Alameda, CA) as previously described (21). Total photon emission from the head region (representing the brain) was quantitated and plotted as fold increase from adult (35d) levels.
Tissue preparation.
Brains were immediately removed from animals, cortex isolated and then divided as follows: 1) Whole portions were placed in RNAlater (Qiagen, Germany) and stored at –20°C for total RNA; and 2) 100 ± 2 mg tissue were diced and sonicated in nine volumes of buffer (22) and stored at –80°C with protease inhibitors (Sigma Chemical Co., St. Louis, MO) for Western blots. Protein concentrations were determined using the Bio-Rad Protein Assay kit (Bio-Rad, Hercules, CA).
RT-PCR.
Total RNA was extracted using the RNAeasy Mini Kit (Qiagen) following the manufacturer's instructions. HO-1, HO-2, and Bach1 mRNA levels were quantified using Quanti-Tect SYBR Green RT-PCR kit (Qiagen), performed in an iCycler iQ Real-Time Detection System (Bio-Rad) using primers shown in Table 1. Standard mRNAs for HO-1, HO-2, and Bach1 were synthesized from pRC-CMV-HO1 and pRC-CMV-HO2 using the Riboprobe In Vitro Transcription System (Promega Corp., Madison, WI). The concentration (g/μL) of the resulted standard RNA was measured by OD260 and the copy number was calculated by following formula:

Copy numbers of HO-1, HO-2, and Bach1 mRNA for each sample was calculated from the standard curve and then normalized to β-actin levels (Figs. 2 and 6).
mRNA and protein levels of HO-1 and HO-2 in the developing mouse cortex. (A) HO-1 (○) and HO-2 (□) mRNA levels (mean ± SD) in the cortex from mice at ages E14, E18, newborn, 2 wks, and 6 wks (n = 3 for each age) were determined by real-time RT-PCR, expressed as copies/μg total RNA, and normalized to β-actin. (B) Western blots of HO-1 protein levels in the developing mouse cortex. Each bar graph represents band intensities compared with 35d levels. *p < 0.0001 vs. all ages; † p < 0.02 vs. 6 wk.
Expression pattern of Bach1 in the developing mouse cortex. Bach1 mRNA levels were determined by real-time RT-PCR in cortex of mice at ages E14, E18, neonate, 2 wks, and 6 wks (n = 3 for each age). Data were calculated as copies/ng total RNA, normalized to β-actin, and shown as fold induction (mean± SD) from adult levels. *p ≤ 0.05.
Western blot.
100 μg of the sonicates was boiled for 10 min in protein loading buffer and electrophoresed. Proteins were transferred to a PDVF membrane (Bio-Rad) using a semidry transblotter. The membrane was probed with HO-1 and HO-2 polyclonal antibodies (1:1000, Stressgen Corp., Victoria, Canada) and protein levels quantitated by densitometry as previously described (21). Coomassie blue staining and/or β-actin immunoblotting were performed to confirm equal loading of samples.
Identification of CpG island.
A CpG island was identified using CpG Plot program (http://www.ebi.ac.uk/emboss/cpgplot/) designed by EMBL-EBI (European Bioinformatics Institute). DNA sequences (≤ 5kb) were screened each time and plotted with the regions as defined CpG islands by the program.
In vitro methylation of the HO-1 BAC clone.
BAC clone (rp21-411019) containing the HO-1 gene was created from the RPCI mouse PAC library 21 (Children's Hospital Oakland Research Institute, CHORI, Oakland, CA) and its sequence created from the GeneBank AC005290. The BAC DNA was tested to be unmethylated. To synthesize a positive control DNA for the methylated CpG island, BAC DNA was treated with SssI methylase (New England BioLab, Beverly, MA) and the completion of CpG methylation was confirmed by digestion with BstUI.
Sodium bisulfite treatment of DNA and methylation-specific PCR (MSP).
