Abstract—
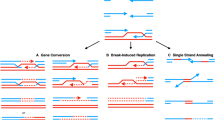
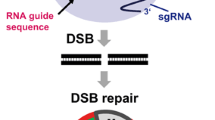
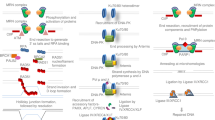
Genome editing is a powerful tool that allows study of the properties of genes or changes to be made to the genetic sequence. Programmable nucleases that can induce double-strand breaks in the genomic sequence of interest have been developed over the past few decades. After initiation of a double-strand break (DSB) in DNA, the DSB can be repaired by the NHEJ (non-homologous end joining), which leads to various errors and gene knockout. Other repair options, HDR (homology directed repair) or SSTR (single-strand template repair), allow researchers to make desired changes in the gene. HDR occurs in the presence of a donor template, in natural conditions the donor template is a sister chromatid. The efficiency of HDR and SSTR is significantly lower than the efficiency of NHEJ in genome editing. Double-stranded, single-stranded and long single-stranded DNAs are used to increase efficiency and to make desired changes in genomic DNA. In this review, we discuss donor molecules that are used for DSB repair using HDR or SSTR during genome editing, their application, and modifications to increase the efficiency of HDR and SSTR.





Similar content being viewed by others
REFERENCES
Durai S., Mani M., Kandavelou K., Wu J., Porteus M.H., Chandrasegaran S. 2005. Zinc finger nucleases: Custom-designed molecular scissors for genome engineering of plant and mammalian cells. Nucleic Acids Res. 33 (18), 3978–5990. https://doi.org/10.1093/nar/gki912
Silva G., Poirot L., Galetto R., Smith J., Montoya G., Duchateau Ph., Pâques F. 2011. Meganucleases and other tools for targeted genome engineering: Perspectives and challenges for gene therapy. Curr. Gene Therapy. 11 (1), 11–27. https://doi.org/10.2174/156652311794520111
Cermak T., Doyle E. L., Christian M., Wang L., Zhang Y., Schmidt C., Baller J.A., Somia N.V., Bogdanove A.J., Voytas D.F. 2011. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39 (12), e82. https://doi.org/10.1093/nar/gkr218
Jinek M., Chylinski K., Fonfara I., Hauer M., Doudna J.A., Charpentier E. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 337 (6096), 816–821. https://doi.org/10.1126/science.1225829
Guha T.K., Wai A., Hausner G. 2017. Programmable genome editing tools and their regulation for efficient genome engineering. Comp. Struct. Biotechnol. J. 15, 146–160. https://doi.org/10.1016/j.csbj.2016.12.006
Symington L.S., Gautier J. 2011. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 45, 247–271. https://doi.org/10.1146/annurev-genet-110410-132435
Chang H., Pannunzio N.R., Adachi N., Lieber M.R. 2017. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat. Rev. Mol. Cell Biol. 18 (8), 495–506. https://doi.org/10.1038/nrm.2017.48
Li T., Huang S., Zhao X., Wright D.A., Carpenter S., Spalding M.H., Weeks D.P., Yang B. 2011. Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes. Nucleic Acids Res. 39 (14), 6315–6325. https://doi.org/10.1093/nar/gkr188
Devkota S. 2018. The road less traveled: Strategies to enhance the frequency of homology-directed repair (HDR) for increased efficiency of CRISPR/Casmediated transgenesis. BMB Rep. 51 (19), 437–443. https://doi.org/10.5483/BMBRep.2018.51.9.187
Richardson C.D., Kazane K.R., Feng S.J., Zelin E., Bray N.L., Schäfer A.J., Floor S.N., Corn J.E. 2018. CRISPR-Cas9 genome editing in human cells occurs via the Fanconi anemia pathway. Nat. Genet. 50 (8), 1132–1139. https://doi.org/10.1038/s41588-018-0174-0
Hockemeyer D., Soldner F., Beard C., Gao Q., Mitalipova M., DeKelver R.C., Katibah G.E., Amora R., Boydston E.A., Zeitler B., Meng X., Miller J.C., Zhang L., Rebar E.J., Gregory Ph.D., et al. 2009. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat. Biotechnol. 27, 851–857. https://doi.org/10.1038/nbt.1562
