Abstract
Human uracil-DNA glycosylase SMUG1 removes uracil residues and some other noncanonical or damaged bases from DNA. Despite the functional importance of this enzyme, its X-ray structure is still unavailable. Previously, we performed homology modeling of human SMUG1 structure and suggested the roles of some amino acid residues in the recognition of damaged nucleotides and their removal from DNA. In this study, we investigated the kinetics of conformational transitions in the protein and in various DNA substrates during enzymatic catalysis using the stopped-flow method based on changes in the fluorescence intensity of enzyme’s tryptophan residues and 2-aminopurine in DNA or fluorescence resonance energy transfer (FRET) between fluorophores in DNA. The kinetic mechanism of interactions between reaction intermediates was identified, and kinetic parameters of the intermediate formation and dissociation were calculated. The obtained data help in elucidating the functions of His239 and Arg243 residues in the recognition and removal of damaged nucleotides by SMUG1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Uracil is one of the most common types of DNA damage which is formed by spontaneous [1-3] or enzymatic [4, 5] deamination of cytosine or erroneous incorporation of uridine residue during replication [6]. In human DNA, uracil is recognized and removed by several DNA glycosylases, including uracil-DNA N-glycosylase (UNG2), thymine-DNA glycosylase (TDG), single-stranded DNA-specific monofunctional DNA glycosylase (SMUG1), and methyl-CpG–binding domain 4 (MBD4) [7, 8]. Even though all uracil-DNA glycosylases hydrolyze N-glycosidic bond involving uracil, they differ in their ability to recognize single-stranded DNA substrates, T/G mismatches [9-13], or certain oxidized bases, such as 5-hydroxymethyluracil, 5-formyluracil, and 5-hydroxyuracil, which are natural derivatives of 5-methylcytosine formed in the process of epigenetic regulation [14].
One of the important problems in DNA repair studies is elucidation of the mechanisms of high-fidelity recognition of damaged bases. Recently, these studies have been focused on various DNA glycosylases in order to understand how a few damaged heterocyclic bases in genomic DNA can be searched for and recognized among numerous unmodified bases [15]. Published data [16] suggest that DNA glycosylases have a common mechanism for the interaction with substrates, despite completely different structures of the DNA-binding and active sites and different amino acid residues involved in the specific recognition of damaged nucleotide and catalysis. For example, most of DNA glycosylases with the known structure bend DNA and evert either damaged nucleotide or nucleotide complementary to it out of the DNA double helix. Usually, the damaged nucleotide is located in the active site pocket of the enzyme, where final nucleotide verification occurs. Some amino acid residues of the enzyme are inserted in the void formed in the DNA molecule and stabilize the extrahelical position of the damaged nucleotide. Next, hydrolysis of the N-glycosidic bond between the damaged heterocyclic base and ribose takes place, with the formation of apurinic/apyrimidinic site (AP site).
At present, the most corroborated mechanism for the recognition of damaged nucleotides by DNA glycosylases at the initial interaction stages suggests that the enzyme inserts into the DNA double helix an amino acid residue (wedge residue) that acts as a damage sensor [17]. This model of DNA damage recognition has been substantially developed in the studies using single-molecule kinetics [18-21] and pre-steady-state kinetics methods [22]. These techniques allow real-time visualization of the enzyme movement on a DNA molecule during the search for a damaged nucleotide and enable studies on the dynamics of conformational rearrangements in the enzyme and DNA (at both initial and later stages of damage recognition) and their catalytic transformations.
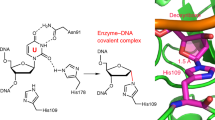
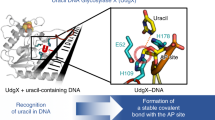
Currently, the structure of human DNA glycosylase SMUG1 remains unknown. Previously [23], we carried out homology modeling based on the structures of SMUG1 enzymes from Geobacter metallireducens (Protein Data Bank [PDB] ID 5H98 and 5H9I) [24] and Xenopus laevis (PDB 1OE4 and 10OE5) [10] and derived the structural model of the human enzyme in a free state and in a complex with uridine-containing DNA (Fig. 1). It was shown that 11 amino acid residues (239-249) of human SMUG1 form an intercalating loop that can serve as the wedge penetrating the DNA double helix near the damaged nucleotide [10, 25]. Residues Arg243 (which occupies the space inside the DNA helix that is formed by uracil eversion) and His239 (which can make contacts with uracil located in the active site and DNA phosphate group) were proposed as functionally important amino acid residues of the intercalating loop. Mutational analysis [23, 25] has revealed that His239 is necessary for the catalytic hydrolysis of the N-glycosidic bond.
