Abstract
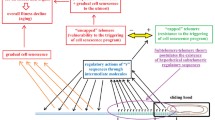
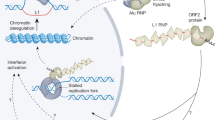
The nature of telomere functioning, which affects the aging of the body, reflects global processes in the genome conditioned by the regulatory influence of transposable elements that are sequentially activated during ontogenesis. This is determined by the fact that centromeres and centromere proteins, telomeres and telomerases, introns and spliceosome components, transcription factors and their binding sites, noncoding RNAs and their targets in protein-coding gene sequences evolutionarily originated from transposable elements. The interrelation of these structural and functional elements of the genome is dynamically changing in individual development and depends on the nature of activations and transpositions of transposable elements. Each species is characterized by a specific set of transposable elements and associated tandem repeats, which reflect the epigenetic tuning of ontogenesis, mainly as a part of centromeres and telomeres; the impact on them is promising for the development of mechanisms of lifespan regulation. This is due to the ability of ribozymes and peptides to interact with specific sequences of DNA nucleotides, especially in tandem repeats. An important approach to the study of the relationship of transposons with telomeres, centromeres, and subtelomeric regions for the regulation of aging may be the study of the role of the peptides and microRNAs that unite them, the complex application of which has high potential for geroprotective effectiveness.
Similar content being viewed by others
REFERENCES
Bondarev, I.E. and Khavinson, V.K., Suppression of alternative telomere lengthening in cancer cells with reverse transcriptase inhibitors, Adv. Gerontol., 2016, vol. 6, no. 4, pp. 272–274.
Mustafin, R.N. and Khusnutdinova, E.K., Noncoding parts of genomes as the basis of epigenetic heredity, Vavilovskii Zh. Genet. Sel., 2017, vol. 21, no. 6, pp. 742–749.
Mustafin, R.N. and Khusnutdinova, E.K., The interaction of transposons with epigenetic factors in aging, Usp. Gerontol., 2017, vol. 30, no. 4, pp. 516–528.
Pavlov, K.I., Mukhin, V.N., Klimenko, V.M., and Anisimov, V.N., The telomere-telomerase system and mental processes in aging, norm and pathology (literature review), Adv. Gerontol., 2017, vol. 7, no. 2, pp. 120–129.
Khavinson, V.Kh., Solovyov, A.Yu., and Shataeva, L.K., Molecular mechanism of interaction between oligopeptides and double-stranded DNA, Bull. Exp. Biol. Med., 2006, vol. 141, no. 4, pp. 457–461.
Khavinson, V.Kh., Peptidnaya regulyatsiya stareniya (Peptide Regulation of Aging), St. Petersburg: Nauka, 2009.
Alter, B.P., Giri, N., Savage, S.A., and Rosenberg, P.S., Cancer in dyskeratosis congenital, Blood, 2009, vol. 113, no. 26, pp. 6549–6557.
Alzohairy, A.M., Gyulai, G., Jansen, R.K., and Bahieldin, A., Transposable elements domesticated and neofunctionaized by eukaryotic genomes, Plasmid, 2013, vol. 69, pp. 1–15.
Arkhipova, I.R., Yushenova, I.A., and Rodriguez, F., Giant reverse transcriptase-encoding transposable elements at telomeres, Mol. Biol. Evol., 2017, vol. 34, no. 9, pp. 2245–2257.
Armanios, M., Syndromes of telomere shortening, Annu. Rev. Genomics Hum. Genet., 2009, vol. 10, pp. 45–61.
Aschacher, T., Sampl, S., Kaser, L., et al., The combined use of known antiviral reverse transcriptase inhibitors AZT and DDI induce anticancer effects at low concentrations, Neoplasia, 2012, vol. 14, pp. 44–53.
Aviv, A. and Shay, J.W., Reflections on telomere dynamics and ageing-related diseases in humans, Philos. Trans. R. Soc., B, 2018, vol. 373, art. ID 20 160 436.