Genomic DNA was isolated, treated with sodium bisulfite and followed by methylation-specific PCR (MSP) and sequencing, as described (http://www3.mdanderson.org/leukemia/methylation). In brief, genomic DNA was isolated using the DNeasy Tissue Kit (Qiagen) and 2-μg were treated with sodium bisulfite using the EZ DNA Methylation kit (ZYMO Research Corp., Orange, CA). Two μL of resulting DNA was used as a template for the nested MSP reaction. The first PCR step was: 95°C 15min; 94°C 30s, 53°C 30s, and 72°C 30s, 35 cycles; 72°C 5min. Program used for the second PCR reaction was: 95°C 15min; 94°C 30s, 67°C 30s, and 72°C 45s, 35 cycles; 72°C 5min. The products were then separated and analyzed by electrophoresis on 1.8% agarose gels.
Quantitative methylation-specific PCR (Q-MSP).
The percentage of methylated to unmethylated DNA was determined by Q-MSP using the QuantiTect SYBR Green Real-time PCR Kit (Qiagen). A BAC plasmid, treated or not treated with SssI methylase, was used to generate a standard curve for methylated or unmethylated PCR, respectively. Dilution of the first PCR product was amplified in real-time using MSP primer sets (Table 1). PCR was run as follows: 95°C 15min; 94°C 15s, 60.5°C 30s, and 72°C 30s, 40 cycles.
Calculations.
All data are presented as mean ± SD. ANOVA analysis was performed to determine significant differences as defined as p < 0.05.
RESULTS
In vivo HO-1 transcription in the developing cortex.
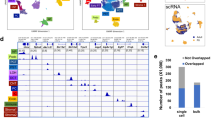
We have observed using in vivo BLI that newborn mice have high HO-1 transcriptional activity in contrast to that reported for adult mice. To characterize the expression pattern, we assessed brain HO-1 transcriptional activity in HO-1-luc mice ranging from neonate to adult using BLI (Fig. 1). HO-1 transcription was highest (20-fold) in 1d-old mice, and then progressively decreased to an average of 4-fold during 2–7d of age, and finally falling to adult levels by 14d of age. Although high individual variations of light emission were found, the downward trend in HO-1 transcription was preserved among all mice imaged.
Transcriptional pattern of HO-1 expression in the developing mouse cortex. In vivo HO-1 transcription levels in the developing mouse cortex were assessed by BLI in mice aged 1–35d of age (n = 3 for each age). Representative whole body images are shown in the same scale (rainbow). Photon emission was quantitated from the head region and normalized to day 35 levels. *p ≤ 0.001.
Down-regulation of HO-1 in the developing cortex.
To investigate the mechanism of the developmental pattern of HO-1 transcription, we analyzed the transcriptional profiles of HO-1 and HO-2 mRNA in wild-type FVB mice, at embryonic ages E14 and E18, 1d (neonate), 2 wks, and 6 wks (Fig. 2A). HO-1 mRNA levels declined with age with the highest expression at E14 (7-fold, relative to 6-wk-old levels, p < 0.0001), followed by a gradual decline during the perinatal period (3-fold, p < 0.02), and reaching the lowest levels at adulthood. In contrast, HO-2 mRNA levels remained relatively stable throughout all ages as expected and was the predominant isoform at all ages (p < 0.005, HO-1 versus HO-2 for each age).
Down-regulation of cortical HO-1 protein was confirmed by Western blot (Fig. 2B). Compared with adult levels, HO-1 protein was the highest (10-fold) at 1d of age, followed by a gradual decrease, reaching lowest levels by day 28. Meanwhile, HO-2 protein levels were fairly constant over the age range studied (data not shown).
Expression of brain HO-1 and HO-2 among adult tissues.
To investigate whether the HO-1 expression profile during brain development was unique to the developing cortex, we assessed HO-1 expression in adult spleen, bone marrow, kidney, liver, heart, testis, lung, and cortex. HO-1 was expressed variably among tissues – with the highest expression found in the spleen and bone marrow. Minimal expression was found in the cortex – more than 5-fold less than that of other tissues (p < 0.05), more specifically, 36-fold less than spleen (p < 0.0001) and 23.5-fold less than bone marrow (p < 0.0001, Fig. 3A). HO-2 mRNA levels in all tissues were relatively stable, except in the testis, which had 5- to 10-fold higher levels than that of other tissues (p < 0.0001).