We also proposed a kinetic mechanism for the interaction of the wild type (WT) SMUG1 with damaged DNA (Scheme 1) [26] using the stopped-flow method for the registration of changes in the fluorescence intensity of (i) Trp residues in the enzyme and (ii) 2-aminopurine (aPu) in DNA and fluorescence resonance energy transfer (FRET) in the FAM/BHQ1 (6-carboxyfluorescein/black hole quencher) pair in model DNA substrates. It was demonstrated that SMUG1 interaction with DNA includes at least two stages (1 and 2) of DNA binding and damaged nucleotide recognition that lead to the formation of catalytically competent complex, in which irreversible catalytic hydrolysis of the N-glycosidic bond (stage 3) with uracil takes place. The last stage 4 of the kinetic mechanism is an equilibrium dissociation of the enzyme–product complex.
Kinetic mechanism of the human WT SMUG1 interaction with damaged DNA (stages 1-4) [26]. E is SMUG1, U is uracil-containing substrate; (E•U)n is enzyme–substrate complex; E•P is enzyme–product (P) complex.
Next, we analyzed fluorescence changes characterizing conformational transitions in both SMUG1 and DNA molecules during the catalytic cycle. At stage 1 of interaction, translocation of the enzyme’s intercalating loop is accompanied by partial melting of the DNA duplex near the damaged nucleotide. Apparently, formation of the enzyme–substrate contacts at this stage induces partial eversion of the damaged base out of the DNA duplex. Stage 2 involves intercalation of amino acid residues of the enzyme’s intercalating loop. Previously [23], FRET study of the conformational dynamics of DNA substrate together with polyacrylamide gel electrophoresis (PAGE) analysis of the kinetics of accumulation of N-glycosylase reaction products revealed that the SMUG1 H239A mutant is catalytically inactive, whereas the R243A mutant retains its full activity. To clarify the role of Arg243 and His239 in the specific recognition of damaged nucleotides and catalysis, here we performed comprehensive pre–steady-state kinetic analysis of conformational changes in the SMUG1 mutants R243A and H239A, substrate DNA, and product of its transformation. For this purpose, we recorded changes in the fluorescence intensity of (i) Trp residues in the enzyme molecule and (ii) aPu situated on the 3′-side of the AP site in DNA, as well as changes in the FRET signal of the FAM/BHQ1 pair introduced at the 3′- and 5′-termini of the DNA duplex. This approach allowed us to monitor in more detail conformational transitions in the enzyme and DNA molecules during their interaction and to clarify the role of functionally important His239 and Arg243 residues in the recognition of damaged bases and their removal by human DNA glycosylase SMUG1.
MATERIALS AND METHODS
Acrylamide, N,N-methylenebisacrylamide, dithiothreitol, urea, EDTA, acetonitrile, glycerol, and Tris were from Sigma-Aldrich (USA).
DNA substrates. DNA substrates and enzyme ligands were 17-mer DNA duplexes presented in Table 1. The oligonucleotides were purified by HPLC on a PRP-X500 ion-exchange column (3.9 × 300 mm; particle size, 12-30 μm; Hamilton Company, USA) followed by reversed-phase chromatography on a Nucleoprep 100-20 C18 column (10 × 250 mm; Macherey-Nagel, Germany). The purity of the obtained preparations was verified by PAGE in denaturing 20% gel.
SMUG1 enzyme. Mutant SMUG1 R243A and H239A proteins were isolated from E. coli Rosetta 2 cells, transformed with the pET28c plasmid carrying the corresponding human uracil-DNA glycosylase gene, as described elsewhere [23, 26]. All enzymatic assays were carried out at 25°C in 50 mM Tris-HCl (pH 7.5) containing 50 mM KCl, 1 mM EDTA, 1 mM dithiothreitol, and 7% (v/v) glycerol.
