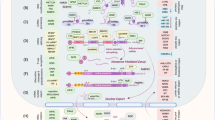
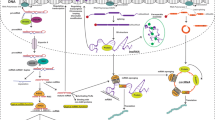
Abstract
RNA interference has become an indispensable research tool to study gene functions in a wide variety of organisms. Because of their high efficacy and specificity, RNA interference-based approaches may also translate into new therapeutic strategies to treat human diseases. In particular, oncogenes such as leukemic fusion proteins, which arise from chromosomal translocations, are promising targets for such gene silencing approaches, because they are exclusively expressed in pre-cancerous and cancerous tissues, and because they are frequently indispensable for maintaining the malignant phenotype. This review summarizes recent developments in targeting leukemia-specific genes and discusses problems and approaches for possible clinical applications.
Similar content being viewed by others
Article PDF
References
Novina CD, Sharp PA . The RNAi revolution. Nature 2004; 430: 161–4.
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC . Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998; 391: 806–11.
Huppi K, Martin SE, Caplen NJ . Defining and assaying RNAi in mammalian cells. Mol Cell 2005; 17: 1–10.
Tuschl T, Zamore PD, Lehmann R, Bartel DP, Sharp PA . Targeted mRNA degradation by double-stranded RNA in vitro. Genes Dev 1999; 13: 3191–7.
Caplen NJ . Gene therapy progress and prospects. Downregulating gene expression: the impact of RNA interference. Gene Ther 2004; 11: 1241–8.
Cheng JC, Moore TB, Sakamoto KM . RNA interference and human disease. Mol Genet Metab 2003; 80: 121–8.
Borkhardt A, Heidenreich O . RNA interference as a potential tool in the treatment of leukaemia. Expert Opin Biol Ther 2004; 4: 1921–9.
Insinga A, Monestiroli S, Ronzoni S, Gelmetti V, Marchesi F, Viale A, et al. Inhibitors of histone deacetylases induce tumor-selective apoptosis through activation of the death receptor pathway. Nat Med 2005; 11: 71–6.
Claus R, Almstedt M, Lubbert M . Epigenetic treatment of hematopoietic malignancies: in vivo targets of demethylating agents. Semin Oncol 2005; 32: 511–20.
Elbashir SM, Lendeckel W, Tuschl T . RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev 2001; 15: 188–200.
Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K, et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005; 436: 740–4.
Meister G, Landthaler M, Patkaniowska A, Dorsett Y, Teng G, Tuschl T . Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol Cell 2004; 15: 185–97.
Okamura K, Ishizuka A, Siomi H, Siomi MC . Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev 2004; 18: 1655–6.
Ambros V . The functions of animal microRNAs. Nature 2004; 431: 350–5.
Hutvagner G, Zamore PD . A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002; 297: 2056–60.
Zeng Y, Cullen BR . RNA interference in human cells is restricted to the cytoplasm. RNA 2002; 8: 855–60.
Liu J, Valencia-Sanchez MA, Hannon GJ, Parker R . MicroRNA-dependent localization of targeted mRNAs to mammalian P-bodies. Nat Cell Biol 2005; 7: 719–23.
Sen GL, Blau HM . Argonaute 2/RISC resides in sites of mammalian mRNA decay known as cytoplasmic bodies. Nat Cell Biol 2005; 7: 633–6.
Thomas M, Gessner A, Vornlocher HP, Hadwiger P, Greil J, Heidenreich O . Targeting MLL-AF4 with short interfering RNAs inhibits clonogenicity and engraftment of t(4;1 1)-positive human leukemic cells. Blood 2005; 106: 3559–66.
Ui-Tei K, Naito Y, Takahashi F, Haraguchi T, Ohki-Hamazaki H, Juni A, et al. Guidelines for the selection of highly effective siRNA sequences for mammalian and chick RNA interference. Nucleic Acids Res 2004; 32: 936–48.
Fedorov Y, King A, Anderson E, Karpilow J, Ilsley D, Marshall W, et al. Different delivery methods-different expression profiles. Nat Methods 2005; 2: 241.
Donze O, Picard D . RNA interference in mammalian cells using siRNAs synthesized with T7 RNA polymerase. Nucleic Acids Res 2002; 30: e46.
Brummelkamp TR, Bernards R, Agami R . Stable suppression of tumorigenicity by virus-mediated RNA interference. Cancer Cell 2002; 2: 243–7.
