-
PDF
- Split View
-
Views
-
Cite
Cite
Hiroshi Katoh, Masahiko Ikeuchi, Targeted Disruption of psbX and Biochemical Characterization of Photosystem II Complex in the Thermophilic Cyanobacterium Synechococcus elongatus, Plant and Cell Physiology, Volume 42, Issue 2, 15 February 2001, Pages 179–188, https://doi.org/10.1093/pcp/pce024
Close - Share Icon Share
Abstract
PSII-X is a small hydrophobic protein, which is universally present in photosystem II (PSII) core complex among cyanobacteria and plants. The role of PSII-X was studied by directed mutagenesis and biochemical analysis in the thermophilic cyanobacterium Synechococcus elongatus. The psbX-disrupted mutant could grow photoautotrophically indicative of non-essential function, while it showed growth defect under low CO2 conditions. An active O2-evolving PSII complex was successfully isolated from the mutant and wild type. Protein composition of the isolated PSII complex was the same as wild type except for the absence of PSII-X. O2 evolution supported by artificial quinones was affected in the psbX-disrupted mutant. At high concentration of 2,6-dichlorobenzoquinone or 2,6-dimethylbenzoquinone, the mutant showed much lower activity than wild type, while not much difference was found at low concentration. These results imply that binding or turnover of quinones at the QB site depends, at least in part, on PSII-X protein in the PSII complex. Gel filtration chromatography of the PSII complex revealed that the dimeric structure of the complex was not greatly affected in the psbX-disrupted mutant.
(Received August 9, 2000; Accepted December 1, 2000).
Introduction
PSII is a multi-subunit pigment-protein complex with the enzymatic activity of light-dependent water-oxidizing plastoquinone reductase. The photochemical and enzymatic reactions catalyzed by PSII are strictly conserved among all oxygenic photosynthetic organisms including cyanobacteria, eukaryotic algae and higher plants, while quite diverse pigment-protein complexes have developed for light-harvesting antenna associated with PSII (Gant 1996, Vermaas 1993). With regard to protein structure, it is known that the PSII complex differs mostly in peripheral subunits between cyanobacteria and higher plants but shares the core parts in common (Ikeuchi 1992). Although photochemical and enzymatic reactions have been extensively characterized and attributed to a few reaction center proteins, roles of other subunits, conserved or not conserved, are still poorly understood.
Functional roles of the subunit proteins of PSII must be studied by a combination of both biochemical and biophysical analysis, and mutational analysis. For the mutational analysis, the mesophilic cyanobacterium Synechocystis sp. PCC 6803 has been widely used because it is naturally transformable and it can grow heterotrophically at the expense of glucose (Williams 1988, Vermaas 1993, Pakrasi 1995). Although there are several reports describing the isolation of an active O2-evolving PSII complex from this organism (Burnap et al. 1989, Kirilovsky et al. 1992, Tang and Diner 1994), the isolation of the active PSII complex from the PSII mutants is not yet fully established. Instead, mutant cells were examined to monitor the effects of directed mutations on the O2-evolving properties (Debus 1992). On the other hand, PSII complex isolated from thermophilic cyanobacteria Synechococcuselongatus or S. vulcanus is stable enough for reconstitution experiments of the O2-evolving activity or crystallization analysis (Shen and Inoue 1993, Zouni et al. 2000). Only introduction of His-tag into psbC has been successfully applied to the thermophilic S. elongatus, which enabled the efficient isolation of the active PSII complex (Sugiura and Inoue 1999). Thus, the targeted mutagenesis should be challenged for analysis of the stable O2-evolving PSII complex.