Kinetic analysis by the stopped-flow method. Fluorescence kinetic curves were obtained using an SX.20 stopped-flow spectrometer (Applied Photophysics, UK) as described elsewhere [27-30]. SMUG1 contains four Trp and four Tyr residues. SMUG1 fluorescence was excited at 290 nm and registered at wavelengths >320 nm (WG-320 filter; Schott, Germany). Under these conditions, the main contribution (>90%) to the protein fluorescence was from Trp residues. When aPu-containing substrates were used, fluorescence was excited at 310 nm and registered at wavelengths >370 nm (LG-370 filter; Corion, USA). To measure the FRET signal of the FAM/BHQ1 pair, FAM fluorescence was excited at 494 nm and registered at wavelengths >515 nm (OG-515 filter; Schott). The instrument dead time was 1.0 ms. Each kinetic curve was averaged from at least three recordings.
Analysis of kinetic data. To identify the minimal kinetic scheme describing the enzyme–substrate interactions and to calculate the rate constants of conformational transitions at stages 1-4 of this scheme, a set of kinetic curves was obtained for different enzyme and substrate concentrations. Quantitative analysis of the experimental results was conducted with the DynaFit software (BioKin, USA) [31] via optimization for the values of parameters in the kinetic scheme, as described before [32-34].
RESULTS
SMUG1 R243A interaction with the stable analog of N-glycosylase reaction product. To study conformational changes in the SMUG1 R243A mutant during its interaction with the product of N-glycosidic bond hydrolysis, we used a model DNA duplex with a tetrahydrofuran analog of the AP site (F-ligand, Table 1). Enzyme interaction with the product analog led to the two-phase decline in Trp fluorescence (Fig. 2a). The signal decrease in the initial region of kinetic curves (30-50 ms) corresponded to the formation of first nonspecific contacts with DNA. The following slight decrease in the fluorescence took place from 1 to 10 s. Conformational changes in SMUG1 R243A during its interaction with the F-ligand were described with Scheme 2, which features two reversible stages. The resulting rate constants are listed in Table 2.
The nature of processes occurring at these stages of enzyme–protein interaction was identified by analyzing the changes in the fluorescence of aPu residue located on the 3′ side of the F-site (F-aPu-ligand, Table 1). This fluorophore is sensitive to changes in the medium dielectric permeability, and its fluorescence decreases with an increase in the microenvironment hydrophobicity [35]. Enzyme interaction with the F-aPu-ligand caused a decrease in the fluorescence intensity of the aPu residue at time points before 200 ms (Fig. 2b), which indicated formation of hydrophobic environment around aPu, e.g., due to the DNA interaction with the amino acid residues of the intercalating loop. The obtained kinetic curves were described by Scheme 3, and the corresponding rate constants are given in Table 2.
At the same time, the kinetic curves obtained by registering the FRET signal during SMUG1 R243A interaction with the FRET-F-ligand demonstrate a decrease in the FRET signal in the initial region, before t ≅ 100-200 ms (Fig. 2c). This suggest a decrease in the distance between the fluorophores, which might happen in the course of DNA bending during its binding to the enzyme. Analysis of kinetic curves showed that this process can be described as a two-stage equilibrium mechanism (Scheme 2) with the rate constants listed in Table 2.
Therefore, SMUG1 R243A interaction with the reaction product analog F includes at least two reversible stages. Analysis of the kinetic curve shapes (Fig. 2) and rate constants (Table 2) suggests that the primary complex formation is accompanied by conformational changes in the enzyme (stage 1, Trp, t ≤ 30-50 ms), followed by alterations in the microenvironment of the aPu residue (stage 1, aPu, t < 100 ms). These sequential conformational changes of the enzyme and DNA substrate indicate that the enzyme–substrate interaction is most likely associated with the movement of the intercalating loop and its interaction with the DNA duplex in the vicinity of the damaged nucleotide and is probably accompanied by eversion of the F-site into the enzyme active site. Formation of contacts between the DNA-binding site and the DNA duplex also induces the bending of the substrate molecule at times t < 100-200 ms (stage 1, FRET data). Subsequent slower processes (stage 2, FRET and Trp data) revealed by changes in Trp fluorescence and FRET signal point to additional conformational rearrangements in the enzyme molecule and F-containing product analog.
Obtained data indicate that each fluorophore detects different stage of conformational rearrangements in the interacting molecules. The second stage is the slowest one and is detectable via Trp fluorescence, which is most likely related to the adaptation of the SMUG1 structure to the structure of the F-containing DNA ligand that models N-glycosylase reaction product.
SMUG1 R243A interaction with uridine-containing DNA. We also investigated the full enzymatic cycle using uridine-containing DNA substrates (Table 1). The enzymatic reaction includes substrate binding by the enzyme, formation of catalytically competent complex, hydrolysis of the N-glycosidic-bond, and enzyme dissociation from the product.
