Abstract
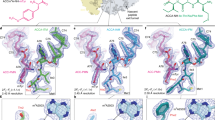
Chloramphenicol amine peptide derivatives containing tripeptide fragments of regulatory “stop peptides”–MRL, IRA, IWP–were synthesized. The ability of the compounds to form ribosomal complexes was studied by displacement of the fluorescent erythromycin analog from its complex with E. coli ribosomes. It was found that peptide chloramphenicol analogs are able to bind to bacterial ribosomes. The dissociation constants were 4.3-10 μM, which is 100-fold lower than the corresponding values for chloramphenicol amine–ribosome complex. Interaction of the chloramphenicol peptide analogs with ribosomes was simulated by molecular docking, and the most probable contacts of “stop peptide” motifs with the elements of nascent peptide exit tunnel were identified.
Similar content being viewed by others
Abbreviations
- Bhoc:
-
N-benzhydryloxycarbonyl
- Boc:
-
tert-butyloxycarbonyl
- BODIPY:
-
(4,4-difluoro-4-bora-5,7-dimethyl)-3a,4a-diaza-s-indacene-3-pentanoic acid
- Caeg:
-
3-(2-aminoethyl)-3-[2-(cytosin-1-yl)acetyl]glycine
- DCC:
-
1,3-dicyclohexylcarbodiimide
- DIPEA:
-
diisopropylethylamine
- Ery:
-
erythromycin
- Fmoc:
-
fluorenylmethyloxycarbonyl
- LCMS:
-
liquid chromatography-mass spectrometry
- PNA:
-
peptide-nucleic acids
- PTC:
-
peptidyl transferase center
- RT:
-
ribosomal tunnel
- TFA:
-
trifluoroacetic acid
References
Ban, N., Nissen, P., Hansen, J., Moore, P. B., and Steitz, T. A. (2000) The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution, Science, 289, 905–920.
Nissen, P., Hansen, J., Ban, N., Moore, P. B., and Steitz, T. A. (2000) The structural basis of ribosome activity in peptide bond synthesis, Science, 289, 920–930.
Harms, J., Schluenzen, F., Zarivach, R., Bashan, A., Gat, S., Agmon, I., Bartels, H., Franceschi, F., and Yonath, A. (2001) High-resolution structure of the large ribosomal subunit from a mesophilic eubacterium, Cell, 107, 679–688.
Bogdanov, A. A., Sumbatyan, N. V., Shishkina, A. V., Karpenko, V. V., and Korshunova, G. A. (2010) Ribosomal tunnel and translation regulation, Biochemistry (Moscow), 75, 1501–1516.
Kolb, V. A. (2010) Properties of intraribosomal part of nascent polypeptide, Biochemistry (Moscow), 75, 1517–1527.
Wilson, D. N., and Beckman, R. (2011) The ribosomal tunnel as a functional environment for nascent polypeptide folding and translational stalling, Curr. Opin. Struct. Biol., 21, 274–282.
Subramanian, S. L., Ramu, H., and Mankin, A. S. (2012) in Antibiotic Discovery and Development (Dougherty, T. J., and Pucci, M. J., eds.) Springer, pp. 455–484.
La Marre, J., Mendes, R. E., Szal, T., Schwarz, S., Jones, R. N., and Mankin, A. S. (2013) The genetic environment of the cfr gene and the presence of other mechanisms account for the very high linezolid resistance of Staphylococcus epidermidis isolate 426-3147L, Antimicrob. Agents Chemother., 57, 1173–1179.
Mankin, A. S. (2006) Nascent peptide in the “birth canal” of the ribosome, Trends Biochem. Sci., 31, 11–13.
Cruz-Vera, L. R., Sachs, M. S., Sguires, C. L., and Yanofsky, C. (2011) Nascent polypeptide sequences that influence ribosome function, Curr. Opin. Microbiol., 14, 160–166.
Ito, K., and Chiba, S. (2013) Arrest peptides: cis-acting modulators of translation, Annu. Rev. Biochem., 82, 171–202.