Hockemeyer D., Wang H., Kiani S., Lai C.S., Gao Q., Cassady J.P., Cost G.J., Zhang L., Santiago Y., Miller J.C., Zeitler B., Cherone J.M., Meng X., Hinkley S.J., Rebar E.J., et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat. Biotechnol. 29, 731–734. https://doi.org/10.1038/nbt.1927
Sommer D., Peters A., Wirtz T., Mai M., Ackermann J., Thabet Y., Schmidt J., Weighardt H., Wunderlich F.T., Degen J., Schultze J.L., Beyer M. 2014. Efficient genome engineering by targeted homologous recombination in mouse embryos using transcription activator-like effector nucleases. Nat. Commun. 5, 3045. https://doi.org/10.1038/ncomms4045
Heyer W.D., Ehmsen K.T., Liu J. 2010. Regulation of homologous recombination in eukaryotes. Annu. Rev. Genet. 44, 113–139. https://doi.org/10.1146/annurev-genet-051710-150955
Yanik M., Ponnam S., Wimmer T., Trimborn L., Müller C., Gambert I., Ginsberg J., Janise A., Domicke J., Wende W., Lorenz B., Stieger K. 2018. Development of a reporter system to explore MMEJ in the context of replacing large genomic fragments. Mol. Therapy. Nucl. Acids. 11, 407–415. https://doi.org/10.1016/j.omtn.2018.03.010
Smirnikhina S.A., Anuchina A.A., Lavrov A.V. 2019. Ways of improving precise knock-in by genome-editing technologies. Hum. Genet. 138 (1), 1–19. https://doi.org/10.1007/s00439-018-1953-5
Lin S., Staahl B.T., Alla R.K., Doudna J.A. 2014. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife. 3, e04766. https://doi.org/10.7554/eLife.04766
Seol J.H., Shim E.Y., Lee S.E. 2018. Microhomology-mediated end joining: Good, bad and ugly. Mutat. Res. 809, 81–87.
González-Marín C., Gosálvez J., Roy R. 2012. Types, causes, detection and repair of DNA fragmentation in animal and human sperm cells. Internat. J. Mol. Sci. 13 (11), 14026–14052. https://doi.org/10.3390/ijms131114026
Liu M., Rehman S., Tang X., Fan Q., Chen D., Ma W. 2019. Methodologies for improving HDR efficiency. Front. Genet. 9 (691). https://doi.org/10.3389/fgene.2018.00691
Zhu Z., Chung W.H., Shim E.Y, Lee S.E., Ira G. 2008. Sgs1 helicase and two nucleases Dna2 and Exo1 resect DNA double-strand break ends. Cell. 34 (6), 981‒994. https://doi.org/10.1016/j.cell.2008.08.037
Falck J., Coates J., Jackson S.P. 2005. Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature. 434 (7033), 605‒611. https://doi.org/10.1038/nature03442
Dupré A., Boyer-Chatenet L., Gautier J. 2006. Two-step activation of ATM by DNA and the Mre11-Rad50-Nbs1 complex. Nat. Struct. Mol. Biol. 13 (5), 451–457. https://doi.org/10.1038/nsmb1090
Helleday T., Lo J., van Gent D.C., Engelward B.P. 2007. DNA double-strand break repair: From mechanistic understanding to cancer treatment. DNA Repair. 6(7), 923–935. https://doi.org/10.1016/j.dnarep.2007.02.006
Liu J., Ede C., Wright W. D., Gore S.K., Jenkins S.S., Freudenthal B.D., Washington M.T., Veaute X., Heyer W.-D. 2017. Srs2 promotes synthesis-dependent strand annealing by disrupting DNA polymerase δ-extending D-loops. eLife. 6, e22195. https://doi.org/10.7554/eLife.22195
McMahill M.S., Sham C.W., Bishop D.K. 2007. Synthesis-dependent strand annealing in meiosis. PLoS Biol. 5 (11), e299. https://doi.org/10.1371/journal.pbio.0050299
Agmon N., Yovel M., Harari Y., Liefshitz B., Kupiec M. 2011. The role of Holliday junction resolvases in the repair of spontaneous and induced DNA damage. Nucleic Acids Res. 39 (16), 7009–7019. https://doi.org/10.1093/nar/gkr277
Davis L., Zhang Y., Maizels N. 2018. Assaying repair at DNA nicks. Methods Enzymol. 601, 71–89. https://doi.org/10.1016/bs.mie.2017.12.001