There are many directed mutants in cyanobacteria or a green alga, Chlamydomonas, where PSII genes were disrupted by genetic engineering (Pakrasi 1995). However, psbX mutant has never been reported. By protein sequencing, the 4.1 kDa hydrophobic protein (named PSII-X) was found in the O2-evolving PSII core complex but not in the reaction center in higher plants or in cyanobacteria (Ikeuchi et al. 1989a, Ikeuchi et al. 1989b, Ikeuchi et al. 1989c, Koike et al. 1989). This protein is encoded in the nuclear genome in higher plants, while it was encoded in the chloroplast genome in algae (Reith and Munholland 1995) as well as in cyanobacterial genome (Kaneko et al. 1996). However, no function had been assigned to PSII-X protein in the PSII complex because of lack of relevant data.
In this communication, we cloned and disrupted psbX of the thermophilic S.elongatus and studied biochemical properties of the PSII complex as well as the mutant phenotype. Results suggested that psbX is dispensable for the photoautotrophic growth but its product may be necessary for normal functioning at the QB site.
Materials and Methods
Strain and culturing conditions
The thermophilic cyanobacterium, S.elongatus, which was derived from a hot spring in Beppu district in Japan (Yamaoka et al. 1978) was grown at 45°C in DTN medium (Mühlenhoff and Chauvat 1996) under continuous illumination with white fluorescence lamps (20 to 50 µE m–2 s–1). Liquid culture was bubbled with air containing 1.0% (v/v) CO2. The psbX-disrupted mutant was maintained with 7 µg ml–1 chloramphenicol but propagated in the absence of antibiotics for analytical experiments.
Construction of the psbX-disrupted mutant
Based on DNA sequence of psbX from S. vulcanus (Tano et al. 1991), the DNA fragment harboring psbX gene was amplified from S. elongatus by polymerase chain reaction (PCR). The nucleotide sequence of the fragment was determined by the dideoxy termination method (Genetic Analyzer Model 310, PE-biosystems, U.S.A.). For construction of the psbX-disrupted DNA, two restriction sites were created within psbX by PCR amplification of upstream and downstream fragments. Oligonucleotide primers, 5′-CTGTGGCAGCCATGAATACC-3′ and 5′-GTCAACTCGAGGGAGGATGCAGACTGA-3′, were used for amplification of the upstream fragment of 905 bp and primers, 5′-TGAAAGAATTCTTTATTGTC-3′ and 5′-GGTACTCCTCCTCTTCCTCC-3′, were used for the downstream fragment of 637 bp. Each fragment was cloned into pT7Blue-T vector (Novagen, U.S.A.) and then combined with a promoter-modified chloramphenicol-resistant cassette. The cassette was derived from the coding region of chloramphenicol acetyl transferase from pACYC184 and two tandemly fused promoters, tac promoter from pKK223-3 and psbA1 promoter from S. elongatus. The resulting plasmid DNA was introduced into S. elongatus cells by electroporation and transformants were selected according to Mühlenhoff and Chauvat (1996), followed by colony isolation on a chloramphenicol-containing agar plate.
77 K fluorescence emission spectra
Low-temperature fluorescence emission spectra of thylakoid membranes (1 µg Chl ml–1) were measured at 77 K using a spectrofluorometer (model RF-5300PC, Shimazdu, Japan). Chlorophyll was excited at 435 nm.
Preparation of thylakoids
The cells at late log phase were harvested by centrifugation at 4,000×g for 10 min. After washing with 0.4 M sorbitol, 20 mM HEPES-NaOH (pH 7.0), 15 mM CaCl2 and 15 mM MgCl2 (hereafter designated as SHCM), cells were resuspended in 20 ml of SHCM medium supplemented with 5 mM aminocaproic acid, 1 mM benzamidine and 1 mM phenylmethylsulfonyl fluoride. The cells were broken with zirconia/silica beads (0.1 mm diameter, Biospec, U.S.A.) in a bead-beater (50 ml volume, Biospec) with three cycles of 30 s homogenization and 2 min cooling. The homogenate was centrifuged at 4,000×g for 10 min to remove cellular debris and then centrifuged at 100,000×g for 1 h to precipitate thylakoid membranes. After washing with SHCM medium once, the membrane pellet was resuspended in 1 M sucrose, 40 mM MES-NaOH (pH 6.5), 15 mM CaCl2, 15 mM MgCl2 and 10 mM NaCl (designated as SMCMN) and subjected to activity measurement or storage at –85°C.