Arenz, S., Meydan, S., Starosta, A. L., Berninghausen, O., Beckmann, R., Vazquez-Laslop, N., and Wilson, D. N. (2014) Drug sensing by the ribosome induces translational arrest via active site perturbation, Mol. Cell, 56, 446–452.
Roy, R. N., Lomakin, I. B., Gagnon, M. G., and Steitz, T. A. (2015) The mechanism of inhibition of protein synthesis by the proline-rich peptide oncocin, Nat. Struct. Mol. Biol., 22, 466–469.
Seefeldt, A. C., Nguyen, F., Antunes, S., Perebaskine, N., Graf, M., Arenz, S., Inampudi, K. K., Douat, C., Guichard, G., Wilson, D. N., and Innis, C. A. (2015) The proline-rich antimicrobial peptide Onc112 inhibits translation by blocking and destabilizing the initiation complex, Nat. Struct. Mol. Biol., 22, 470–475.
Hansen, J. L., Moore, P. B., and Steitz, T. A. (2003) Structures of five antibiotics bound at the peptidyl transferase center of the large ribosomal subunit, J. Mol. Biol., 330, 1061–1075.
Schlunzen, F., Zarivach, R., Harms, J., Bashan, A., Tocilj, A., Albrecht, R., Yonath, A., and Franceschi, F. (2001) Structural basis for the interaction of antibiotics with the peptidyl transferase center in eubacteria, Nature, 413, 814–821.
Lu, J., Hua, Z., Kobertz, W. R., and Detsch, C. (2013) Nascent peptide side-chains induce rearrangements in distinct locations of the ribosomal tunnel, J. Mol. Biol., 411, 499–510.
Woolstenhulme, C. J., Parajuli, S., Healey, D. W., Valverde, D. P., Petersen, E. N., Starosta, A. L., Guydosh, N. R., Johnson, W. E., Wilson, D. N., and Buskirk, A. R. (2013) Nascent peptides that block protein synthesis in bacteria, Proc. Natl. Acad. Sci. USA, 110, 878–887.
Mamos, P., Krokidis, M. G., Papadas, A., Karahalios, P., Starosta, A. L., Wilson, D. N., Kalpaxis, D. L., and Dinos, G. P. (2013) On the use of the antibiotic chloramphenicol to target polypeptide chain mimics to the ribosomal exit tunnel, Biochimie, 95, 1765–1772.
Arenz, S., Ramu, H., Gupta, P., Berninghausen, O., Beckmann, R., Vazquez-Laslop, N., Mankin, A. S., and Wilson, D. N. (2014) Molecular basis for erythromycindependent ribosome stalling during translation of the ErmBL leader peptide, Nat. Commun., 5, 3501.
Fischer, N., Neumann, P., Konevega, A. L., Bock, L. V., Ficner, R., Rodnina, M. V., and Stark, H. (2015) Structure of the E. coli ribosome–EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM, Nature, 520, 567–570.
Sumbatyan, N. V., Korshunova, G. A., and Bogdanov, A. A. (2003) Peptide derivatives of antibiotics tylosin and desmycosin, protein synthesis inhibitors, Biochemistry (Moscow), 68, 1156–1158.
Starosta, A. L., Karpenko, V. V., Shishkina, A. V., Mikolajka, A., Sumbatyan, N. V., Schluenzen, F., Korshunova, G. A., Bogdanov, A. A., and Wilson, D. N. (2010) Interplay between the ribosomal tunnel, nascent chain, and macrolides influences drug inhibition, Chem. Biol., 17, 504–514.
Shishkina, A., Makarov, G., Tereshchenkov, A., Korshunova, G., Sumbatyan, N., Golovin, A., Svetlov, M., and Bogdanov, A. (2013) Conjugates of amino acids and peptides with 5-O-mycaminosyltylonolide and their interaction with the ribosomal exit tunnel, Bioconj. Chem., 24, 1861–1869.
Vazquez-Laslop, N., Ramu, H., and Mankin, A. (2011) in Ribosomes: Structure, Function and Dynamics (Rodnina, M. V., Wintermeyer, W., and Green, R., eds.) Springer, WienN.Y., pp. 377–392.