The kinetic curves characterizing changes in Trp fluorescence (Fig. 3a) demonstrate a decrease in the fluorescence signal at t ≈ 0.3 s followed by the increase to a plateau at t ≈ 5 s. Comparison of these kinetic curves with the curves obtained for the enzyme–product binding (Fig. 2a) suggests that during formation of the catalytic complex with the U-substrate, the curves display similar changes in Trp fluorescence in their initial parts. The increase in Trp fluorescence intensity at the time points ~0.5-10 s, which is absent in the case of F-ligand, can be ascribed to the N-glycosidic bond hydrolysis and dissociation of the enzyme–product complex. Earlier, it has been shown for the WT SMUG1 [26] that dissociation of the enzyme–product complex is the rate-limiting stage of the enzymatic process. Therefore, it is unlikely that dissociation of the enzyme–product complex occurs within the 0.5-10 s time interval. Indeed, analysis of kinetic curves revealed that they correspond to the reaction Scheme 4, which includes two reversible stages of substrate binding, followed by the irreversible catalytic stage resulting in the formation of the enzyme–product complex (Table 3).
Based on the changes in the fluorescence of aPu in the U-aPu-substrate (Fig. 3b) within the first 100 ms, the kinetics of conformational changes in DNA in the reaction with SMUG1 R234A was similar to the kinetics of enzyme interaction with the product analog F (Fig. 2b), indicating that hydrolysis of the N-glycosidic bond does not require major changes in the substrate geometry. Besides, in the case of the U-aPu-substrate, we observed a slight increase in the fluorescence intensity at later time points (≤10 s), which was probably related to slow dissociation of the enzyme–product complex. The obtained kinetic curves were described by the Scheme 5, which allowed to estimate the dissociation rate constant koff for the enzyme–product complex (Table 3).
FRET curves for the SMUG1 R243A interaction with the FRET-U-substrate (Fig. 3c) feature a decrease in the FAM fluorescence in the time interval <10 s. In this case, the decline of the FRET signal can be divided into two phases: rapid decrease before t ≈ 100 ms (as with the FRET-F-ligand) and subsequent slow decrease before t ≈ 3 s. It can be theorized that the FRET curves do not reflect the catalytic process, because changes in the aPu fluorescence show that hydrolysis of the N-glycosidic bond does not require additional conformational changes in DNA. The resulting kinetic curves were described with Scheme 6, which features two equilibrium stages of DNA binding (Table 3).
It should be noted that the initial parts of kinetic curves were similar for the F-ligands and U-substrates, regardless of the fluorophore used (Figs. 2 and 3). This finding suggests that the corresponding processes of the enzyme–substrate complex formation include identical stages and are not affected by the nature of damaged nucleotide.
SMUG1 H239A interaction with the uridine-containing DNA. As reported previously [23], the activity of the SMUG1 H239A mutant toward U-containing DNA is substantially lower than that of the WT enzyme or the R243A mutant. Nonetheless, formation of the enzyme–substrate complex between SMUG1 H239A and FRET-U-substrate was registered as changes in the FRET signal within a time interval of 100 s (Fig. 4). The analysis of the enzyme activity by PAGE demonstrated only an insignificant accumulation of the reaction product amount within this period. Therefore, it can be concluded that the processes revealed by the stopped-flow method correspond to the DNA binding. As shown in Fig. 4, the kinetic curves describing this interaction were characterized by a three-phase decrease in the FRET signal, which was interpreted with Scheme 7 that includes three reversible stages (Table 4).
DISCUSSION
Analysis of model structures of the WT SMUG1 enzyme and its complex with DNA [23, 26] showed that amino acid residues His239 and Arg243 should alter their conformation for intercalation into DNA and formation of contacts with the everted damaged base (Fig. 1). At the same time, the H239A and R243A substitutions should not substantially disrupt the network of protein–DNA contacts, despite the lost ability for the formation of hydrogen bonds with DNA. Furthermore, the SMUG1 R243A mutant retains the catalytic activity, whereas the H239A mutant almost completely lacks it [23].
In this work, the role of His239 and Arg243 residues was investigated by registering conformational changes in the enzyme and model DNA substrates during the enzyme–ligand interaction. To identify conformational changes in the SMUG1 R243A and SMUG1 H239A proteins in the course of DNA binding, we used model DNA duplexes containing either uridine or F-site.
