-
PDF
- Split View
-
Views
-
Cite
Cite
Shin-Feng Fu, Wan-Chi Chou, Dinq-Ding Huang, Hao-Jen Huang, Transcriptional Regulation of a Rice Mitogen-Activated Protein Kinase Gene, OsMAPK4, in Response to Environmental Stresses, Plant and Cell Physiology, Volume 43, Issue 8, 15 August 2002, Pages 958–963, https://doi.org/10.1093/pcp/pcf111
Close - Share Icon Share
Abstract
Mitogen-activated protein kinase (MAPK) cascades play important roles in signal transduction of extracellular stimuli in eukaryotes. However, stimulatory signals for plant MAPKs have not been well elucidated. Here, a cDNA clone, termed Oryza sativaMAPK4 (OsMAPK4), from rice encoding a protein that showed homology with the eukaryotic MAPKs was isolated. According to the phylogenetic analysis, OsMAPK4 belongs to subgroup IV MAPK in plants. OsMAPK4 transcripts were expressed strongly in mature leaves and weakly in young leaves and panicles. The gene was also differentially expressed in roots at different developmental stages. In addition, the mRNA level of OsMAPK4 was up-regulated under sugar starvation, high salinity and cold treatments. These results suggest that this OsMAPK4 functions not only in developmental programs but also in stress-signaling pathways.
(Received December 14, 2001; Accepted June 5, 2002)
Plants are frequently exposed to unfavorable environmental stresses such as drought, extreme temperature, wounding and pathogen infection. Like all living organisms, plants must sense the signals and respond to the external stimuli that influence their growth and development. Progress in understanding signal transduction showed that reversible protein phosphorylation plays a pivotal role in response to extracellular signals (Somssich 1997). One of the major pathways by which environmental stimuli are transduced intracellularly is the mitogen-activated protein kinases (MAPKs) cascade. It plays a crucial role in regulating physiological and biochemical responses associated with extracellular stimuli. MAPKs are a family of conserved serine/threonine protein kinases that have been adapted throughout evolution to form parts of diverse signal-transduction pathways in all eukaryotes (Thomas 1992). MAPKs are activated by a highly conserved mechanism of sequential phosphorylation events in which a MAPK kinase kinase (MAPKKK) activates a MAPK kinase (MAPKK) that then activates a MAPK by phosphorylation of both tyrosine and threonine residues in the TXY motif (Herskowitz 1995, Nishihama and Machida 2000).
The biological processes in which MAPKs are involved are best understood in yeast and mammals. These organisms can be viewed as a model for understanding the role of MAPK cascades in plant systems. Six MAPKs are coded by the yeast genome, which has been fully sequenced. Five of them have been assigned to specific pathways involved in mating, filamentation, adaptation to osmotic stress, cell wall remodeling and sporulation (Madhani and Fink 1998). In mammalian cells, ERK (also known as MAPK), JNK and p38 MAPK have been reported (Gotoh et al. 1991, Han et al. 1994, Kyriakis et al. 1994). The ERKs have been shown to be involved in signaling hormones, neurotransmitters and signals for differentiation (Marshall 1994). The JNK1 and p38 MAPKs are activated in response to environmental stresses. Recently, a new MAPK termed Big MAPK (BMK1) or ERK5 was identified as a novel member of the MAPK family (Zhou et al. 1995). MAPKs may phosphorylate a number of different substrates in vivo, such as transcription factors and cytoskeletal proteins; and, a wide range of cellular signals have been found to feed into the pathways in which they operate (Lange-Carter et al. 1993).
