Abstract
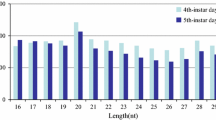
MicroRNAs (miRNAs) are a class of non-protein coding small RNAs that regulate a gene expression at the post-transcriptional level. Using in silico screening, we found that the 3′-untranslated regions of the P25 gene mRNA are perfectly complementary to nucleotides 2–8 at the 5′ end of the miRNA-2b (miR-2b). The expression of miR-2b and the P25 gene in posterior silk gland of the fifth instar larval silkworm was investigated using real-time PCR detection method. The results indicated that expression of the P25 gene was very high in the posterior silk gland during the fifth instar larvae, whereas a level of miR-2b sharply decreased until reaching the lowest one on the 8th day. The expression patterns of miR-2b and P25 gene indicate that miR-2b might act as a fine-tuning regulator of expression of the P25 gene at the post-transcriptional level.
Similar content being viewed by others
References
Ambros V. 2004. The functions of animal microRNAs. Nature. 431, 350–355.
Lee R.C., Feinbaum R.L., Ambros V. 1993. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 75, 843–854.
Brennecke J., Stark A., Russell R.B., Cohen S.M. 2005. Principles of microRNA-target recognition. PLoS Biol. 3, e85.
Huang Y., Zou Q., Wang S.P., Tang S.M., Zhang G.Z., Shen X.J. 2010. The discovery approaches and detection methods of microRNAs. Mol. Biol. Rep. doi: 10.1007/s11033-010-0532-1.
He L., Hannon G.J. 2004. MicroRNAs: small RNAs with a big role in gene regulation. Nature Rev. Genet. 5, 522–531.
Lewis B.P., Burge C.B., Bartel D.P. 2005. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 120, 15–20.
Huang Y., Shen X.J., Zou Q., Wang S.P., Tang S.M., Zhang G.Z. 2010. Biological functions of microRNAs: A review. J. Physiol. Biochem. doi: 10.1007/s11033-010-0532-1.
Couble P., Moine A., Garel A., Prudhomme J.C. 1983. Developmental variations of a nonfibroin mRNA of Bombyx mori silkgland, encoding for a low-molecular-weight silk protein. Dev. Biol. 97, 398–407.
Sprague K.U. 1975. The Bombyx mori silk proteins: characterization of large polypeptides. Biochemistry. 14, 925–931.
Yamaguchi K., Kikuchi Y., Takagi T., Kikuchi A., Oyama F. 1989. Primary structure of the silk fibroin light chain determined by cDNA sequencing and peptide analysis. J. Mol. Biol. 210, 127–139.
Kimura K., Oyama F., Ueda H., Mizuno S., Shimura K. 1985. Molecular cloning of the fibroin light chain complementary DNA and its use in the study of the expression of the light chain gene in the posterior silk gland of Bombyx mori. Experientia. 41, 1167–1171.
Inoue S., Tanaka K., Arisaka F., Kimura S., Ohtomo K. 2000. Silk fibroin of Bombyx mori is secreted, assembling a high molecular mass elementary unit consisting of H-chain, L-chain, and P25, with a 6: 6: 1 molar ratio. J. Biol. Chem. 275, 40517–40528.
Grzelak K. 1995. Control of expression of silk protein genes. Comp. Biochem. Physiol. B: Biochem. Mol. Biol. 110, 671–681.
Okamoto H., Ishikawa E., Suzuki Y. 1982. Structural analysis of sericin genes: Homologies with fibroin gene in the 5′ flanking nucleotide sequences. J. Biol. Chem. 257, 15192–15199.
Cao J., Tong C., Wu X., Lv J., Yang Z. 2008. Identification of conserved microRNAs in Bombyx mori (silk-worm) and regulation of fibroin L chain production by microRNAs in heterologous system. Insect. Biochem. Mol. Biol. 38, 1066–1071.
Huang Y., Zou Q., Song H., Song F., Wang L., Zhang G., Shen X. 2010. A study of miRNAs targets prediction and experimental validation. Protein Cell. 1, 979–986.
Min H., Yoon S. 2010. Got target? Computational methods for microRNA target prediction and their extension. Exp. Mol. Med. 42, 233–244.