Gumbart, J., Schreiner, E., Wilson, D., Beckmann, R., and Schulten, K. (2012) Mechanism of SecM-mediated stalling in the ribosome, Biophys. J., 103, 331–341.
Nielsen, P. E. (1998) Structural and biological properties of peptide nucleic acid (PNA), Pure Appl. Chem., 70, 105–110.
Lundin, K. E., Good, L., Stromberg, R., Graslund, A., and Smith, C. I. E. (2006) in Advances in Genetics (Hall, J. C., ed.), vol. 56, Academic Press, Waltham, pp. 1–51.
Good, L., and Nielsen, P. E. (1998) Inhibition of translation and bacterial growth by peptide nucleic acid targeted to ribosomal RNA, Proc. Natl. Acad. Sci. USA, 3, 2073–2076.
Rebstock, M. C., Crooks, H. M., Controulis, J., and Bartz, Q. R. (1949) Chloramphenicol (Chloromycetin). IV. Chemical Studies, J. Am. Chem. Soc., 71, 2458–2462.
Trott, O., and Olson, A. J. (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comput. Chem., 31, 455–461.
Dunkle, J. A., Xiong, L., Mankin A. S., and Cate, J. H. D. (2010) Structures of the Escherichia coli ribosome with antibiotics bound near the peptidyl transferase center explain spectra of drug action, Proc. Natl. Acad. Sci. USA, 107, 17152–17157.
Stewart, J. J. (2013) Optimization of parameters for semiempirical methods VI: more modifications to the NDDO approximations and re-optimization of parameters, J. Mol. Model., 19, 1–32.
Yan, K., Hunt, E., Berge, J., May, E., Copeland, R. A., and Gontarek, R. R. (2005) Fluorescence polarization method to characterize macrolide–ribosome interactions, Antimicrob. Agents Chemother., 49, 3367–3372.
Wang, Z. X. (1995) An exact mathematical expression for describing competitive binding of two different ligands to a protein molecule, FEBS Lett., 360, 111–114.
Shishkina, A. V., Tereshchenkov, A. G., Sumbatyan, N. V., Korshunova, G. A., and Bogdanov, A. A. (2013) Characterization of tylosin-related macrolides–ribosome interactions by fluorescence polarization method, FEBS J., 280 (Suppl. 1), 356.
Tereshchenkov, A., Sergeeva, V., Shishkina, A., Sumbatyan, N., and Bogdanov, A. (2014) in EMBO Conference Series: Chemical Biology 2014, Mera Druck GmbH, Sanghausen, pp. 263–263.
Tereshchenkov, A. G. (2013) in Kazan Summer School on Chemoinformatics, Innovation Publishing House “Butlerov Heritage”, Kazan, pp. 33–33.
Seidelt, B., Innis, C. A., Wilson, D. N., Gartmann, M., Armache, J.-P., Villa, E., Trabuco, L. G., Becker, T., Mielke, T., Schulten, K., Steitz, T. A., and Beckmann, R. (2009) Structural insight into nascent polypeptide chainmediated translational stalling, Science, 326, 1412–1415.
Sohmen, D., Chiba, S., Shimokawa-Chiba, N., Innis, C. A., Berninghausen, O., Beckmann, R., Ito, K., and Wilson, D. N. (2015) Structure of the Bacillus subtilis 70S ribosome reveals the basis for species-specific stalling, Nat. Commun., 6, 6941.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published in Russian in Biokhimiya, 2016, Vol. 81, No. 4, pp. 538-547.
Originally published in Biochemistry (Moscow) On-Line Papers in Press, as Manuscript BM15-322, January 31, 2016.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Tereshchenkov, A.G., Shishkina, A.V., Tashlitsky, V.N. et al. Interaction of chloramphenicol tripeptide analogs with ribosomes. Biochemistry Moscow 81, 392–400 (2016). https://doi.org/10.1134/S000629791604009X
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S000629791604009X