Isolation of PSII core complex
Thylakoids (2 mg Chl ml–1) were solubilized with 1.8% (w/v) heptyl-β-d-thioglucoside (Sigma, U.S.A.) in SMCMN at room temperature for 10 min. The unsolubilized materials were removed by centrifugation at 72,000×g for 20 min at 4°C. The supernatant was diluted with an equal volume of a solution of 40 mM MES-NaOH (pH 6.5), 15 mM CaCl2, 15 mM MgCl2 and 10 mM NaCl and, then, centrifuged at 56,000×g for 20 min to remove PSI complex. Finally, the crude PSII particles were precipitated by centrifugation at 440,000×g for 1 h.
The crude PSII particles were solubilized with 2.0% (w/v) n-dodecyl-β-d-maltoside in SMCMN for 5 min (0.5 mg Chl ml–1) at 0°C and subjected to anion exchange column chromatography (HitrapQ, ÄKTA System, Amersham Pharmacia Biotech, Sweden). After loading, the column was washed with 200 mM NaCl in 40 mM MES-NaOH (pH 6.0) and 0.05% (w/v) n-dodecyl-β-d-maltoside. Then, the PSII complex was eluted with the linear gradient from 200 mM to 320 mM NaCl in the same medium. Fractions around 300 mM NaCl containing PSII complex were collected, diluted fivefold with 40 mM MES-NaOH (pH 6.0), 10 mM CaCl2 and 10 mM MgCl2 and precipitated by centrifugation at 541,000×g for 2 h. The PSII complex, thus obtained, was suspended in 40 mM MES-NaOH (pH 6.0), 50 mM NaCl and 25% (v/v) glycerol and stored at –85°C.
Gel filtration chromatography
Size of the PSII complex was estimated by gel filtration chromatography using Superdex-200 (PC3.2/30, SMART System, Amersham Pharmacia Biotech). The PSII complex was eluted with 30 mM MES-NaOH (pH 6.0), 150 mM NaCl and 0.03% (w/v) n-dodecyl-β-d-maltoside at a flow rate of 10 µl min–1. Molecular mass of the PSII complex was estimated according to the gel filtration calibration kit (Amersham Pharmacia Biotech).
SDS-urea-PAGE
PSII complex was solubilized with 2% (w/v) lithium dodecylsulfate, 60 mM dithiothreitol and 60 mM Tris-HCl (pH 8.8) at 0°C and subjected to SDS-urea-PAGE with the 16–22% (w/v) linear gradient of polyacrylamide gel containing 7.5 M urea as described by Ikeuchi and Inoue (1988).
Assay of oxygen evolving activity
Oxygen evolving activity of thylakoids (final 30 µg Chl ml–1) or PSII particles (final 4 µg Chl ml–1) was measured in SMCMN medium at 25°C using a Clark-type oxygen electrode with saturating light of about 3,000 µE m–2 s–1 in the presence of 2,6DCBQ or 2,6DMBQ as electron acceptors. Activity of intact cells (final 10 µg Chl ml–1) was measured in BG11 medium (Stanier et al. 1971) supplemented with 20 mM HEPES-NaOH (pH 7.0) in the presence or absence of artificial electron acceptors.