More than 20 cDNAs encoding putative MAPKs have been cloned from different plants such as Arabidopsis (Mizoguchi et al. 1993), tobacco (Wilson et al. 1995), Petunia (Decroocq-Ferrant et al. 1995), oat (Huttly and Phillips 1995) and barley (Knetsch et al. 1996). Research on many of the MAPKs identified in plants suggests that MAPK cascades have a wider variety of roles in plants than those in animals and yeast (Mizoguchi et al. 1997). From an analysis of the sequence homology of predicted amino acid sequences, plant MAPKs can be further divided into at least four subgroups (Jonak et al. 1999). Within each subgroup, the high degree of sequence similarity appears to reflect a functional conservation between plant species (Jonak et al. 1999). According to available information, MAPKs in subgroups I and II are involved in signaling pathogen infections and abiotic stresses, whereas at least some of the MAPKs in subgroup III are involved in cell cycle regulation (Jonak et al. 1999). However, knowledge about the functions of subgroup IV is just emerging (Decroocq-Ferrant et al. 1995, Wilson et al. 1997).
Considering the conserved MAPK cascade present in many organisms, it is likely that the plant MAPK is activated posttranslationally by MAPK kinase (Morris et al. 1997). Interestingly, however, increased transcript levels of genes encoding a MAPK in plants have been observed in response to environmental stresses (Jonak et al. 1996, Mizoguchi et al. 1996, Seo et al. 1995) and hormonal treatments (Huttly and Phillips 1995, Marcote and Carbonell 2000). For instance, drought and cold induce mRNA accumulation of MMK4 in alfalfa (Jonak et al. 1996), and ATMPK3 in Arabidopsis (Mizoguchi et al. 1996). Similarly, the WIPK transcript encoding MAPK in tobacco accumulates after wounding (Seo et al. 1995). Gibberellins and cytokinins regulate PsMAPK3 mRNA levels in pea ovary shortly after fruit set is induced (Marcote and Carbonell 2000). In addition, Banno et al. (1993) and Nakashima et al. (1998) found a parallel correlation between the induction of tobacco MAPKKK, NPK1, gene expression and the occurrence of mitotic cell division and the formation of root primordia. Thus, the increase in expression of these MAPK and MAPKKK genes may contribute to the stress and growth response by increasing the amount of proteins available for activation (Mizoguchi et al. 1997).
Despite the large number of MAPK cDNAs isolated from plants, the MAPK activated by a particular stimulus, including stresses and hormones, has been identified in three subgroups, I, II and III (Jonak et al. 1999). Relatively little is known about the functions of members of subgroup IV. During pollen or ovule development, specific changes in transcript levels have been reported for two members of subgroup IV, NtF3 from tobacco (Wilson et al. 1997) and PMEK1 from Petunia (Decroocq-Ferrant et al. 1995). In addition, the mRNA level of ATMPK1, subgroup IV, was affected by salt-stress treatment (Mizoguchi et al. 1996).
In this study, we isolated and characterized a plant MAPK from rice, OsMAPK4. OsMAPK4 protein was most closely related to ATMPK1 and NtF3 of Arabidopsis and tobacco, respectively, and all belong to subgroup IV. This report describes the developmental regulation of the OsMAPK4 gene in response to the abiotic stimuli, sugar starvation, cold and salt stresses, leading to MAPK activation at the transcriptional level. The possible roles of subgroup IV MAPKs in signal transduction under environmental stresses are also discussed.
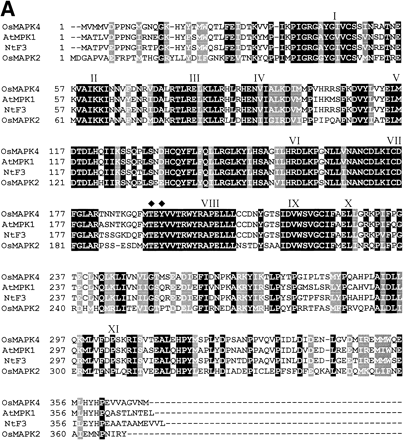
The EST database from the Rice Genome Research Program of the National Institute of Agrobiological Resources was screened for potential cDNAs of rice MAPKs using published MAPKs from other plants (Jonak et al. 1999). This cDNA clone, termed OsMAPK4 (AJ251330), was identified as potentially coding for the desired kinase and was chosen for further study. An alignment of the predicted amino acid sequences of OsMAPK4 with the cloned MAPKs from various plants is shown in Fig. 1A. The OsMAPK4 protein contained all eleven conserved amino acids and peptide motifs characteristic of 11 subdomains of protein kinases with serine/threonine specificity. Southern blot analysis in which genomic DNA digested with ClaI, XhoI, HindIII or EcoRI was hybridized to the radiolabeled OsMAPK4-specific probe indicated that the OsMAPK4 protein is encoded by a single-copy gene (data not shown).