Abbas-Terki T, Blanco-Bose W, Deglon N, Pralong W, Aebischer P . Lentiviral-mediated RNA interference. Hum Gene Ther 2002; 13: 2197–201.
Druker BJ, Talpaz M, Resta DJ, Peng B, Buchdunger E, Ford JM, et al. Efficacy and safety of a specific inhibitor of the BCR-ABL tyrosine kinase in chronic myeloid leukemia. N Engl J Med 2001; 344: 1031–7.
Barthe C, Gharbi MJ, Lagarde V, Chollet C, Cony-Makhoul P, Reiffers J, et al. Mutation in the ATP-binding site of BCR-ABL in a patient with chronic myeloid leukaemia with increasing resistance to STI571. Br J Haematol 2002; 119: 109–11.
Barthe C, Cony-Makhoul P, Melo JV, Mahon JR . Roots of clinical resistance to STI-571 cancer therapy. Science 2001; 293: 2163.
Skorski T, Nieborowska-Skorska M, Nicolaides NC, Szczylik C, Iversen P, Iozzo RV, et al. Suppression of Philadelphia 1 leukemia cell growth in mice by BCR-ABL antisense oligodeoxynucleotide. Proc Natl Acad Sci USA 1994; 91: 4504–8.
Wilda M, Fuchs U, Wossmann W, Borkhardt A . Killing of leukemic cells with a BCR/ABL fusion gene by RNA interference (RNAi). Oncogene 2002; 21: 5716–24.
Scherr M, Battmer K, Winkler T, Heidenreich O, Ganser A, Eder M . Specific inhibition of bcr-abl gene expression by small interfering RNA. Blood 2003; 101: 1566–9.
Rapozzi V, Xodo LE . Efficient silencing of bcr/abl oncogene by single- and double-stranded siRNAs targeted against b2a2 transcripts. Biochemistry 2004; 43: 16134–41.
Li MJ, McMahon R, Snyder DS, Yee JK, Rossi JJ . Specific killing of Ph+ chronic myeloid leukemia cells by a lentiviral vector-delivered anti-bcr/abl small hairpin RNA. Oligonucleotides 2003; 13: 401–9.
Scherr M, Battmer K, Schultheis B, Ganser A, Eder M . Stable RNA interference (RNAi) as an option for anti-bcr-abl therapy. Gene Ther 2005; 12: 12–21.
Withey JM, Marley SB, Kaeda J, Harvey AJ, Crompton MR, Gordon MY . Targeting primary human leukaemia cells with RNA interference: Bcr-Abl targeting inhibits myeloid progenitor self-renewal in chronic myeloid leukaemia cells. Br J Haematol 2005; 129: 377–80.
Zhelev Z, Bakalova R, Ohba H, Ewis A, Ishikawa M, Shinohara Y, et al. Suppression of bcr-abl synthesis by siRNAs or tyrosine kinase activity by Glivec alters different oncogenes, apoptotic/ antiapoptotic genes and cell proliferation factors (microarray study). FEBS Lett 2004; 570: 195–204.
Ohba H, Zhelev Z, Bakalova R, Ewis A, Omori T, Ishikawa M, et al. Inhibition of bcr-abl and/or c-abl gene expression by small interfering, double-stranded RNAs: cross-talk with cell proliferation factors and other oncogenes. Cancer 2004; 101: 1390–403.
Bartova E, Harnicarova A, Pachernik J, Kozubek S . Nuclear topography and expression of the BCR/ABL fusion gene and its protein level influenced by cell differentiation and RNA interference. Leuk Res 2005; 29: 901–13.
Wohlbold L, van der Kuip H, Miething C, Vornlocher HP, Knabbe C, Duyster J, et al. Inhibition of bcr-abl gene expression by small interfering RNA sensitizes for imatinib mesylate (STI571). Blood 2003; 102: 2236–9.
Golub TR, Barker GF, Lovett M, Gilliland DG . Fusion of PDGF receptor beta to a novel ets-like gene, tel, in chronic myelomono-cytic leukemia with t(5;12) chromosomal translocation. Cell 1994; 77: 307–16.
Chen J, Wall NR, Kocher K, Duclos N, Fabbro D, Neuberg D, et al. Stable expression of small interfering RNA sensitizes TEL-PDGFbetaR to inhibition with imatinib or rapamycin. J Clin Invest 2004; 113: 1784–91.