Interaction of catalytically active SMUG1 R243A with the uridine-containing DNA substrate or F-site-containing DNA product was detected via changes in the fluorescence of protein Trp residues, aPu residue in DNA, or FRET pair FAM/BHQ1 (Figs. 2 and 3). Comparative analysis of the conformational dynamics of the SMUG1 H239A mutant (which has low catalytic activity) during its interaction with the DNA product also revealed changes in the fluorescence of Trp residues and aPu and in the FRET signal, while formation of the enzyme–substrate complex with the uridine-containing DNA was successfully registered only by the FRET method (Fig. 4).
Overall, our experimental data on the conformational transitions in the SMUG1 mutants R243A and H239A and DNA, along with the data obtained for the WT enzyme [26], allowed us to propose that SMUG1 undergoes several stages of conformational changes during specific DNA binding and catalysis (Scheme 1) [26] (Table 5). The general kinetic mechanism involves two reversible stages corresponding to the formation of catalytically competent complex. These stages were successfully detected with all three types of fluorophores used for signal registration, thus indicating mutual conformational changes in both the enzyme and the DNA substrate. The first binding stage includes translocation of the enzyme’s intercalating loop, local melting of DNA near the damaged nucleotide, and eversion of the damaged base from the DNA duplex. The movement of the intercalating loop is probably related to the intercalation of amino acid residue (damage sensor) into the DNA duplex, which is a process responsible for the initial recognition of damaged nucleotide. We believe this residue to be Arg243, which is consistent with conclusions in [23]. At the second stage, the catalytically active complex is formed, in which amino acid residues of the intercalating loop are completely inserted into DNA and form all possible contacts with the everted nucleotide, including contacts between His239 and phosphate groups. The reaction then proceeds to the third irreversible stage. Previously, it has been reported that dissociation of the enzyme–product complex is as a rate-limiting stage of catalysis by the WT enzyme [26]. Because the R243A substitution reduces the number of hydrogen bonds formed with DNA, it destabilizes both the enzyme–substrate complex and the complex of the enzyme with the reaction product, which increases the enzyme turnover rate. Comparison of the rate constants of individual stages for WT SMUG1 based on Trp fluorescence [26] with the data for the R243A mutant (Table 3) shows that their catalytic activities are comparable (k3 = 1.7 and 2.0 s1 for WT and R243A enzymes, respectively). In contrast, the rates of the first and second stages, which characterize substrate recognition and binding, are substantially higher for WT SMUG1 than for the R243A mutant. Similar conclusion can be drawn from the kinetic data obtained using aPu fluorescence and FRET, both for the R243A mutant and the H239A mutant. At the same time, the enzyme affinity for the DNA substrate is unchanged. Therefore, we can conclude that Arg243 participates in the recognition of the damaged nucleotide but is not required at the catalytic stage, whereas His239 is necessary for both recognition of the damaged nucleotide and catalysis.
Abbreviations
- AP site:
-
apurinic/apyrimidinic site
- aPu:
-
2-aminopurine
- BHQ1:
-
black hole fluorescence quencher
- F:
-
(2R,3S)-2-(hydroxymethyl)-3-hydroxytetrahydrofuran
- FAM:
-
6-carboxyfluorescein
- FRET:
-
fluorescence resonance energy transfer
- PAGE:
-
polyacrylamide gel electrophoresis
- SMUG1:
-
single-stranded DNA-specific monofunctional DNA glycosylase
- WT:
-
wild typ
REFERENCES
Lari, S.-U., Chen, C.-Y., Vertéssy, B. G., Morré, J., and Bennett, S. E. (2006) Quantitative determination of uracil residues in Escherichia coli DNA: contribution of ung, dug, and dut genes to uracil avoidance, DNA Repair (Amst), 5, 1407-1420.
Lindahl, T. (1993) Instability and decay of the primary structure of DNA, Nature, 362, 709-715.
Lewis, C. A., Crayle, J., Zhou, S., Swanstrom, R., Wolfenden, R., and Wolfenden, R. (2016) Cytosine deamination and the precipitous decline of spontaneous mutation during Earth’s history, Proc. Natl. Acad. Sci. USA, 113, 8194-8199.
Jaszczur, M., Bertram, J. G., Pham, P., Scharff, M. D., and Goodman, M. F. (2013) AID and Apobec3G haphazard deamination and mutational diversity, Cell. Mol. Life Sci., 70, 3089-108.