It has been proposed that from an analysis of sequence homology of the predicted amino acid sequences, plant MAPKs can be grouped into at least four distinct families. The significance of the branching into different families is not yet fully understood, but suggests that MAPKs within one branch can serve similar functions. A phylogenetic tree was constructed based on the sequence alignment. OsMAPK4 can be grouped into subgroup IV (Fig. 1B). Comparison of the predicted protein sequences of the OsMAPK4 with MAPKs of other plants shows that OsMAPK4 is most homologous to the Arabidopsis ATMPK1, which also encodes a subfamily IV MAPK (Mizoguchi et al. 1993).
The expression of the OsMAPK4 gene was analyzed by Northern hybridization of gene-specific probe to total RNA from several rice tissues. The results showed that OsMAPK4 is distributed abundantly in mature leaves of 3-month-old plants and poorly in young leaves of 10-day-old seedlings and panicles (Fig. 2A). To determine whether OsMAPK4 was playing a role in root development, we used Northern blots to follow the accumulation of the transcript during root development. Different stages of root development were selected as follows: roots of 10-day-old (10 d), 20-day-old (20 d), 30-day-old (30 d) and 40-day-old (40 d) rice (Fig. 2B). Only a trace of OsMAPK4 transcript was detected at 10 d. The transcript level in root gradually increased at 20 d, became highest at 30 d, and then decreased drastically at 40 d (Fig. 2B).
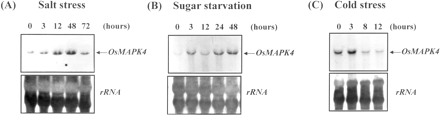
To characterize the expression of the OsMAPK4 gene in response to environmental stresses, total RNA extracted from suspension cells treated with 300 mM NaCl or cultured in the absence of sucrose were subjected to Northern blot. As shown in Fig. 3A, the mRNA level of OsMAPK4 was elevated within 3 h by high-salinity stress, continued to accumulate throughout 48 h, and then decreased at 72 h. Furthermore, the effect of sucrose starvation on the transcription of OsMAPK4 was also determined. The level of OsMAPK4 transcript increased significantly after 3 h of sucrose starvation (Fig. 3B). To gain further insight into the possible function of OsMAPK4 in plants, we exposed seedlings of rice to low-temperature stress. After exposure to cold stress, the OsMAPK4 transcript increased immediately within 3 h and decreased at 12 h (Fig. 3C).
In this study, we isolated and characterized a member of plant subgroup IV MAPKs, identified as OsMAPK4, from rice. Comparison of the amino acid sequences of OsMAPK4 with other cloned plant MAPKs showed high conservation of the central region, including all eleven domains that are necessary for the catalytic function of serine/threonine protein kinase. The N- and C-terminal extensions outside the eleven subdomains are more divergent than the catalytic core, but are still conserved in a specific subgroup of plant MAPKs (Hirt 1997).