Results
Cloning and disruption of psbX gene
It is known that the DNA sequences of the two thermophilic cyanobacteria, S. vulcanus and S. elongatus, which were isolated from the separate hot springs in Japan (Yamaoka et al. 1978, Koike and Inoue 1983), are closely related to each other (for example, see Shimizu et al. 1992, Mühlenhoff et al. 1993). As expected, the DNA sequence of psbX as well as flanking regions from S. elongatus (Fig. 1A) was almost identical to that from S. vulcanus (Tano et al. 1991). Only 1 bp in psbX was different but the deduced amino acid was conserved. Compared with the N-terminal amino acid sequence of the 4.1 kDa PSII-X protein from S. vulcanus (Ikeuchi et al. 1989b), it was suggested that psbX codes for N-terminal transit of 10 amino acid residues, which was not present in the mature protein (Fig. 1B). Such a short transit sequence at N-terminus has not yet been found in psbX in other cyanobacteria (Kaneko et al. 1996) or algae such as Cyanophoraparadoxa (Stirewalt et al. 1995), but not unusual in general as will be described in the Discussion.
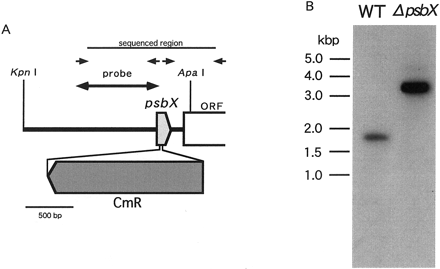
To disrupt psbX, a chloramphenicol-resistant cassette was inserted within psbX at restriction sites, which were created by PCR (Fig. 2A). After introduction by electroporation, chloramphenicol-resistant transformants were selected and propagated under photoautotrophic conditions for 2–3 months in the presence of antibiotics. To confirm segregation, Southern blot analysis (Fig. 2B) was carried out with a probe of 637 bp PCR product which carried a part of psbX. Upon digestion with ApaI and KpnI, the 1.7 kbp fragment was detected in the wild type, whereas the 3.4 kbp fragment carrying the 1.7 kbp cassette was solely detected in the mutant. This indicated that the mutant was homozygous and ready for further characterization. It is of note that psbX was dispensable for the photoautotrophic growth.
Effects of psbX disruption on growth
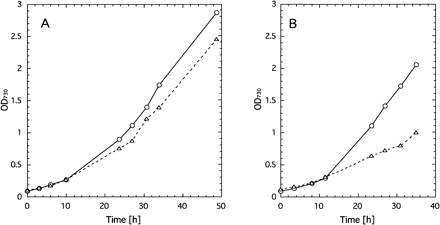
Photoautotrophic growth of the psbX mutant was slightly but reproducibly slower than that of wild type under any conditions of temperature and CO2. Especially, with 0.3% (v/v) CO2 at 50°C, the mutant grew much slower than wild type (Fig. 3). Taking into account that PSII-X protein is a component of the O2-evolving PSII core complex (Ikeuchi 1992), the growth properties of the psbX mutant may suggest that PSII complex depleted of PSII-X is not so stable under stressed conditions such as low CO2.
Effects of psbX disruption on content of photosystems
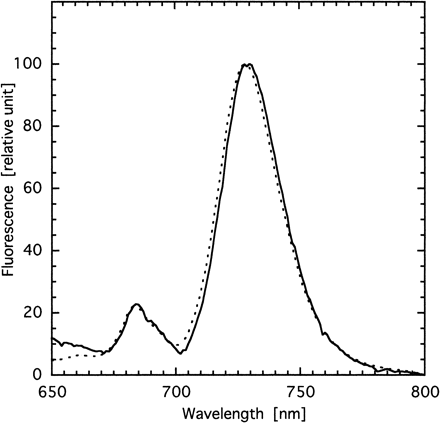
To examine whether or not the photosystem stoichiometry was affected by disruption of psbX, we measured chlorophyll fluorescence emission spectra of thylakoids at 77 K (Fig. 4). Clearly, there was no significant difference in the ratio of F695 and F725 between the mutant and wild type. This suggests that the disruption of psbX did not affect the assembly process of the PSII complex.
