Abstract
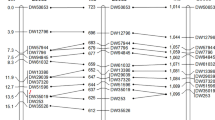
The genetic map of chromosome 5B has been constructed by using microsatellite (SSR) analysis of 381 plants from the F2 population produced by cross of the Chinese Spring (CS) and Renan cultivars. Initially, 180 SSR markers for the common wheat 5B chromosome have been used for analysis of these cultivars. The 32 markers able to detect polymorphism between these cultivars have been located on the genetic map of chromosome 5B. Cytogenetic mapping has involved a set of CS 5B chromosome deletion lines. Totally, 51 SSR markers have been located in ten regions (deletion bins) of this chromosome by SSR analysis of these deletion lines. Five genes—TaCBFIIIc-B10, Vrn-B1, Chi-B1, Skr, and Ph1—have been integrated into the cytogenetic map of chromosome 5B using the markers either specific of or tightly linked to the genes in question. Comparison of the genetic and cytogenetic maps suggests that recombination is suppressed in the pericentromeric region of chromosome 5B, especially in the short arm segment. The 18 markers localized to deletion bins 5BL16-0.79-1.00 and 5BL18-0.66-0.79 have been used to analyze common wheat introgression lines L842, L5366-180, L73/00i, and L21-4, carrying fragments of alien genomes in the terminal region of 5B long arm. L5366-180 and L842 lines carry a fragment of the Triticum timopheevii 5GL chromosome, while L73/00i and L21-4 lines, a fragment of the Aegilops speltoides 5SL chromosome. As has been shown, the translocated fragments in these four lines are of different lengths, allowing bin 5BL18-0.66-0.79 to be divided into three shorter regions. The utility of wheat introgression lines carrying alien translocations for increasing the resolution of cytogenetic mapping is discussed.
Similar content being viewed by others
References
Sears, E.R., Nullisomic-tetrasomic combinations in hexaploid wheat, in Chromosome Manipulations and Plant Genetics, London: Oliver and Boyd, 1966, pp. 29–45.
Lazo, G.R., Chao, S., Hummel, D.L., et al., Development of an expressed sequence tag resource for wheat (Triticum aestivum L.): unigene analysis, probe selection and bioinformatics for a 16.000-locus bin map, Genetics, 2004, vol. 168, pp. 585–593.
Paux, E., Sourdille, P., Salse, J., et al., Physical map of the 1-Gigabase bread wheat chromosome 3B, Science, 2008, vol. 322, pp. 101–104.
Endo, T.R. and Gill, B.S., The deletion stocks of common wheat, J. Hered., 1996, vol. 87, no. 4, pp. 295–307.
Sourdille, P., Singh, S., Cadalen, T., et al., Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.), Funct. Integr. Genomics, 2004, vol. 4, pp. 12–25.
Qi, L., Echalier, B., Friebe, B., and Gill, B.S., Molecular characterization of a set of wheat deletion stocks for use in chromosome bin mapping of ESTs, Funct. Integr. Genomics, 2003, vol. 3, pp. 39–55.
Kalavacharla, V., Hossain, K., Gu, Y., et al., High-resolution radiation hybrid map of wheat chromosome 1D, Genetics, 2006, vol. 173, pp. 1089–1099.
Schlegel, R., Current list of wheats with rye and alien introgression, 2013, vol. 01–13, pp. 1–18. http://www.rye-gene-map.de/rye-introgression
Sakata, M., Nasuda, S., and Endo, T.R., Dissection of barley chromosome 4H in common wheat by the gametocidal system and cytological mapping of chromosome 4H with EST markers, Genes Genet. Syst., 2010, vol. 85, pp. 19–29.
Adonina, I.G., Petrash, N.V., Timonova, E.M., et al., Construction and study of leaf rust-resistant common wheat lines with translocations of Aegilops speltoides Tausch. genetic material, Russ. J. Genet., 2012, vol. 48, no. 4, pp. 488–494.
Timonova, E.M., Leonova, I.N., Röber, M.S., and Salina, E.A., Marker-assisted development and characterization of a set of Triticum aestivum lines carrying different introgressions from T. timopheevii genome, Mol. Breed., 2013, vol. 31, no. 1, pp. 123–136.
Alfarez, W., Bouguennec, A., Balfourier, F., et al., Fine mapping and marker development for the crossability gene Skr on chromosome 5BS of hexaploid wheat (Triticum aestivum L.), Genetics, 2009, vol. 183, pp. 469–481.
Griffiths, S., Sharp, R., Foote, T.N., et al., Molecular characterization of Ph1 as a major chromosome pairing locus in polyploid wheat, Nature, 2006, vol. 439, pp. 749–752.
Badawi, M., Danyluk, J., Boucho, B., et al., The CBF gene family in hexaploid wheat and its relationship to the phylogenetic complexity of cereal CBFs, Mol. Genet. Genomics, 2007, vol. 277, pp. 533–554.