Ritter U, Damm-Welk C, Fuchs U, Bohle RM, Borkhardt A, Woessmann W . Design and evaluation of chemically synthesized siRNA targeting the NPM-ALK fusion site in anaplastic large cell lymphoma (ALCL). Oligonucleotides 2003; 13: 365–73.
Walters DK, Stoffregen EP, Heinrich MC, Deininger MW, Druker BJ . RNAi-induced down-regulation of FLT3 expression in AML cell lines increases sensitivity to MLN518. Blood 2005; 105: 2952–4.
Ptasznik A, Nakata Y, Kalota A, Emerson SG, Gewirtz AM . Short interfering RNA (siRNA) targeting the Lyn kinase induces apoptosis in primary, and drug-resistant, BCR-ABL1(+) leukemia cells. Nat Med 2004; 10: 1187–9.
Downing JR . The AML1-ETO chimaeric transcription factor in acute myeloid leukaemia: biology and clinical significance. Br J Haematol 1999; 106: 296–308.
Westendorf JJ, Yamamoto CM, Lenny N, Downing JR, Selsted ME, Hiebert SW . The t(8;21) fusion product, AML-1-ETO, associates with C/EBP-alpha, inhibits C/EBP-alpha-dependent transcription, and blocks granulocytic differentiation. Mol Cell Biol 1998; 18: 322–33.
Heidenreich O, Krauter J, Riehle H, Hadwiger P, John M, Heil G, et al. AML1/MTG8 oncogene suppression by small interfering RNAs supports myeloid differentiation of t(8;21)-positive leukemic cells. Blood 2003; 101: 3157–63.
Pui CH, Schrappe M, Ribeiro RC, Niemeyer CM . Childhood and adolescent lymphoid and myeloid leukemia. Hematology (Am Soc Hematol Educ Program) 2004: 118–45.
Rowley JD . The role of chromosome translocations in leukemogenesis. Semin Hematol 1999; 36: 59–72.
So CW, Lin M, Ayton PM, Chen EH, Cleary ML . Dimerization contributes to oncogenic activation of MLL chimeras in acute leukemias. Cancer Cell 2003; 4: 99–110.
Srinivasan RS, Nesbit JB, Marrero L, Erfurth F, LaRussa VF, Hemenway CS . The synthetic peptide PFWT disrupts AF4-AF9 protein complexes and induces apoptosis in t(4;11) leukemia cells. Leukemia 2004; 18: 1364–72.
Zeisig DT, Bittner CB, Zeisig BB, Garcia-Cuellar MP, Hess JL, Slany RK . The eleven-nineteen-leukemia protein ENL connects nuclear MLL fusion partners with chromatin. Oncogene 2005; 24: 5525–32.
Armstrong SA, Golub TR, Korsmeyer SJ . MLL-rearranged leukemias: insights from gene expression profiling. Semin Hematol 2003; 40: 268–73.
Domer PH, Fakharzadeh SS, Chen CS, Jockel J, Johansen L, Silverman GA, et al. Acute mixed-lineage leukemia t(4;11)(q21;q23) generates an MLL-AF4 fusion product. Proc Natl Acad Sci USA 1993; 90: 7884–8.
Armstrong SA, Staunton JE, Silverman LB, Pieters R, den Boer ML, Minden MD, et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet 2002; 30: 41–7.
Pui CH, Relling MV, Downing JR . Acute lymphoblastic leukemia. N Engl J Med 2004; 350: 1535–48.
Elbashir SM, Martinez J, Patkaniowska A, Lendeckel W, Tuschl T . Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J 2001; 20: 6877–88.
Saxena S, Jonsson ZO, Dutta A . Small RNAs with imperfect match to endogenous mRNA repress translation. Implications for off-target activity of small inhibitory RNA in mammalian cells. J Biol Chem 2003; 278: 44312–9.
Chen CZ, Li L, Lodish HF, Bartel DP . MicroRNAs modulate hematopoietic lineage differentiation. Science 2004; 303: 83–6.
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A, et al. RAS is regulated by the let-7 microRNA family. Cell 2005; 120: 635–47.
Bridge AJ, Pebernard S, Ducraux A, Nicoulaz AL, Iggo R . Induction of an interferon response by RNAi vectors in mammalian cells. Nat Genet 2003; 34: 263–4.