Effects of psbX disruption on oxygen evolution
We assayed O2 evolution activity with a Clark-type electrode. Without addition of artificial acceptors, no significant difference was detected in O2 evolution, which reflects CO2 fixation (not shown). When artificial quinones were added, O2 evolution activity, which was supported by PSII, was clearly different between the mutant and wild type. With less than 1 mM 2,6DCBQ, the psbX mutant showed activities almost identical to wild type, while higher concentrations of 2,6DCBQ did not give much enhancement to the mutant compared to wild type (Fig. 5). This seems to suggest that the affinity of PSII with quinone was somewhat affected by the absence of PSII-X protein. However, it must be mentioned that cellular activity of O2 evolution depends not only on the properties of PSII but also on permeability of the outer and plasma membranes.
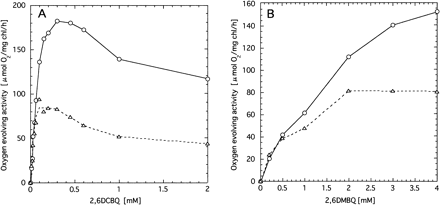
To avoid permeability barriers of cells, we isolated active thylakoid membranes. Clearly, the O2 evolution activities of the mutant thylakoids were much lower than those of wild type at a concentration of 2,6DCBQ above 0.1 mM (Fig. 6A). By contrast, 2,6DCBQ below 0.1 mM hardly gave any differences between the mutant and wild type. Thus, the concentration of 2,6DCBQ to achieve the maximum activity of the mutant PSII was about ten times lower in the thylakoids than in the cells. Moreover, not only the mutant but also wild-type thylakoids showed a peak, although the optimum concentration of 2,6DCBQ was 2 to 3 times higher in the wild type than in the mutant. Effects of 2,6DMBQ on O2-evolving activity were also different between wild type and the mutant (Fig. 6B). Namely, at higher concentrations the activity of the mutant was significantly lower than wild type, while almost comparable at lower concentrations.
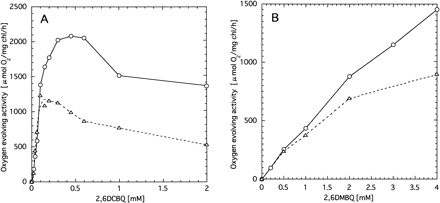
Isolation of photosystem II core complex
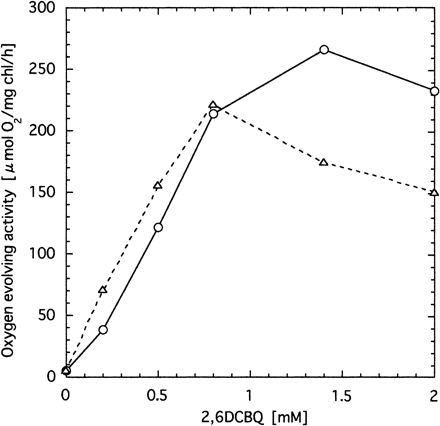
To study further the effects of psbX disruption, we isolated O2-evolving PSII core complex. Thylakoid membranes were solubilized with heptyl-β-d-thioglucoside and the crude PSII particles were isolated by differential ultracentrifugation. The recovery was about 3% on a chlorophyll basis. The effects of 2,6DCBQ or 2,6DMBQ on the O2-evolving activity were again different between wild type and the mutant (Fig. 7). Namely, at high concentrations the activity of the mutant PSII was clearly lower than that of wild type, while comparable at low concentration. The maximum activity of wild type (2,077 µmol O2 (mg Chl)–1 h–1) was achieved at 0.45 mM 2,6DCBQ, while that of the psbX mutant (1,277 µmol O2 (mg Chl)–1 h–1) was obtained around 0.1–0.2 mM 2,6DCBQ. Although features of the quinone-dependent O2 evolution of the PSII complex was very similar to those of thylakoid membranes, about ten times higher specific activities were obtained for the PSII particles, which can be accounted for by removal of major chlorophyll-binding proteins of Photosystem I.