Tissue-specific expression HO-1 and HO-2 mRNA and protein levels in adult mice. HO-1 (left) and HO-2 (right) mRNA (A) and protein (B) levels from adult spleen (▪), bone marrow ( ), kidney (
), kidney ( ), liver (
), liver ( ), heart (
), heart ( ), testis (
), testis ( ), lung (
), lung ( ), and cortex (
), and cortex ( ) were determined by real-time RT-PCR and Western blot. *p ≤ 0.02, † p < 0.0001 vs. all tissues; **p = 0.025 vs. liver. Each bar graph represents the mean value of n = 3 Western blots.
) were determined by real-time RT-PCR and Western blot. *p ≤ 0.02, † p < 0.0001 vs. all tissues; **p = 0.025 vs. liver. Each bar graph represents the mean value of n = 3 Western blots.
When HO-1 protein levels were determined (Fig. 3B), no protein was detected in the cortex. Spleen showed 2–3-fold more HO-1 protein than that of liver, kidney, and testis, and 15-fold more than that of lung. The bands from heart samples were slightly up-shifted, suggesting some post-translational modifications of HO-1 protein in heart. HO-2 protein has a different pattern, with the highest levels in adult testis followed by adult brain (Fig. 3B).
When the relative ratio of HO-1 to HO-2 mRNA levels in each tissues were compared (Table 2), HO-1 mRNA levels were higher in the spleen (3.3-fold, p < 0.05) and bone marrow (1.6-fold, not significant) only. In contrast, HO-2 was the predominant isozyme in the cortex (13.2-fold, p < 0.01), testis (10.7-fold, p < 0.005), liver (1.9-fold, p < 0.05), and lung (1.5-fold, p < 0.05).
Identification of a CpG island in HO-1 promoter.
A 15-kb upstream region of HO-1 coding sequence has been shown to play a critical role in the transcriptional regulation of HO-1 (11). We identified a CpG-rich cluster of 283-bp in the promoter and transcriptional start sites consisting of 25 CpG dinucleotides located between 110 bp upstream and 172 bp downstream of the transcriptional start site (Fig. 4). This DNA sequence possesses all the properties of a target for methylation, which includes a composition of C and G of over 50% and an observed/expected ratio of over 0.6 for a minimum of 200 bases. The region contains several putative transcriptional factor binding sites, including Sp-1, C/EBP, AP-4 MYC/MAX, TATA, and AP-2. Thirteen CpG dinucleotides are located downstream of the transcriptional start site and among them, three in Exon 1.
CpG island in the HO-1 promoter region. A 283-bp region consisting of 25 CpG dinucleotides was identified in the HO-1 promoter, located between 110-bp upstream and 172-bp downstream from the transcriptional start site. DE1 and DE2, distal enhancers-1 and -2, respectively; PE, proximal enhancer; and P, promoter.
When we screened for the putative CpG rich cluster in the upstream region of HO-2 coding sequence, no CpG islands were found.
Hypomethylation of the HO-1 CpG island.
To study the possible involvement of DNA methylation in the down-regulation of HO-1 expression, MSP was performed using two sets of PCR primers, each of which can only recognize either methylated or unmethylated DNA (Fig. 5A). Figure 5B shows the presence of only unmethylated bands and the absence of methylated bands in all tissues, which suggests that the HO-1 CpG island is hypomethylated and preserved throughout all tissues even in the developing cortex. After performing MSP sequencing, we found that all of 25 CpG dinucleotides were uniformly unmethylated (data not shown).