Scacheri PC, Rozenblatt-Rosen O, Caplen NJ, Wolfsberg TG, Umayam L, Lee JC, et al. Short interfering RNAs can induce unexpected and divergent changes in the levels of untargeted proteins in mammalian cells. Proc Natl Acad Sci USA 2004; 101: 1892–7.
Hornung V, Guenthner-Biller M, Bourquin C, Ablasser A, Schlee M, Uematsu S, et al. Sequence-specific potent induction of IFN-alpha by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nat Med 2005; 11: 263–70.
Kim DH, Longo M, Han Y, Lundberg P, Cantin E, Rossi JJ . Interferon induction by siRNAs and ssRNAs synthesized by ph-age polymerase. Nat Biotechnol 2004; 22: 321–5.
Song E, Lee SK, Wang J, Ince N, Ouyang N, Min J, et al. RNA interference targeting Fas protects mice from fulminant hepatitis. Nat Med 2003; 9: 347–51.
Soutschek J, Akinc A, Bramlage B, Charisse K, Constien R, Donoghue M, et al. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 2004; 432: 173–8.
Schiffelers RM, Ansari A, Xu J, Zhou Q, Tang Q, Storm G, et al. Cancer siRNA therapy by tumor selective delivery with ligand-targeted sterically stabilized nanoparticle. Nucleic Acids Res 2004; 32: e149.
Urban-Klein B, Werth S, Abuharbeid S, Czubayko F, Aigner A . RNAi-mediated gene-targeting through systemic application of polyethylenimine (PEI)-complexed siRNA in vivo. Gene Ther 2005; 12: 461–6.
Takeshita F, Minakuchi Y, Nagahara S, Honma K, Sasaki H, Hirai K, et al. Efficient delivery of small interfering RNA to bone-metastatic tumors by using atelocollagen in vivo. Proc Natl Acad Sci USA 2005; 102: 12177–82.
Yano J, Hirabayashi K, Nakagawa S, Yamaguchi T, Nogawa M, Kashimori I, et al. Antitumor activity of small interfering RNA/ cationic liposome complex in mouse models of cancer. Clin Cancer Res 2004; 10: 7721–6.
Landen CN Jr, Chavez-Reyes A, Bucana C, Schmandt R, Deavers MT, Lopez-Berestein G, et al. Therapeutic EphA2 gene targeting in vivo using neutral liposomal small interfering RNA delivery. Cancer Res 2005; 65: 6910–8.
Thomas M, Gessner A, Vornlocher HP, Hadwiger P, Greil J, Heidenreich O . Targeting MLL-AF4 with short interfering RNAs inhibits clonogenicity and engraftment of t(4;11)-positive human leukemic cells. Blood 2005; 106: 3559–66.
Heidenreich O, Krauter J, Riehle H, Hadwiger P, John M, Heil G, et al. AML1/MTG8 oncogene suppression by small interfering RNAs supports myeloid differentiation of t(8;21)-positive leukemic cells. Blood 2003; 101: 3157–63.
Martinez N, Drescher B, Riehle H, Cullmann C, Vornlocher HP, Ganser A, et al. The oncogenic fusion protein RUNX1-CBFA2T1 supports proliferation and inhibits senescence in t(8;21)-positive leukaemic cells. BMC Cancer 2004; 4: 44.
Author information
Authors and Affiliations
Corresponding author
Additional information
Project supported by grants from the Deutsche Krebshilfe (10-2104-He2), the Deutsche José Carreras Leukämie-Stiftung (DJCLS-R03/10), and the Wilhelm-Sander-Stiftung (2003.169.1).
Rights and permissions
About this article
Cite this article
Thomas, M., Greil, J. & Heidenreich, O. Targeting leukemic fusion proteins with small interfering RNAs: recent advances and therapeutic potentials. Acta Pharmacol Sin 27, 273–281 (2006). https://doi.org/10.1111/j.1745-7254.2006.00282.x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1111/j.1745-7254.2006.00282.x
Keywords
This article is cited by
-
SMRI: A New Method for siRNA Design for COVID-19 Therapy
Journal of Computer Science and Technology (2022)
-
In vivo CRISPR/Cas9 targeting of fusion oncogenes for selective elimination of cancer cells
Nature Communications (2020)
-
Growth inhibitory effect of dihydroartemisinin on Bcr/Abl+ chronic myeloid leukemia K562 cells involve AKT, ERK and NF-κB modulation
Journal of Cancer Research and Clinical Oncology (2012)