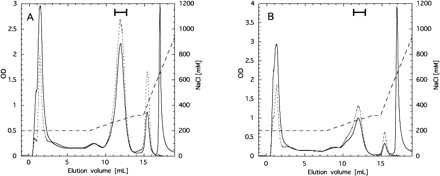
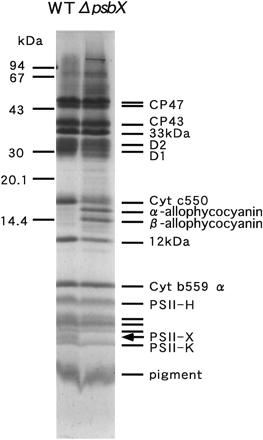
The PSII core complex was further purified using a HitrapQ column chromatography following solubilization with dodecylmaltoside. The PSII complex of both wild type and the mutant was eluted around 300 mM NaCl (Fig. 8). SDS-urea-PAGE unequivocally demonstrated that the complex from the mutant was depleted of the 4.1 kDa band (Fig. 9), which had been already assigned as PSII-X protein according to N-terminal amino acid sequencing (Ikeuchi et al. 1989b). The complete absence of this band in the mutant PSII indicates that no more PSII protein with blocked N-terminus comigrated underneath the band. It is of note that the rest of polypeptide composition of the mutant PSII was practically identical to that of wild type except the absence of the 4.1 kDa band. This suggests that PSII-X protein is dispensable for assembly and functioning of the rest of PSII complex. However, intensity of 15 kDa band of cytochrome c550 and 12 kDa band of PSII-U protein was slightly lower in the mutant PSII than in wild type. It must be mentioned that the quinone dependence of O2-evolving activity of the purified PSII was markedly changed compared with the crude particles (not shown). This suggests that free plastoquinone was largely lost or the QB site was greatly modified in the purified complex. Thus, we did not study further the O2-evolving activities of the purified complexes from the mutant and wild type.
Molecular size of PSII complex
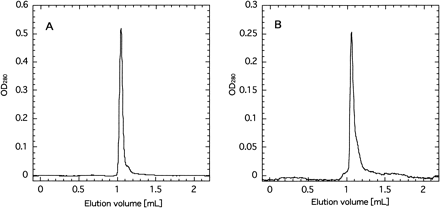
It is well known that the PSII complex exists as dimer not only in cyanobacteria but also in higher plants (Boekema et al. 1995), although the physiological significance of the dimerization has not yet been elucidated. Zheleva et al. (1998) reported that the dimeric PSII complex retained several low molecular proteins, which were not present in the monomer. We examined by gel filtration chromatography the molecular size of the PSII complex, which was depleted of PSII-X (Fig. 10). Both the wild type and mutant gave practically the same elution profiles peaking at 1.05 ml with a slight shoulder around 1.15 ml. The molecular sizes of the peak and the shoulder were calculated as 500–550 kDa and 330 kDa, respectively. The size around 500–550 kDa is consistent with the value (580 kDa) of the PSII dimer from the same organism in the literature (Sugiura and Inoue 1999), while the shoulder may suggest the presence of the PSII monomer, which might have been produced from the native dimer during the isolation procedures.
It is of note that the shoulder around 1.15 ml of the putative monomer was significantly larger in the mutant PSII than the wild type. Provided that the dimeric PSII is a native form in thylakoid membranes (Boekema et al. 1995), our results suggested that the PSII complex depleted of the PSII-X protein is less stable than the wild-type PSII, although we do not know whether the monomeric PSII was produced from the dimer during the isolation procedures or was already present in the thylakoid membranes in situ.
Discussion
This is the first report to describe the psbX-disrupted mutant. psbX was dispensable for the normal photoautotrophic growth. Taking advantage of thermostability of proteins from this cyanobacterium, we successfully isolated active thylakoids and PSII complexes from the mutant as well as wild type. Results of O2-evolving properties, protein composition and size of the complex were presented as evidence that PSII-X protein largely plays a functional role but a much smaller structural role in the PSII complex.