Methylation-specific PCR for assessing methylation patterns of HO-1. (A) Primer design for MSP-PCR and methylation-specific sequencing. Each filled circle represents a CpG dinucleotide and its position in the island. The position and direction of primers for the nested PCR are shown with arrows; (B) Methylation status in the (top panel) developing cortex: E14 and E18 (n = 6), neonate (n = 3), 2 wk- and 6-wk-old mice (n = 3), and in the spleen, bone marrow, liver, kidney, testis, lung, and cortex of adult mice (6-wk-old, n = 3, middle panel) as assessed by MSP-PCR. Bottom panels show validation of the method using control DNA: HO-1 PAC plasmid DNA (unmethylated DNA, U) or the plasmid DNA treated with SssI methylase (methylated DNA, M).
Because every tissue is a mixed population of different cell types, it is virtually impossible to detect and quantify the small percentage of the methylated DNA in the pool of mostly unmethylated DNA using the traditional MSP method. Therefore, we adapted a method called Q-MSP using real-time PCR (23,24), a method that is very sensitive, capable of detecting 1 molecule of methylated DNA in 104 of unmethylated DNA. Using this methodology, we found that most samples consisted of completely unmethylated DNA. Only embryonic samples (E14 and E18 cortex, and E18 liver) contained a low percentage of methylated DNA, ranging from only 0.1–15% (Table 3). This low frequency of DNA methylation was not found in any of the adult tissues studied.
Bach1 expression in the developing mouse cortex.
To explore another possible mechanism for the observed down-regulation of HO-1 in the adult cortex, we investigated the role of Bach1, the only identified repressor of HO-1 transcription (25). We hypothesized that expression of Bach1 levels should increase with increasing age. When Bach1 mRNA levels in the developing cortex were measured using real-time RT-PCR, we found that Bach1 expression was highest at age E14 and decreased progressively during development (Fig. 6).
DISCUSSION
Characterization of the developmental expression profiles of HO isozymes is fundamental for the understanding of their physiology functions and roles in age-related diseases. In this study, we identified age-related expression patterns of HO isozymes in the mouse cortex. HO-1 expression is high in early fetal ages (E10 and 14), progressively decreases in perinatal period, and then reaches lowest levels by adulthood; while HO-2, the predominant isoform in brain, remains relatively stable throughout development (Fig. 2). This ontogeny of the mouse cortex is unique and distinct from the expression profiles of other tissues, such as liver (26) and spleen. Although HO-1 in the neonatal cortex has a similar mRNA expression level as the neonatal liver (data not shown), its adult level is significantly less (87%) than that of the adult liver. Moreover, cortical HO-1 is the lowest among all other adult tissues tested suggesting that HO-1 expression is highly suppressed during cortical maturation. Our data are consistent with the findings of Bergeron et al. (27), which showed the minimal HO-1 staining in adult rat brain endothelium, white matter, and cortex by immunocytochemistry.
We speculated that suppression of HO-1 transcription in the adult cortex might be controlled in a cell type- and region-specific manner. Others have shown that in brains exposed to stresses, such as hyperosmotic insult (28), heme administration (29), and ischemic injury (30), HO-1 expression is induced only in astrocytes and microglia. HO-1 silencing phenomenon was also reported in the study by Chauveau et al. (31), in which HO-1 expression drastically decreased during human and rat dendritic cell maturation induced in vitro as well as in human tissue. They also found that prevention of HO-1 silencing could inhibit LPS-induced dendritic cell phenotypic maturation and secretion of pro-inflammatory cytokines, suggesting the necessity of HO-1 silencing in some cell types for their function.
There are several mechanisms for gene silencing, one of which is DNA methylation of CpG islands. Our data shows that hypomethylation of the HO-1 CpG island in all tissues tested, suggesting that methylation does not occur, even in the less populated cells in the adult cortex. In addition, by sequencing this region, no mCpG dinucleotides in the CpG island (data not shown) were found, indicating that there is no T-DMRs in HO-1 promoter. Therefore, we conclude that CpG island methylation is not associated with HO-1 down-regulation, and other mechanisms must be involved. To further investigate other mechanisms, which may regulate this down-regulation, we measured the expression of Bach1, the only identified transcriptional repressor of HO-1 (25). We found that Bach1 mRNA levels actually paralleled that of HO-1 and therefore does not appear to silence HO-1 expression in the adult cortex (Fig. 6). We speculate that reduction of transcriptional activators, such as Nrf-2, might be involved in the down-regulation. Other possibilities might be that there is a limited amount of heme in adult neuronal cells or induction of multi-drug resistant protein (MDR) during cortical maturation, which functions as a filter to block harmful agents from entering cells.