The roles of MAPK pathways in developmental processes have been studied in a wide range of cell or tissue types and in a diverse set of organisms. In yeast, SMK1 encodes a developmentally regulated MAPK that is required for morphogenesis of the spore wall (Wagner et al. 1997). In rats, ERK transcripts are developmentally regulated within the nervous system, suggesting that they play unique physiological roles in response to different repertoires of signals (Boulton et al. 1991). One of the most studied developmental systems is that of PC-12 cells in which nerve growth factor (NGF) activates the MAPK cascade, leading to differentiation (Qui and Green 1992). In plants, the synthesis of tobacco MAPK, NtF3, occurs at the late-bicellular stage of maturation, as evident by an increase of mRNA synthesis (Wilson et al. 1997). Furthermore, high transcript levels of Arabidopsis MAPKK are found in old leaves, while much lower levels of expression are detected in young leaves (Hamal et al. 1999). Tomato MAPKK gene is also up-regulated during fruit development (Hackett et al. 1998). In this study, OsMAPK4 mRNA was expressed abundantly in mature leaves, and very weakly in young leaves (Fig. 2A). These observations suggest that the gene is predominantly expressed in leaves at a late developmental stage. This study also showed that the accumulation of the OsMAPK4 transcript is increasingly coincident with the developmental stage of roots from 10 to 30 d after seed germination, and the level of OsMAPK4 expression is significantly reduced at 40 d (Fig. 2B). Therefore, the OsMAPK4 system described here probably is involved in leaf and root development.
The activation of MAPK pathways in plants has been studied by specific treatments, such as wounding, salt, cold, drought and application of an elicitor (Hirt 1997). The mRNA level of ArabidopsisATMPK1, subgroup IV, was increased by salt-stress treatment, whereas it was transcriptionally unaffected by cold stress (Mizoguchi et al. 1996). In this study, OsMAPK4 falls within a fourth subgroup of MAPKs and is related to ATMPK1. The expression of OsMAPK4 gene was induced significantly by salt and cold stresses (Fig. 3). Therefore, it is suggested that plant MAPKs within the same subgroup are probably differentially regulated in monocots and dicots. Carbohydrates are major respiratory substrates in plant cells. Sugar starvation can occur in plants during leaf senescence (Hensel et al. 1993), extreme temperatures and leaf shedding, all of which significantly reduces photosynthesis (Tassi et al. 1992). Sugar starvation generally triggers physiological and cellular changes (Journet et al. 1986, Yu 1999). However, little is known about the signaling transduction pathway controlling these sugar starvation-related physiological responses (Yu 1999). In yeast, glucose starvation stimulated a stress-activated, MAPK pathway (Takeda et al. 1995). It has also been shown that glucose deprivation activates MAPK and induces cell death in human breast cells (Gupta et al. 1997). In this study, it is shown that the transcript of the OsMAPK4 gene accumulated under sugar starvation (Fig. 3B). This suggests that the MAPK functioned in response to sugar starvation among eukaryotes.
In plants, distinct subgroups of MAPKs have evolved for transmitting different types of signals such as most members of subgroup I and II for stress factors and subgroup III for mitogenic stimuli (Jonak et al. 1999). Very little is known about the functions of members of subgroup IV that include NtF3 in tobacco (Wilson et al. 1993), ATMAPK-1, -2 and -7 in Arabidopsis (Mizoguchi et al. 1993), and PMEK1 in Petunia (Decroocq-Ferrant et al. 1995). It has been reported that the NtF3 gene is expressed during anther development (Wilson et al. 1993). In Arabidopsis, the mRNA level of ATMPK1 increased slightly under salt stress (Mizoguchi et al. 1996). Mizoguchi et al. (1994) also suggested that auxin might function as an activator of ATMPK2. Similarly, the PMEK1 gene expression is clearly induced by auxin treatment and may be involved in the regulation of the cell cycle (Tréhin et al. 1998). In this study, we showed that abiotic stresses, salt, cold, and sugar starvation, induce transcriptional regulation of OsMAPK4 in rice. Moreover, the gene appears to be a part of the developmental program of leaves and roots. Based on these results, it is suggested that MAPKs in subgroup IV play a role in developmental processes as well as in the response to some environmental stresses.
Acknowledgments
We thank the members of the Japan Rice Genome Research Program for their kind donation of clone D41944. We are deeply grateful to Professor Toshio Murashige for critical reading of the manuscript. This work was supported by a grant from National Science Council (NSC 89-2311-B-006-005) of the ROC.