Gallagher D.N., Pham N., Tsai A.M., Janto A.N., Choi J., Ira G., Haber J.E. 2020. A Rad51-independent pathway promotes single-strand template repair in gene editing. PLoS Genet. 16 (10), e1008689. https://doi.org/10.1371/journal.pgen.1008689
Gallagher D.N., Haber J.E. 2021. Single-strand template repair: Key insights to increase the efficiency of gene editing. Curr. Genet. 67, 747–753. https://doi.org/10.1007/s00294-021-01186-z
Hu Z., Zhou M., Wu Y., Li Z., Liu X., Wu L., Liang D. 2019. ssODN-mediated in-frame deletion with CRISPR/Cas9 restores FVIII function in hemophilia A‑patient-derived iPSCs and ECs. Mol. Therapy. Nucl. Acids. 17, 198–209. https://doi.org/10.1016/j.omtn.2019.05.019
Bennett H., Aguilar-Martinez E., Adamson A.D. 2021. CRISPR-mediated knock-in in the mouse embryo using long single stranded DNA donors synthesised by biotinylated PCR. Methods (San Diego, CA). 191, 3–14. https://doi.org/10.1016/j.ymeth.2020.10.012
Cristea S., Freyvert Y., Santiago Y. 2013. In vivo cleavage of transgene donors promotes nuclease-mediated targeted integration. Biotechnol. Bioengin. 110 (3), 871–880. https://doi.org/10.1002/bit.24733
Fueller J., Herbst K., Meurer M., Gubicza K., Kurtulmus B., Knopf J.D., Kirrmaier D., Buchmuller B.C., Pereira G., Lemberg M.K., Knop M. 2020. CRISPR-Cas12a-assisted PCR tagging of mammalian genes. J. Cell Biol. 219 (6), e201910210. https://doi.org/10.1083/jcb.201910210
Chen F., Pruett-Miller S.M., Davis G.D. 2015. Gene editing using ssODNs with engineered endonucleases. Methods Mol. Biol. 1239, 251–265. https://doi.org/10.1007/978-1-4939-1862-1_14
Myagkaya N.O., Marochkin N.A., Norbobaeva M.B., Serba T.V., Starodub M.A., Kurbedinov R.A., Kostenko S.A., Zayats I.V. 2020. A review of the CRISPR/Cas9 method for genome editing. Molodoi Uchenyi. 12 (302), 81–85.
Designing Homologous Repair Templates. New England BioLabs Inc. https://international.neb.com/applications/genome-editing/designing-homologous-repair-templates.
Mao C.Z., Zheng L., Zhou Y.M., Wu H.-Y., Xia J.-B., Liang C.-Q., Guo X.-F., Peng W.-T., Zhao H., Cai W.-B., Kim S.-K., Park K.-S., Cai D.-Q., Qi X.-F. 2018. CRISPR/Cas9-mediated efficient and precise targeted integration of donor DNA harboring double cleavage sites in Xenopus tropicalis. FASEB J. fj201800093. https://doi.org/10.1096/fj.201800093
Pineault K.M., Novoa A., Lozovska A., Wellik D.M., Mallo M. 2019. Two CRISPR/Cas9-mediated methods for targeting complex insertions, deletions, or replacements in mouse. Methods X. 6, 2088–2100. https://doi.org/10.1016/j.mex.2019.09.003
Roth T.L., Puig-Saus C., Yu R., Shifrut E., Carnevale J., Li P. J., Hiatt J., Saco J., Krystofinski P., Li H., Tobin V., Nguyen D.N., Lee M.R., Putnam A.L., Ferris A.L., Chen J.W., et al. 2018. Reprogramming human T cell function and specificity with non-viral genome targeting. Nature. 559 (7714), 405–409. https://doi.org/10.1038/s41586-018-0326-5
Quadros R.M., Miura H., Harms D.W., Akatsuka H., Sato T., Aida T., Redder R., Richardson G.P., Inagaki Y., Sakai D., Buckley S.M., Seshacharyulu P., Batra S.K., Behlke M.A., Zeiner S.A., et al. 2017. Easi-CRISPR: A robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biol. 18 (1), 92. https://doi.org/10.1186/s13059-017-1220-4
Miura H., Gurumurthy C.B., Sato T., Sato M., Ohtsuka M. 2015. CRISPR/Cas9-based generation of knockdown mice by intronic insertion of artificial microRNA using longer single-stranded DNA. Sci. Rep. 5, 12799. https://doi.org/10.1038/srep12799
Leonetti M.D., Sekine S., Kamiyama D., Weissman J.S., Huang B. 2016. A scalable strategy for high-throughput GFP tagging of endogenous human proteins. Proc. Natl. Acad. Sci. U. S. A. 113 (25), E3501–E3508. https://doi.org/10.1073/pnas.1606731113