In neonatal mice, an increase in HO-1 expression may be the result of an abundance of heme, both as a substrate and inducer of HO-1, released from senescing fetal erythrocytes, which have a shortened life span during neonatal period. In addition, Bergeron et al. (27) has shown that HO-1 expression was cell-type specific in the brain, which suggests that HO-1 up-regulation may be due to the differentiation, proliferation and redistribution of neuronal cells in the developing mouse cortex. Moreover, the inducibility of HO-1 can also be influenced by endocrine factors (26).
We have observed co-expression of both isozymes in all the adult tissues collected, but their relative levels were quite different. HO-2 expression exceeded that of HO-1 in most organs, especially in the cortex (at all ages) and testis; whereas, in the spleen and bone marrow, HO-1 was present more abundantly than HO-2 (Table 2). Besides their function in maintaining cellular homeostasis, each isozyme has its unique tissue-specific role. In the cortex, the two isozymes display distinct regional and cellular distributions with HO-1 present only in a few cell subtypes and HO-2 being very widespread (32–35). The inducible HO-1 plays a mostly protective role in the CNS, while the constitutively-expressed HO-2 is important for maintaining neuronal function, probably by producing CO, a putative neurotransmitter (36,37). High expression levels of HO-2 in the testis may be associated with rodent male reproduction (38). HO-1 is expressed abundantly in spleen and bone marrow, where senescent red blood cells are sequestered and heme degraded. Moreover, HO-1 and its byproducts have anti-inflammatory and anti-apoptotic properties, all which function in immune cell maturation and hematopoiesis.
Although HO-1 and HO-2 have been extensively studied, the physiologic relevance of HO-1 and HO-2 and their reciprocal interrelationship is not quite fully understood. Our data suggested that these two isozymes are regulated independently in developing and adult tissues with unique physiologic roles. Yet, their combined activities control tissue heme catabolism. Therefore proper balance of the expression of both isozymes is key to maintaining tissue integrity and homeostasis.
Abbreviations
- BLI:
-
bioluminescence imaging
- HO:
-
heme oxygenase
- MSP:
-
methylation-specific PCR
- Q-MSP:
-
quantitative methylation-specific PCR
References
Tenhunen R, Marver HS, Schmid R 1968 The enzymatic conversion of heme to bilirubin by microsomal heme oxygenase. Proc Natl Acad Sci USA 61: 748–755
Abraham NG, Lin JH, Schwartzman ML, Levere RD, Shibahara S 1988 The physiological significance of heme oxygenase. Int J Biochem 20: 543–558
Elbirt KK, Bonkovsky HL 1999 Heme oxygenase: recent advances in understanding its regulation and role. Proc Assoc Am Physicians 111: 438–447
Maines MD 2003 Bile pigments: newcomers to the cell signaling arena. Toxicol Sci 71: 9–10
Christova T, Diankova Z, Setchenska M 2000 Heme oxygenase – carbon monoxide signalling pathway as a physiological regulator of vascular smooth muscle cells. Acta Physiol Pharmacol Bulg 25: 9–17
Abraham NG, Lutton JD, Levere RD 1985 Heme metabolism and erythropoiesis in abnormal iron states: role of delta-aminolevulinic acid synthase and heme oxygenase. Exp Hematol 13: 838–843
Brown A, Lutton JD, Nelson J, Abraham NG, Levere RD 1987 Microenvironmental cytokines and expression of erythroid heme metabolic enzymes. Blood Cells 13: 123–136