Corresponding author: E-mail, haojen@mail.ncku.edu.tw; Fax, +886-6-274-2583.
Fig. 1 Alignment of the predicted OsMAPK4 protein with MAPKs from other plants (A) and the phylogenetic relationship of OsMAPK4 to the cloned plant MAPKs (B). Roman numerals indicate the eleven major conserved subdomains of protein kinases identified previously (Hanks et al. 1988). Tyrosine and threonine residues, which must be phosphorylated for MAPK activation, are indicated by diamonds. The black shading indicates identical residues, and gray shading indicates conservative substitutions. Hyphens indicated gaps introduced to optimize alignments. The protein sequences shown in these diagrams are listed in the GenBank database under the following accession numbers: OsMAPK4 (AJ251330), OsMAPK2 (AJ250311), ATMPK1 (D14713), NtSIPK (U94192), NtF4 (X83880), SIMK (X66469), PsD5 (X70703), ATMPK6 (D21842), ZmMPK5 (AB016802), WIPK (D61377), IbMAPK (AF149424), ATMPK3 (D21839), SAMK (X82270), PsMAPK3 (AF153061), ZmMPK4 (AB016801), ASMAP1 (X79993), WCK-1 (AF079318), NtF6 (X83879), NRK1 (AB055515), ATMPK5 (D21841), ATMPK4 (D21840), MMK2 (X82268), ATMPK7 (D21843), ATMPK2 (D14714), PaMAPK (AF134730), NtF3 (X69971), PMEK1 (X83440). Similarity searches of nucleotide sequence were performed at the National Center Biotechnology Information using the BLAST network service (Altschul et al. 1990). The alignments were generated by the Clustal X program (Thompson et al. 1994). Phylogenetic analysis was performed using MEGA 1.02 software (Kumar et al. 1994).
Fig. 2 Northern hybridization analysis of OsMAPK4 transcripts in various organs of rice (A) and temporal expression pattern of OsMAPK4 during root development (B). Each lane was loaded with 10 µg of total RNA extracted from young leaves (YL) of 10-day-old seedlings, mature leaves (ML) of 3-month-old plants, young panicles (YP) at the stage before heading, and mature panicles (MP) at the heading stage. The RNA was isolated from roots at four different stages of 10-day-old (10 d), 20-day-old (20 d), 30-day-old (30 d) and 40-day-old (40 d) rice. As the OsMAPK4-specific probe, a 286-bp fragment was amplified by PCR using primers for forward: 5′-GAG TGA ATA TGT GAC AGG CA-3′ and reverse: 5′-AGC ATC TAA CAT TAC AAG CC-3′. The transcripts are shown by arrows at the right. The rRNA transcript level served as the internal loading control. Northern blot was performed according to Sambrook et al. (1989). Total RNA was extracted from leaves, roots, panicles, seedlings and suspension-cultured cells by using the RNeasy kit (QIAGEN, Hilden, Germany).
Fig. 3 The effect of salt stress (A), sucrose starvation (B) and cold stress (C) on the mRNA gene expression of OsMAPK4 in rice. RNA samples were prepared from treated rice suspension cells (A, B) and 7-day-old seedlings (C), which were sampled at indicated times after each stress treatment. Blots were probed with a radiolabeled 3′ end cDNA fragment of OsMAPK4. The transcripts are shown by arrows at the right. The rRNA transcript level served as the internal loading control.
Abbreviations
- ERK
extracellular-regulated protein kinase
- EST
expressed sequence tag
- JNK
c-Jun N-terminal kinase
- MAPK
mitogen-activated protein kinase.
The nucleotide sequence reported in this paper has been submitted to DDBJ, EMBL, GenBank under the accession number AJ251330 (OsMAPK4).