Mendes N.D., Freitas A.T., Sagot M.F. 2009. Current tools for the identification of miRNA genes and their targets. Nucleic Acids Res. 37, 2419–2433.
Rehmsmeier M., Steffen P., Hochsmann M., Giegerich R. 2004. Fast and effective prediction of microRNA/target duplexes. RNA. 10, 1507–1517.
Miranda K.C., Huynh T., Tay Y., Ang Y.S., Tam W.L. 2006. A pattern-based method for the identification of microRNA binding sites and their corresponding heteroduplexes. Cell. 126, 1203–1217.
Yu X., Zhou Q., Li S.C., Luo Q., Cai Y. 2008. The silkworm (Bombyx mori) microRNAs and their expressions in multiple developmental stages. PLoS One. 3, e2997.
Schmittgen T.D., Livak K.J. 2008. Analyzing real-time PCR data by the comparative C(T) method. Nature Protoc. 3, 1101–1108.
Mardanova E.S., Zamchuk L.A., Ravin N.V. 2007. The 5′ untranslated region of the maize alcohol dehydrogenase gene provides efficient translation of mRNA in plants under stress conditions. Mol. Biol. (Moscow). 41, 914–919.
Lai E.C., Tomancak P., Williams R.W., Rubin G.M. 2003. Computational identification of Drosophila microRNA genes. Genome Biol. 4, R42.
Lewis B.P., Shih I.H., Jones-Rhoades M.W., Bartel D.P., Burge C.B. 2003. Prediction of mammalian microRNA targets. Cell. 115, 787–798.
Datta A., Ghosh A.K., Kundu S.C. 2001. Differential expression of the fibroin gene in developmental stages of silkworm, Antheraea mylitta (Saturniidae). Comp. Biochem. Physiol. B: Biochem. Mol. Biol. 129, 197–204.
Liu S., Zhang L., Li, Q., Zhao P., Duan J., Cheng D., Xiang Z., Xia Q. 2009. MicroRNA expression profiling during the life cycle of the silkworm (Bombyx mori). BMC Genomics. 10, 455.
Chaudhuri K., Chatterjee R. 2007. MicroRNA detection and target prediction: Integration of computational and experimental approaches. DNA Cell Biol. 26, 321–337.
Li L., Xu J., Yang D., Tan X., Wang H. 2010. Computational approaches for microRNA studies: A review. Mamm. Genome. 21, 1–12.
Sethupathy P., Megraw M., Hatzigeorgiou A.G. 2006. A guide through present computational approaches for the identification of mammalian microRNA targets. Nature Methods. 3, 881–886.
Xia W., Cao G., Shao N. 2009. Progress in miRNA target prediction and identification. Sci. China C Life Sci. 52, 1123–1130.
Vasudevan S., Tong Y., Steitz J.A. 2007. Switching from repression to activation: microRNAs can up-regulate translation. Science. 318, 1931–1934.
Vasudevan S., Steitz J.A. 2007. AU-rich-element-mediated upregulation of translation by FXR1 and argonaute 2. Cell. 128, 1105–1118.
Buchan J.R., Parker R. 2007. The two faces of miRNA. Science. 318, 1877–1878.
Ason B., Darnell D.K., Wittbrodt B., Berezikov E., Kloosterman W.P. 2006. Differences in vertebrate microRNA expression. Proc. Natl. Acad. Sci. U. S. A. 103, 14385–14389.
Aboobaker A.A., Tomancak P., Patel N., Rubin G.M., Lai E.C. 2005. Drosophila microRNAs exhibit diverse spatial expression patterns during embryonic development. Proc. Natl. Acad. Sci. U. S. A. 102, 18017–18022.
Lagos-Quintana M., Rauhut R., Yalcin A., Meyer J., Lendeckel W. 2002. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Rights and permissions
About this article
Cite this article
Huang, Y., Zou, Q., Shen, X.J. et al. Differential expression of microRNA-2b with potential target coding P25 in the fifth instar larvae posterior silk gland of the silkworm. Mol Biol 45, 576–581 (2011). https://doi.org/10.1134/S0026893311040133
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893311040133