Cruse I, Maines MD 1988 Evidence suggesting that the two forms of heme oxygenase are products of different genes. J Biol Chem 263: 3348–3353
Shibahara S, Yoshizawa M, Suzuki H, Takeda K, Meguro K, Endo K 1993 Functional analysis of cDNAs for two types of human heme oxygenase and evidence for their separate regulation. J Biochem (Tokyo) 113: 214–218
Hayashi S, Omata Y, Sakamoto H, Higashimoto Y, Hara T, Sagara Y, Noguchi M 2004 Characterization of rat heme oxygenase-3 gene. Implication of processed pseudogenes derived from heme oxygenase-2 gene. Gene 336: 241–250
Alam J, Camhi S, Choi AM 1995 Identification of a second region upstream of the mouse heme oxygenase-1 gene that functions as a basal level and inducer-dependent transcription enhancer. J Biol Chem 270: 11977–11984
Choi AM, Alam J 1996 Heme oxygenase-1: function, regulation, and implication of a novel stress-inducible protein in oxidant-induced lung injury. Am J Respir Cell Mol Biol 15: 9–19
Kitamuro T, Takahashi K, Ogawa K, Udono-Fujimori R, Takeda K, Furuyama K, Nakayama M, Sun J, Fujita H, Hida W, Hattori T, Shirato K, Igarashi K, Shibahara S 2003 Bach1 functions as a hypoxia-inducible repressor for the heme oxygenase-1 gene in human cells. J Biol Chem 278: 9125–9133
Raju VS, McCoubrey WK, Maines MD 1997 Regulation of heme oxygenase-2 by glucocorticoids in neonatal rat brain: characterization of a functional glucocorticoid response element. Biochim Biophys Acta 1351: 89–104
Williams SE, Wootton P, Mason HS, Bould J, Iles DE, Riccardi D, Peers C, Kemp PJ 2004 Hemoxygenase-2 is an oxygen sensor for a calcium-sensitive potassium channel. Science 306: 2093–2097
Bird A 2002 DNA methylation patterns and epigenetic memory. Genes Dev 16: 6–21
Jones PA, Takai D 2001 The role of DNA methylation in mammalian epigenetics. Science 293: 1068–1070
Imamura T, Ohgane J, Ito S, Ogawa T, Hattori N, Tanaka S, Shiota K 2001 CpG island of rat sphingosine kinase-1 gene: tissue-dependent DNA methylation status and multiple alternative first exons. Genomics 76: 117–125
Shiota K 2004 DNA methylation profiles of CpG islands for cellular differentiation and development in mammals. Cytogenet Genome Res 105: 325–334
Zhang W, Feng JQ, Harris SE, Contag PR, Stevenson DK, Contag CH 2001 Rapid in vivo functional analysis of transgenes in mice using whole body imaging of luciferase expression. Transgenic Res 10: 423–434
Morioka I, Wong RJ, Abate A, Vreman HJ, Contag CH, Stevenson DK 2006 The systemic effects of orally administered zinc and tin (IV) metalloporphyrins on heme oxygenase expression in mice. Pediatr Res 59: 667–672
Vreman HJ, Stevenson DK 1999 Detection of heme oxygenase activity by measurement of CO, In: Maines MD, Costa LG, Reed DJ, Sassa S, Sipes IG (eds) Current Protocols in Toxicology. John Wiley & Sons, Inc, New York, pp 9.2.1–9.2.10
Chan MW, Chu ES, To KF, Leung WK 2004 Quantitative detection of methylated SOCS-1, a tumor suppressor gene, by a modified protocol of quantitative real time methylation-specific PCR using SYBR green and its use in early gastric cancer detection. Biotechnol Lett 26: 1289–1293
Thomassin H, Kress C, Grange T 2004 MethylQuant: a sensitive method for quantifying methylation of specific cytosines within the genome. Nucleic Acids Res 32: e168–176
Kanezaki R, Toki T, Yokoyama M, Yomogida K, Sugiyama K, Yamamoto M, Igarashi K, Ito E 2001 Transcription factor BACH1 is recruited to the nucleus by its novel alternative spliced isoform. J Biol Chem 276: 7278–7284