Rebhandl, S., Hümer, M., Greil, R., and Geisberger, R. (2015) AID/APOBEC deaminases and cancer, Oncoscience, 2, 320.
Ladner, R. D. (2001) The role of dUTPase and uracil-DNA repair in cancer chemotherapy, Curr. Protein Pept. Sci., 2, 361-370.
Jacobs, A. L., and Schar, P. (2012) DNA glycosylases: in DNA repair and beyond, Chromosoma, 121, 1-20.
Visnes, T., Doseth, B., Pettersen, H. S., Hagen, L., Sousa, M. M., Akbari, M., Otterlei, M., Kavli, B., Slupphaug, G., and Krokan, H. E. (2009) Uracil in DNA and its processing by different DNA glycosylases, Philos. Trans. R. Soc. Lond. B Biol. Sci., 364, 563-568.
Kavli, B., Sundheim, O., Akbari, M., Otterlei, M., Nilsen, H., Skorpen, F., Aas, P. A., Hagen, L., Krokan, H. E., and Slupphaug, G. (2002) hUNG2 is the major repair enzyme for removal of uracil from U : A matches, U : G mismatches, and U in single-stranded DNA, with hSMUG1 as a broad specificity backup, J. Biol. Chem., 277, 39926-39936.
Wibley, J. E. A., Waters, T. R., Haushalter, K., Verdine, G. L., and Pearl, L. H. (2003) Structure and specificity of the vertebrate anti-mutator uracil-DNA glycosylase SMUG1, Mol. Cell, 11, 1647-1659.
Hashimoto, H., Hong, S., Bhagwat, A. S., Zhang, X., and Cheng, X. (2012) Excision of 5-hydroxymethyluracil and 5-carboxylcytosine by the thymine DNA glycosylase domain: its structural basis and implications for active DNA demethylation, Nucleic Acids Res., 40, 10203-10214.
Hashimoto, H., Zhang, X., and Cheng, X. (2012) Excision of thymine and 5-hydroxymethyluracil by the MBD4 DNA glycosylase domain: structural basis and implications for active DNA demethylation, Nucleic Acids Res., 40, 8276-8284.
Sjolund, A. B., Senejani, A. G., and Sweasy, J. B. (2013) MBD4 and TDG: multifaceted DNA glycosylases with ever expanding biological roles, Mutat. Res., 743-744, 12-25.
Ito, S., and Kuraoka, I. (2015) Epigenetic modifications in DNA could mimic oxidative DNA damage: a double-edged sword, DNA Repair (Amst.), 32, 52-57.
Kuznetsov, N. A., and Fedorova, O. S. (2020) Kinetic milestones of damage recognition by DNA glycosylases of the Helix-hairpin-Helix structural superfamily, Adv. Exp. Biol. Med., (in press).
Brooks, S. C., Adhikary, S., Rubinson, E. H., and Eichman, B. F. (2013) Recent advances in the structural mechanisms of DNA glycosylases, Biochim. Biophys. Acta, 1834, 247-271.
Dunn, A. R., Kad, N. M., Nelson, S. R., Warshaw, D. M. and Wallace, S. S. (2011) Single Qdot-labeled glycosylase molecules use a wedge amino acid to probe for lesions while scanning along DNA, Nucleic Acids Res., 39, 7487-7498.
Lee, A. J., Warshaw, D. M., and Wallace, S. S. (2014) Insights into the glycosylase search for damage from single-molecule fluorescence microscopy, DNA Repair, 20, 23-31.
Nelson, S. R., Dunn, A. R., Kathe, S. D., Warshaw, D. M., and Wallace, S. S. (2014) Two glycosylase families diffusively scan DNA using a wedge residue to probe for and identify oxidatively damaged bases, Proc. Natl. Acad. Sci. USA, 111, E2091-E2099.
Lee, A. J., and Wallace, S. S. (2016) Visualizing the search for radiation-damaged DNA bases in real time, Radiat. Phys. Chem. Oxf. Engl. 1993, 128, 126-133.
Lee, A. J., and Wallace, S. S. (2017) Hide and seek: how do DNA glycosylases locate oxidatively damaged DNA bases amidst a sea of undamaged bases? Free Radic. Biol. Med., 107, 170-178.
Kuznetsov, N. A., and Fedorova, O. S (2016) Thermodynamic analysis of fast stages of specific lesion recognition by DNA repair enzymes, Biochemistry (Moscow), 81, 1136-1152.