Miura H., Quadros R.M., Gurumurthy C.B., Ohtsuka M. 2018. Easi-CRISPR for creating knock-in and conditional knockout mouse models using long ssDNA donors. Nat. Protocols. 13 (1), 195–215. https://doi.org/10.1038/nprot.2017.153
Harrison G.P., Mayo M.S., Hunter E., Lever A.M. 1998. Pausing of reverse transcriptase on retroviral RNA templates is influenced by secondary structures both 5' and 3' of the catalytic site. Nucleic Acids Res. 26 (14), 3433–3442. https://doi.org/10.1093/nar/26.14.3433
Bai H., Liu L., An K., X. Lu, Harrison M., Zhao Y., Yan R., Lu Z., Li S., Lin S., Liang F., Qin W. 2020. CRISPR/Cas9-mediated precise genome modification by a long ssDNA template in zebrafish. BMC Genomics. 21 (1), 67. https://doi.org/10.1186/s12864-020-6493-4
Hwang W.Y., Fu Y., Reyon D., Maeder M.L., Kaini P., Sander J.D., Joung J.K., Peterson R.T., Yeh J.-R.J. 2013. Heritable and precise zebrafish genome editing using a CRISPR-Cas system. PLoS One. 8 (7), e68708 https://doi.org/10.1371/journal.pone.0068708
Richardson C.D., Ray G.J., DeWitt M.A., Curie G.L., Corn J.E. 2016. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat. Biotechnol. 34 (3), 339–344. https://doi.org/10.1038/nbt.3481
O'Brien A.R., Wilson L., Burgio G., Bauer D.C. 2019. Unlocking HDR-mediated nucleotide editing by identifying high-efficiency target sites using machine learning. Sci. Rep. 9 (1), 2788. https://doi.org/10.1038/s41598-019-39142-0
Lim D., Sreekanth V., Cox K.J., Law B.K., Wagner B.K., Karp J.M., Choudhary A. 2020. Engineering designer beta cells with a CRISPR-Cas9 conjugation platform. Nat. Commun. 11 (1), 4043. https://doi.org/10.1038/s41467-020-17725-0
Harmsen T., Klaasen S., van de Vrugt H., Te Riele H. 2018. DNA mismatch repair and oligonucleotide end-protection promote base-pair substitution distal from a CRISPR/Cas9-induced DNA break. Nucleic Acids Res. 46 (6), 2945–2955. https://doi.org/10.1093/nar/gky076
Shao Y., Guan Y., Wang L., Qiu Z., Liu M., Chen Y., Wu L., Li Y., Ma X., Liu M., Li D. 2014. CRISPR/Cas-mediated genome editing in the rat via direct injection of one-cell embryos. Nat. Protocols. 9 (10), 2493–2512. https://doi.org/10.1038/nprot.2014.171
Irion U., Krauss J., Nüsslein-Volhard C. 2014. Precise and efficient genome editing in zebrafish using the CRISPR/Cas9 system. Development. 141 (24), 4827–4830. https://doi.org/10.1242/dev.115584
Wang B., Karaçay B. Megamer single-stranded donor templates (ssDNA or ssODNs) for successful homology-directed repair (HDR) in genome editing applications. https://eu.idtdna.com/pages/education/decoded/ article/megamer-single-stranded-donor-templates-(ssdna-or-ssODNs)-for-successful-homology-directed-repair-(HDR)-in-genome-editing-applications.
Yoshimi K., Oka Y., Miyasaka Y., Kotani Y., Yasumura M., Uno Y., Hattori K., Tanigawa A., Sato M., Oya M., Nakamura K., Matsushita N., Kobayashi K., Mashimo T. 2021. Combi-CRISPR: Combination of NHEJ and HDR provides efficient and precise plasmid-based knock-ins in mice and rats. Hum. Genet. 140 (2), 277–287. https://doi.org/10.1007/s00439-020-02198-4
Highly Specific Gene Knockins of Long Sequences Using CRISPR/Cas9 and a Single-stranded DNA Donor Template. Takara Bio USA, Inc. https://catalog.takara-bio.co.jp/PDFS/Highly-specific-gene-knockins-of-long-sequences-using-CRISPR_Cas9-and-a-single-stranded-DNA-donor-template.pdf.
Zhao Y., Zheng Z., Cohen C.J., Gattinoni L., Pal-mer D.C., Restifo N.P., Rosenberg S.A., Morgan R.A. 2006. High-efficiency transfection of primary human and mouse T lymphocytes using RNA electroporation. Mol. Ther: J. Am. Soc. Gene Ther. 13 (1), 151–159. https://doi.org/10.1016/j.ymthe.2005.07.688
Mello C.C., Kramer J.M., Stinchcomb D., Ambros V. 1991. Efficient gene transfer in C. elegans: Extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10 (12), 3959–3970.