Nicot, N., Chiquet, V., Gandon, B., et al., Study of simple sequence repeat (SSR) markers from wheat expressed sequence tags (ESTs), Theor. Appl. Genet., 2004, vol. 109, pp. 800–805.
Lander, E.S., Green, P., Abrahamson, J., et al., Mapmaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations, Genomics, 1987, vol. 1, pp. 174–181.
Kosambi, D.D., The estimation of map distances from recombination values, Ann. Eugen., 1944, vol. 12, no. 1, pp. 172–175.
Gill, K.S., Gill, B.S., Endo, T.R., and Boyko, E.V., Identification and high-density mapping of gene-rich regions in chromosome group 5 of wheat, Genetics, 1996, vol. 143, pp. 1001–1012.
Gadaleta, A., Giancaspro, A., Giove, S.L., et al., Development of a deletion and genetic linkage map for the 5A and 5B chromosomes of wheat (Triticum aestivum), Genome, 2012, vol. 55, pp. 417–427.
Akhunov, E.D., Goodyear, A.W., Geng, S., et al., The organization and rate of evolution of wheat genomes are correlated with recombination rates along chromosome arms, Genome Res., 2003, vol. 13, pp. 753–763.
Erayman, M., Sandhu, D., Sidhu, D., et al., Demarcating the gene-rich regions of the wheat genome, Nucleic Acids Res., 2004, vol. 32, no. 12, pp. 3546–3565.
Saintenac, C., Falque, M., Martin, O.C., et al., Detailed recombination studies along chromosome 3B provide new insights on crossover distribution in wheat (Triticum aestivum L.), Genetics, 2009, vol. 181, pp. 393–403.
Faris, J.D., Haen, K.M., and Gill, B.S., Saturation mapping of a gene-rich recombination hot spot region in wheat, Genetics, 2000, vol. 154, pp. 823–835.
Vrána, J., Kubaláková, M., Simková, H., et al., Flow sorting of mitotic chromosomes in common wheat (Triticum aestivum L.), Genetics, 2000, vol. 156, pp. 2033–2041.
Goyal, A., Bandopadhyay, R., Sourdille, P., et al., Physical molecular maps of wheat chromosomes, Funct. Integr. Genomics, 2005, vol. 5, no. 4, pp. 260–263.
Akhunov, E.D., Akhunova, A.R., Linkiewicz, A.M., et al., Synteny perturbations between wheat homoeologous chromosomes caused by locus duplications and deletions correlate with recombination rates, Proc. Natl. Acad. Sci. U.S.A., 2003, vol. 100, pp. 10836–10841.
Francki, M.G., Walker, E., Crawford, A.C., et al., Comparison of genetic and cytogenetic maps of hexaploid wheat (Triticum aestivum L.) using SSR and DArT markers, Mol. Genet. Genomics, 2009, vol. 281, no. 2, pp. 181–191.
Sarma, R.N., Fish, L., Gill, B.S., and Snape, J.W., Physical characterization of the homoeologous group 5 chromosomes of wheat in terms of rice linkage blocks, and physical mapping of some important genes, Genome, 2000, vol. 43, no. 1, pp. 191–198.
Kynast, R.G., Okagaki, R.J., Galatowitsch, M.W., et al., Dissecting the maize genome by using chromosome addition and radiation hybrid lines, Proc. Natl. Acad. Sci. U.S.A., 2004, vol. 101, pp. 9921–9926.
Gao, W., Chen, Z.J., Yu, J.Z., et al., Widecross whole genome radiation hybrid mapping of cotton (Gossypium hirsutum L.), Genetics, 2004, vol. 167, pp. 1317–1329.
Tsuchida, M., Fukushima, T., Nasuda, S., et al., Dissection of rye chromosome 1R in common wheat, Genes Genet. Syst., 2008, vol. 83, pp. 43–53.
Ashida, T., Nasuda, S., Sato, K., and Endo, T.R., Dissection of barley chromosome 5H in common wheat, Genes Genet. Syst., 2007, vol. 82, pp. 123–133.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © E.M. Timonova, O.B. Dobrovol’skaya, E.M. Sergeeva, L.L. Bildanova, P. Sourdille, C. Feuillet, E.A. Salina, 2013, published in Genetika, 2013, Vol. 49, No. 12, pp. 1376–1384.
The article was translated by the authors.
Rights and permissions
About this article
Cite this article
Timonova, E.M., Dobrovol’skaya, O.B., Sergeeva, E.M. et al. A comparative genetic and cytogenetic mapping of wheat chromosome 5B using introgression lines. Russ J Genet 49, 1200–1206 (2013). https://doi.org/10.1134/S1022795413120132
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795413120132