It is rather difficult to point out exactly which step is functionally affected in the mutant, since the overall activities of O2 evolution depend on rates of both sides of the reaction center. Analysis of the acceptor side is further complicated by the fact that the artificial quinones accept electrons at the QB site as well as at the site of plastoquinone pool with distinct affinity and maximum velocity (Vmax). Our preliminary results of concentration dependences of artificial quinones revealed that profiles of the mutant were somewhat affected but more or less similar to those of wild type in this work and in the literature (Satoh et al. 1995). The marked difference between the mutant and wild type was found in the activities at high concentrations of quinones (Fig. 6, 7). One can calculate Vmax at the QB site without other parameters by extrapolating activities to saturating concentration of 2,6DCBQ, since contribution of electron transfer at the site of plastoquinone pool becomes negligible (Satoh et al. 1995). Based on the double reciprocal plot of the data in Fig. 7A (not shown), Vmax of 2,6DCBQ at the QB site of the mutant PSII (500 µmol O2 (mg Chl)–1 h–1) was much lower than Vmax of wild-type PSII (1,230 µmol O2 (mg Chl)–1 h–1). This suggests that PSII-X protein is functionally involved in the QB site of PSII. On the other hand, the fact that activities of the mutant supported by lower concentrations of 2,6DCBQ were comparable to those of wild type implies that the rate on the donor side or at the site of plastoquinone pool was not much affected in the mutant. A similar tendency was also observed for 2,6DMBQ. However, we do not know whether or not the affinity of the QB site or the site of plastoquinone pool with the artificial quinones was changed by the mutation. More detailed studies are needed to better understand the functional role of PSII-X protein in PSII.
Although psbX was dispensable for the normal photoautotrophic growth, the mutant showed growth defect under stressed conditions such as low CO2 (Fig. 3). Since the low CO2 conditions often cause the photoinhibition of PSII via overreduction of the electron transport chain, the growth defect suggested that PSII-X may play an important role in protection of PSII possibly at the QB site. In this context, it is of note that mRNA level of psbX was specifically elevated upon high light treatment within 15 min, in addition to the marked elevation of well-known psbA transcripts as shown in DNA microarray analysis in Synechocystis sp. PCC 6803 (Y. Hihara, A. Kamei, M. Ikeuchi, unpublished). This suggests that PSII-X protein may turn over rapidly like D1 protein during recovery process for photoinhibition. These results also agree with the observation that PSII-X protein is located in the close vicinity of the PSII reaction center complex as shown by crosslinking to the large subunit of cytochrome b559 (Shi et al. 1999).
PSII-X is one of the small hydrophobic proteins associated with the O2-evolving PSII core complex in cyanobacteria as well as in higher plants (Ikeuchi et al. 1989b, Ikeuchi et al. 1989c). As it has a single stretch of hydrophobic residues (Fig. 1B), it is thought to span the thylakoid membrane once. In higher plants, psbX is encoded by the nuclear genome and its deduced product has a bipartite presequence like PSII-W protein and subunit II of coupling factor ATP synthase (Michl et al. 1994, Lorkovic et al. 1995, Kim et al. 1996). Membrane topology studies of these proteins including PSII-X showed that the mature N-termini are translocated into the thylakoid lumen. In vitro transport assay using chloroplasts from higher plants suggested that the precursor proteins were inserted into the thylakoid membrane via an unusual Sec-independent pathway and processed in the lumen (Thompson et al. 1998). However, such a lumen-targeting presequence was not always found in cyanobacterial or algal chloroplast genes, even though the same membrane topology is expected for the homologous PSII-X or subunit II of ATP synthase. This suggested that an additional lumen-targeting signal is present within the mature portion irrespective of the presence or absence of the presequence. It may be of note that PSII-X from only S. elongatus and higher plants has a positively-charged residue on the N-terminal side (Lys residue at position 7 in mature PSII-X in S. elongatus) of the membrane-spanning segment, while those from other cyanobacteria or algae have no charged residues in this region (Fig. 1). This may suggest that the transit sequence in S. elongatus as well as the bipartite presequence in higher plants facilitates the targeting of such a charged portion across the membrane.