Forbes D.J., Kirschner M.W., Newport J.W. 1983. Spontaneous formation of nucleus-like structures around bacteriophage DNA microinjected into Xenopus eggs. Cell. 34 (1), 13–23. https://doi.org/10.1016/0092-8674(83)90132-0
Stinchcomb D.T., Shaw J.E., Carr S.H., Hirsh D. 1985. Extrachromosomal DNA transformation of Caenorhabditis elegans. Mol. Cell. Biol. 5 (12), 3484–3496. https://doi.org/10.1128/mcb.5.12.3484-3496.1985
Ghanta K.S., Mello C.C. 2020. Melting dsDNA donor molecules greatly improves precision genome editing in Caenorhabditis elegans. Genetics. 216 (3), 643–650. https://doi.org/10.1534/genetics.120.303564
Levi T., Sloutskin A., Kalifa R., Juven-Gershon T., Gerlitz O. 2020. Efficient in vivo introduction of point mutations using ssODN and a Co-CRISPR approach. Biol. Proc. Online. 22, 14. https://doi.org/10.1186/s12575-020-00123-7
Boel A., De Saffel H., Steyaert W., Callewaert B., De Paepe A., Coucke P.J., Willaert A. 2018. CRISPR/ Cas9-mediated homology-directed repair by ssODNs in zebrafish induces complex mutational patterns resulting from genomic integration of repair-template fragments. Dis. Models Mech. 11 (10), dmm035352. https://doi.org/10.1242/dmm.035352
Gratz S.J., Cummings A.M., Nguyen J.N., Hamm D.C., Donohue L.K., Harrison M.M., Wildonger J., O’Connor-Giles K.M. 2013. Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics. 194 (4), 1029–1035. https://doi.org/10.1534/genetics.113.152710
Wang H., Yang H., Shivalila C.S., Dawlaty M.M., Cheng A.W., Zhang F., Jaenisch R. 2013. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell. 153 (4), 910–918. https://doi.org/10.1016/j.cell.2013.04.025
Guide-it Long ssDNA Production System v. 2. Takara Bio USA, Inc. https://www.takarabio.com/products/ gene-function/gene-editing/crispr-cas9/long-ssdna-for-knockins.
Bassett A.R., Tibbit C., Ponting C.P., Liu J.L. 2013. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep. 4 (1), 220–228. https://doi.org/10.1016/j.celrep.2013.06.020
Codner G.F., Mianné J., Caulder A., Loeffler J., Fell R., King R., Allan A.J., Mackenzie M., Pike F.J., McCabe C.V., Christou S., Joynson S., Hutchison M., Stewart M.E., Kumar S, et al. 2018. Application of long single-stranded DNA donors in genome editing: Generation and validation of mouse mutants. BMC Biol. 16 (1), 70. https://doi.org/10.1186/s12915-018-0530-7
Yoshimi K., Kunihiro Y., Kaneko T. 2016. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat. Commun. 7, 10431. https://doi.org/10.1038/ncomms10431
Ranawakage D.C., Okada K., Sugio K., Kawaguchi Y., Kuninobu-Bonkohara Y., Takada T., Kamachi Y. 2021. Efficient CRISPR-Cas9-mediated knock-in of composite tags in zebrafish using long ssDNA as a donor. Front. Cell Dev. Biol. 8, 598634. https://doi.org/10.3389/fcell.2020.598634
Funding
This study was conducted within the state assignment set out by the Ministry of Science and Higher Education of the Russian Federation for the Research Centre for Medical Genetics.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by M. Novikova
Abbreviations: ZFN, zinc finger nuclease; TALEN, transcription activator-like effector nuclease; CRISPR, clustered regularly interspaced short palindromic repeats; PAM, protospacer adjacent motif; a-NHEJ, alternative non-homologous end joining; lssDNA, long single-strand DNA; dsDNA, double-strand DNA; DSB, double-strand break; с-NHEJ, classic non-homologous end joining; MMEJ, microhomology-mediated end joining; HDR, homology directed repair; NHEJ, non-homologous end joining; ssODNs, single-strand oligodeoxyribonucleotides; SSTR, single-strand template repair; SDSA, synthesis-dependent strand annealing.
Rights and permissions
About this article
Cite this article
Volodina, O.V., Smirnikhina, S.A. The Choice of a Donor Molecule in Genome Editing Experiments in Animal Cells. Mol Biol 56, 372–381 (2022). https://doi.org/10.1134/S002689332203013X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S002689332203013X