Iakovlev, D. A., Alekseeva, I. V., Vorobjev, Y. N., Kuznetsov, N. A., and Fedorova, O. S. (2019) The role of active-site residues Phe98, HiS239, and Arg243 in DNA binding and in the catalysis of human uracil–DNA glycosylase SMUG1, Molecules, 24, doi: 10.3390/molecules24173133.
Zhang, Z., Shen, J., Yang, Y., Li, J., Cao, W., and Xie, W. (2016) Structural basis of substrate specificity in geobacter metallireducens SMUG1, ACS Chem. Biol., 11, 1729-1736.
Matsubara, M., Tanaka, T., Terato, H., Ohmae, E., Izumi, S., Katayanagi, K., and Ide, H. (2004) Mutational analysis of the damage-recognition and catalytic mechanism of human SMUG1 DNA glycosylase, Nucleic Acids Res., 32, 5291-5302.
Kuznetsova, A. A., Iakovlev, D. A., Misovets, I. V., Ishchenko, A. A., Saparbaev, M. K., Kuznetsov, N. A., and Fedorova, O. S. (2017) Pre-steady-state kinetic analysis of damage recognition by human single-strand selective monofunctional uracil-DNA glycosylase SMUG1, Mol. Biosyst., 13, 2638-2649.
Kuznetsov, N. A., Faleev, N. G., Kuznetsova, A. A., Morozova, E. A., Revtovich, S. V., Anufrieva, N. V., Nikulin ,A. D. D., Fedorova, O. S., and Demidkina, T. V. (2015) Pre-steady-state kinetic and structural analysis of interaction of methionine gamma-lyase from Citrobacter freundii with inhibitors, J. Biol. Chem., 290, 671-681.
Kladova, O. A., Krasnoperov, L. N., Kuznetsov, N. A., and Fedorova, O. S. (2018) Kinetics and thermodynamics of DNA processing by wild type DNA-glycosylase endo III and its catalytically inactive mutant forms, Genes (Basel), 9, doi: 10.3390/genes9040190.
Kladova, O. A., Kuznetsova, A. A., Fedorova, O. S., and Kuznetsov, N. A. (2017) Mutational and kinetic analysis of lesion recognition by Escherichia coli endonuclease VIII, Genes (Basel), 8, 1-13.
Yakovlev, D. A., Kuznetsova, A. A., Fedorova, O. S., and Kuznetsov, N. A. (2017) Search for modified DNA sites with the human methyl-CpG-binding enzyme MBD4, Acta Naturae, 9, 88-98.
Kuzmic, P. (1996) Program DYNAFIT for the analysis of enzyme kinetic data: application to HIV proteinase, Anal. Biochem., 237, 260-273.
Alekseeva, I. V., Davletgildeeva, A. T., Arkova, O. V., Kuznetsov, N. A., and Fedorova, O. S. (2019) The impact of single-nucleotide polymorphisms of human apurinic/apyrimidinic endonuclease 1 on specific DNA binding and catalysis, Biochimie, 163, 73-83.
Kladova, O. A., Kuznetsov, N. A. and Fedorova, O. S. (2019) Thermodynamic parameters of endonuclease VIII interactions with damaged DNA, Acta Naturae, 11, 29-37.
Miroshnikova, A. D., Kuznetsova, A. A., Kuznetsov, N. A., and Fedorova, O. S. (2016) Thermodynamics of damaged DNA binding and catalysis by human AP endonuclease 1, Acta Naturae, 8, 103-110.
Rachofsky, E. L., Osman, R., and Ross, J. B. A. (2001) Probing structure and dynamics of DNA with 2-aminopurine: effects of local environment on fluorescence, Biochemistry, 40, 946-956.
Funding
This work was supported by the Russian Science Foundation (project No. 16-14-10038) with partial governmental support for equipment use (project AAAA-A17-117020210022-4).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
This article does not contain studies with human participants or animals performed by any of the authors. The authors declare no conflict of interests.
Rights and permissions
Open access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License ( http://creativecommons.org/licenses/by/4.0/ ), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Iakovlev, D., Alekseeva, I., Kuznetsov, N. et al. Role of Arg243 and His239 Residues in the Recognition of Damaged Nucleotides by Human Uracil-DNA Glycosylase SMUG1. Biochemistry Moscow 85, 594–603 (2020). https://doi.org/10.1134/S0006297920050089
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297920050089