Belancio, V.P., Roy-Engel, A.M., and Deininger, P.L., All you need to know about retroelements in cancer, Semin. Cancer Biol., 2010, vol. 20, no. 4, pp. 200–210.
Bertuch, A.A., The molecular genetics of the telomere biology disorders, RNA Biol., 2016, vol. 13, no. 8, pp. 696–706.
Biscotti, M.A., Canapa, A., Forconi, M., et al., Transcription of tandemly repetitive DNA: functional roles, Chromosome Res., 2015, vol. 23, pp. 463–477.
Blackburn, E.H. and Gall, J.G., A tandemly repeated sequence at the termini of the extrachromosomal ribosomal RNA genes in Tetrahymena,J. Mol. Biol., 1978, vol. 120, pp. 33–53.
Bolzan, A.D., Interstitial telomeric sequences in vertebrate chromosomes: origin, function, instability and evolution, Mutat. Res., 2017, vol. 3, pp. 51–65.
Calado, R.T. and Young, N.S., Telomere diseases, N. Engl. J. Med., 2009, vol. 361, no. 24, pp. 2353–2365.
Casacuberta, E., Drosophila: retrotransposons making up telomeres, Viruses, 2017, vol. 9, art. ID E192.
Cesare, A.J. and Reddel, R.R., Alternative lengthening of telomeres: models, mechanisms and implications, Nat. Rev. Genet., 2010, vol. 11, pp. 319–330.
Chan, F.L. and Wong, L.H., Transcription in the maintenance of centromere chromatin identity, Nucleic Acids Res., 2012, vol. 40, pp. 11 178–11 188.
Cheng, Z.J. and Murata, M., A centromeric tandem repeat family originating from a part of Ty3/gypsy-retroelement in wheat and its relatives, Genetics, 2003, vol. 164, pp. 665–672.
Cheong, C., Hong, K.U., and Lee, H.W., Mouse models for telomere and telomerase biology, Exp. Mol. Med., 2003, vol. 35, pp. 141–153.
Cherif, H., Tarry, J.L., Ozanne, S.E., and Hales, C.N., Ageing and telomeres: a study into organ- and gender-specific telomere shortening, Nucleic Acids Res., 2003, vol. 31, pp. 1576–1583.
Choi, J., Southworth, L.K., Sarin, K.Y., et al., TERT promotes epithelial proliferation through transcriptional control of a Myc- and Wnt-related developmental program, PLoS Genet., 2008, vol. 4, no. 1, p. e10.
Couzigou, J.M., Lauressergues, D., Becard, G., and Comier, J.P., miRNA-encoded peptides (miPEPs): A new tool to analyze the role of miRNAs in plant biology, RNA Biol., 2015, vol. 12, pp. 1178–1180.
de Sotero-Caio, C.G., Cabral-de-Mello, D.C., Calixto, M.D.S., et al., Centromeric enrichment of LINE-1 retrotransposons and its significance for the chromosome evolution of Phyllostomid bats, Chromosome Res., 2017, vol. 25, pp. 313–325.
Feschotte, C., Transposable elements and the evolution of regulatory networks, Nat. Rev. Genet., 2008, vol. 9, pp. 397–405.
Garavis, M., Gonzalez, C., and Villasante, A., On the origin of the eukaryotic chromosome: the role of noncanonical DNA structures in telomere evolution, Genome Biol. Evol., 2013, vol. 5, pp. 1142–1150.
Gent, J.I. and Dawe, R.K., RNA as a structural and regulatory component of the centromere, Annu. Rev. Genet., 2012, vol. 46, pp. 443–453.
Gill, Z., Nieuwoudt, M., and Ndifon, W., The Hayflick limit and age-related adaptive immune, Gerontology, 2018, vol. 64, pp. 135–139.
Gim, J., Ha, H., Ahn, K., et al., Genome-wide identification and classification of microRNAs derived from repetitive elements, Genomic Inf., 2014, vol. 12, no. 4, pp. 261–267.
Gladyshev, E.A. and Arkhipova, I.R., A subtelomeric non-LTR retrotansposon Hebe in the bdelloid rotifer Adineta vaga is subject to inactivation by deletions but not 5'truncations, Mobile DNA, 2010, vol. 1, p. 12.