References
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. and Lipman, D.J. (
Banno, H., Hirano, K., Nakamura, T., Irie, K., Nomoto, S., Matsumoto, K. and Machida, Y. (
Boulton, T.G., Nye, S.H., Robbins, D.J., Ip, N.Y., Radziejewska, E., Morgenbesser, S.D., DePinho, R.A., Panayotatos, N., Cobb, M.H. and Yancopoulos, G.D. (
Decroocq-Ferrant, V., Decroocq, S., van Went, J., Schmidt, E. and Kreis, M. (
Gotoh, Y., Moriyama, K., Matsuda, S., Okumura, E., Kishimoto, T., Kawasaki, H., Suzuki, K., Yahara, I., Sakai, H. and Nishida, E. (
Gupta, A.K., Lee, Y.J., Galoforo, S.S., Berns, C.M., Martinez, A.A., Corry, P.M., Wu, X. and Guan, K.L. (
Hackett, R.M., Oh, S.A., Morris, P.C. and Grierson, D. (
Hamal, A., Jouannic, S., Leprine, A.S., Kreis, M. and Henry, Y. (
Han, J., Lee, J.D., Bibbs, L. and Ulevitch, R.J. (
Hanks, S.K., Quinn, A.M. and Hunter, T. (
Hensel, L.L., Grbic, V., Baumgarten, D.A. and Bleecker, A.B. (
Hirt, H. (
Huttly, A.K. and Phillips, A.L. (
Jonak, C., Kiegerl, S., Ligterink, W., Barker, P.J., Huskisson, N.S. and Hirt, H. (
Jonak, C., Ligterink, W. and Hirt, H. (
Journet, E.P., Bligny, R. and Douce, R. (
Knetsch, M.L.W., Wang, M., Snaar-Jagalska, B.E. and Heimovaara-Dijkstra, S. (
Kumar, S., Tamura, K. and Nei, M. (
Kyriakis, J.M., Banerjee, P., Nikolakaki, E., Dai, T., Rubie, E.A., Ahmad, M.F., Avruch, J. and Woodgett, J.R. (
Lange-Carter, C.A., Pleiman, C.M., Gardner, A.M., Blumer, K.J. and Johnson, G.L. (
Madhani, H.D. and Fink, G.R. (
Marcote, M.J. and Carbonell, J. (
Marshall, C.J. (
Mizoguchi, T., Gotoh, Y., Nishida, E., Yamaguchi-Shinozaki, K., Hayashida, N., Iwasaki, T., Kamada, H. and Shinozaki, K. (
Mizoguchi, T., Hayashida, N., Yamaguchi-Shinozaki, K., Kamada, H. and Shinozaki, K. (
Mizoguchi, T., Irie, K., Hirayama, T., Hayashida, N., Yamaguchi-Shinozaki, K., Matsumoto, K. and Shinozaki, K. (
Mizoguchi, T., Irie, K. and Shinozaki, K. (
Morris, P.C., Guerrier, D., Leung, J. and Giraudat, J. (
Nakashima, M., Hirano, K., Nakashima, S., Banno, H., Nishihama, R. and Machida, Y. (
Nishihama, R. and Machida, Y. (
Qui, M.S. and Green, S.H. (
Sambrook, J., Fritsch, E.F. and Maniatis, T. (
Seo, S., Okamoto, M., Seto, H., Ishizuka, K., Sano, H. and Ohashi, Y. (
Takeda, T., Toda, T., Kominami, K., Kohnosu, A., Yanagida, M. and Jones, N. (
Tassi, F., Maestri, E., Restivo, F.M. and Harmiroi, N. (
Thompson, J.D., Higgins, D.G. and Gibson, T.J. (
Tréhin, C., Planchais, S., Glab, N., Perennes, C. and James, T. (
Wagner, M., Pierce, M. and Winter, E. (
Wilson, C., Anglmayer, R., Vicente, O. and Heberle-Bors, E. (
Wilson, C., Eller, N., Gartner, A., Vicente, O. and Heberle-Bors, E. (
Wilson, C., Voronin, V., Touraev, A., Vicente, O. and Heberle-Bors, E. (
Yu, S.M. (