Lin JH, Villalon P, Nelson JC, Abraham NG 1989 Expression of rat liver heme oxygenase gene during development. Arch Biochem Biophys 270: 623–629
Bergeron M, Ferriero DM, Sharp FR 1998 Developmental expression of heme oxygenase-1 (HSP32) in rat brain: an immunocytochemical study. Brain Res Dev Brain Res 105: 181–194
Turner CP, Bergeron M, Matz P, Zegna A, Noble LJ, Panter SS, Sharp FR 1998 Heme oxygenase-1 is induced in glia throughout brain by subarachnoid hemoglobin. J Cereb Blood Flow Metab 18: 257–273
Richmon JD, Fukuda K, Maida N, Sato M, Bergeron M, Sharp FR, Panter SS, Noble LJ 1998 Induction of heme oxygenase-1 after hyperosmotic opening of the blood-brain barrier. Brain Res 780: 108–118
Nimura T, Weinstein PR, Massa SM, Panter S, Sharp FR 1996 Heme oxygenase-1 (HO-1) protein induction in rat brain following focal ischemia. Brain Res Mol Brain Res 37: 201–208
Chauveau C, Remy S, Royer PJ, Hill M, Tanguy-Royer S, Hubert FX, Tesson L, Brion R, Beriou G, Gregoire M, Josien R, Cuturi MC, Anegon I 2005 Heme oxygenase-1 expression inhibits dendritic cell maturation and proinflammatory function but conserves IL-10 expression. Blood 106: 1694–1702
Sun Y, Rotenberg MO, Maines MD 1990 Developmental expression of heme oxygenase isozymes in rat brain. Two HO-2 mRNAs are detected. J Biol Chem 265: 8212–8217
Ewing JF, Maines MD 1995 Distribution of constitutive (HO-2) and heat-inducible (HO-1) heme oxygenase isozymes in rat testes: HO-2 displays stage-specific expression in germ cells. Endocrinology 136: 2294–2302
Scapagnini G, D'Agata V, Calabrese V, Pascale A, Colombrita C, Alkon D, Cavallaro S 2002 Gene expression profiles of heme oxygenase isoforms in the rat brain. Brain Res 954: 51–59
Ewing JF, Maines MD 1997 Histochemical localization of heme oxygenase-2 protein and mRNA expression in rat brain. Brain Res Brain Res Protoc 1: 165–174
Snyder SH, Jaffrey SR, Zakhary R 1998 Nitric oxide and carbon monoxide: parallel roles as neural messengers. Brain Res Brain Res Rev 26: 167–175
Maines MD 2000 The heme oxygenase system and its functions in the brain. Cell Mol Biol 46: 573–585
McCoubrey WK, Eke B, Maines MD 1995 Multiple transcripts encoding heme oxygenase-2 in rat testis: developmental and cell-specific regulation of transcripts and protein. Biol Reprod 53: 1330–1338
Acknowledgements
We thank Drs. Stacy Burns-Guydish and Ichiro Morioka for their help with tissue sample collection. We also thank Drs. Weisheng Zhang and Aida Abate for their critical reviews of the manuscript. We are indebted to Dr. Kazuhiko Igarashi for providing us with the Bach1 antibody.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is supported by the National Institutes of Health grants HL58013 and HL68703 and unrestricted funds from the Hess Research Fund, the Mary L. Johnson Research Fund and Philips Medical Corp.
Rights and permissions
About this article
Cite this article
Zhao, H., Wong, R., Nguyen, X. et al. Expression and Regulation of Heme Oxygenase Isozymes in the Developing Mouse Cortex. Pediatr Res 60, 518–523 (2006). https://doi.org/10.1203/01.PDR.0000242374.21415.f5
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1203/01.PDR.0000242374.21415.f5
This article is cited by
-
Deletion of the hemopexin or heme oxygenase-2 gene aggravates brain injury following stroma-free hemoglobin-induced intracerebral hemorrhage
Journal of Neuroinflammation (2016)
-
Optical imaging in vivo with a focus on paediatric disease: technical progress, current preclinical and clinical applications and future perspectives
Pediatric Radiology (2011)