Greider, C.W. and Blackburn, E.H., Identification of a specific telomere terminal transferase activity in Tetrahymena extracts, Cell, 1985, vol. 43, pp. 405–413.
Guttman, M. and Rinn, J.L., Modular regulatory principles of large non-coding RNAs, Nature, 2012, vol. 482, pp. 339–346.
Hara, T., Hirai, Y., Jahan, I., et al., Tandem repeat sequences evolutionarily related to SVA-type retrotransposons are expanded in centromere region of the western hoolock gibbon, a small ape, J. Hum. Genet., 2012, vol. 57, pp. 760–765.
Hastie, N.D., Dempster, M., Dunlop, M.G., et al., Telomere reduction in human colorectal carcinoma and with ageing, Nature, 1990, vol. 346, no. 6287, pp. 866–868.
Jankowska, M., Fuchs, J., Klocke, E., et al., Holokinetic centromeres and efficient telomere healing enable rapid karyotype evolution, Chromosoma, 2015, vol. 124, pp. 519–528.
Johnson, R. and Guigo, R., The RIDL hypothesis: transposable elements as functional domains of long noncoding RNAs, RNA, 2014, vol. 20, no. 7, pp. 959–976.
Joly-Lopez, Z. and Bureau, T.E., Exaptation of transposable element coding sequences, Curr. Opin. Genet. Dev., 2018, vol. 49, pp. 34–42.
Klutstein, M., Fennell, A., Farnandez-Alvarez, A., and Cooper, J.P., The telomere bouquet regulates meiotic centromere assembly, Nat. Cell. Biol., 2015, vol. 17, pp. 458–469.
Kordyukova, M., Olovnikov, I., and Kalmykova, A., Transposon control mechanisms in telomere biology, Curr. Opin. Genet. Dev., 2018, vol. 49, pp. 56–62.
Kubiak, M.R. and Makalowska, I., Protein-coding genes’ retrocopies and their functions, Viruses, 2017, vol. 9, art. ID E80.
Kulski, J.K., Anzai, T., and Inoko, H., ERVK9, transposons and the evolution of MHC class I duplicons within the alpha-block of the human and chimpanzee, Cytogenet. Genome Res., 2005, vol. 110, pp. 181–192.
Lauressergues, D., Couzigou, J.M., Clemente, H.S., et al., Primary transcripts of microRNAs encode regulatory peptides, Nature, 2015, vol. 520, no. 7545, pp. 90–93.
Lewis, K.A. and Wuttke, D.S., Telomerase and telomere-associated proteins: structural insights into mechanism and evolution, Structure, 2012, vol. 20, no. 1, pp. 28–39.
Lieberman, P.M., Retrotransposon-derived p53 binding sites enhance telomere maintenance and genome protection, BioEssays, 2016, vol. 38, pp. 943–949.
Linardopoulou, E.V., Parghi, S.S., Friedman, C., et al., Human subtelomeric WASH genes encode a new subclass of the WASP family, PLoS Genet., 2007, vol. 3, p. e237.
Lu, D., Davis, M.P., Abreu-Goodger, C., et al., MiR-25 regulates Wwp2 and Fbxw7 and promotes reprogramming of mouse fibroblast cells to iPSCs, PLoS One, 2012, vol. 7, no. 8, p. e40938.
Lv, S., Pan, L., and Wang, G., Commentary: primary transcripts of microRNAs encode regulatory peptides, Front. Plant Sci., 2016, vol. 7, p. 1436.
McGurk, M.P. and Barbash, D.A., Double insertion of transposable elements provides a substrate for the evolution of satellite DNA, Genome Res., 2018, vol. 28, no. 5, pp. 714–725.
Maeda, T., Horiuchi, T., and Makino, N., Epigenetic status of subtelomere of peripheral leukocytes corresponds to cardiographic parameters with a sex association, Geriatr. Gerontol. Int., 2018, vol. 18, pp. 1415–1419.
Mestrovic, N., Mravinac, B., Pavlek, M., et al., Structural and functional liaisons between transposable elements and satellite DNAs, Chromosome Res., 2015, vol. 23, no. 3, pp. 583–596.
Mueller, C., Aschacher, T., Wolf, B., and Bergmann, M., A role of LINE-1 in telomere regulation, Front. Biosci., 2018, vol. 23, pp. 1310–1319.
Piqueret-Stephan, L., Ricoul, M., Hempel, W.M., and Sabatier, L., Replication timing of human telomeres is conserved during immortalization and influenced by respective subtelomeres, Sci. Rep., 2016, vol. 6, p. 32510.
Qu, G., Kaushal, P.S., Wang, J., et al., Structure of a group II intron in complex with its reverse transcriptase, Nat. Struct. Mol. Biol., 2016, vol. 23, no. 6, pp. 549–557.
Riethman, H., Ambrosini, A., and Paul, S., Human subtelomere structure and variation, Chromosome Res., 2005, vol. 13, pp. 505–515.
Rosen, M., Castillejo-Lopez, C., and Edstrom, J.E., Telomere terminating with centromere-specific repeats is closely associated with a transposon derived gene in Chronomus pallidivittatus,Chromosoma, 2002, vol. 110, no. 8, pp. 532–541.
Rubtsova, M., Narayking, Y., Vasilkova, D., et al., Protein encoded in human telomerase RNA is involved in cell protective pathways, Nucleic Acids Res., 2018, vol. 46, pp. 8966–8977.
Sarin, K.Y., Cheung, P., Gilison, D., et al., Conditional telomerase induction causes proliferation of hair follicle stem cells, Nature, 2005, vol. 436, no. 7053, pp. 1048–1052.
Sharma, A., Wolfgruber, T.K., and Presting, G.G., Tandem repeats derived from centromeric retrotransposons, BMC Genomics, 2013, vol. 14, p. 142.
Slijepcevic, P., Mechanisms of the evolutionary chromosome plasticity: integrating the ‘centromere-from-telomere’ hypothesis with telomere length regulation, Cytogenet. Genome Res., 2016, vol. 148, pp. 268–278.
Szostak, J.W. and Blackburn, E.H., Cloning yeast telomeres on linear plasmid vectors, Cell, 1982, vol. 29, no. 1, pp. 245–255.
Tan, S., Cardoso-Moreira, M., Shi, W., et al., LTR-mediated retroposition as a mechanism of RNA-based duplication in metazoans, Genome Res., 2016, vol. 26, no. 12, pp. 1663–1675.
Tutton, S., Azzam, G.A., Stong, N., et al., Subtelomeric p53 binding prevents accumulation of DNA damage at human telomeres, EMBO J., 2016, vol. 35, pp. 193–207.
Villasante, A., Abad, J.P., and Mendez-Logo, M., Centromeres were derived from telomeres during the evolution of the eukaryotic chromosome, Proc. Natl. Acad. Sci. U.S.A., 2007, vol. 104, pp. 10 542–10 547.
Volff, J.N., Turning junk into gold: domestication of transposable elements and the creation of new genes in eukaryotes, BioEssays, 2006, vol. 28, pp. 913–922.
Xiong, W., Dooner, H.K., and Du, C., Rolling-circle amplification of centromeric Helitrons in plant genomes, The Plant. J., 2016, vol. 88, pp. 1038–1045.
Zhong, C.X., Marshall, J.B., Topp, C., et al., Centromeric retroelements and satellites interact with maize kinetochore protein CENH3, Plant Cell, 2002, vol. 14, no. 11, pp. 2825–2836.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest. The author declares that there is no conflict of interest.
Statement on animal welfare. This article does not contain any studies involving animals performed by any of the authors.
Statement of compliance with standards of research involving humans as subjects. This article does not contain any studies involving humans as subjects of research.
Additional information
Translated by D. Novikova
Rights and permissions
About this article
Cite this article
Mustafin, R.N. Interrelation of Telomeres with Transposable Elements in Aging. Adv Gerontol 10, 101–108 (2020). https://doi.org/10.1134/S2079057020020113
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S2079057020020113