The transit sequence of PSII-X (10 amino acid residues) is much shorter than those of the typical lumen-targeting hydrophilic proteins such as 33 kDa O2-evolving protein and plastocyanin (longer than 20 residues). To date, the short transit sequences have been found in several membrane proteins in some but not all cyanobacteria. For example, psaK (four residues in S. elongatus, Mühlenhoff et al. 1993, seven residues in Synechocystis, Ikeuchi unpublished results), slr1676 in Synechocystis sp. PCC 6803 (seven residues, Ikeuchi unpublished results) and psaI only in Anabaenavariabilis ATCC 29413 (11 residues, Sonoike et al. 1992). These transit sequences share common features in sequence, namely, a cluster of hydrophobic residues and lack of charged residues. Although the membrane topology of these proteins is not known, the unique short sequences may direct translocation of the mature N-terminus across the thylakoid membrane in some common way. Further study on the topology of these membrane proteins would reveal a common role of the unique short transit sequences.
Three dimensional structure of the PSII complex is one of the goals to understand the molecular process of water splitting. Recently, crystallization and 4 Å diffraction of X-ray was reported by using O2-evolving PSII complex from the same thermophilic cyanobacterium S. elongatus (Zouni et al. 2000). Further analysis of this complex may reveal the fine structure of PSII but 4 Å resolution may not be sufficient for full assignment of the electron density map. In the case of PSI complex, crosslinking experiments, mutant analysis and single particle analysis etc. also helped the interpretation of the X-ray crystallographic data. Our preparation of PSII complex devoid of PSII-X may be used as an alternative material for the structural analysis or other characterization.
Structure of O2-evolving PSII core complex and CP47-D1-D2 complex are now being analyzed mainly with two-dimensional crystals (Hankamer et al. 1999, Rhee et al. 1997, Rhee et al. 1998). The resolution was not so high that only membrane-spanning α-helices were visualized, enabling us to assign D1, D2, CP47 and CP43 proteins. However, there are several more α-helices located at the peripheral parts of the reaction center. Some of them were found at the interface of the PSII dimer, suggesting that some membrane-spanning subunit(s) of low molecular mass may be responsible for the dimerization of PSII. Our data of the psbX mutant excluded the possible involvement of PSII-X in dimerization of the PSII complex. We recently cloned psbI, psbK and psbT genes from S. elongatus and created their disruption mutants (H. Katoh, M. Iwai, M. Ikeuchi, unpublished). Further analysis of the PSII complexes from these mutants will give us a clue for structural organization of the PSII complex.
Acknowledgements
This work was supported by Grants-in-Aid for Scientific Research (to M.I.) from the Ministry of Education, Science and Culture, Japan, by the Program for Promotion of Basic Research Activities for Innovative Biosciences of Japan (to M.I.) and by a grant for Scientific Research from the Human Frontier Science program (to M.I.).
Corresponding author: E-mail, mikeuchi@ims.u-tokyo.ac.jp; Fax, +81-3-5454-4337.
Fig. 1 psbX of Synechococcus elongatus. Panel A: Nucleotide sequence of only the coding region of psbX together with deduced amino acid sequence is presented. Difference in nucleotide sequence of S. vulcanus (black box) was placed below the sequence of S. elongatus. The deduced sequence corresponding to N-terminal protein sequence of the4.1 kDa PSII-X from S. vulcanus is underlined. Panel B: Sequence alignment of PSII-X protein. Amino acid residues conserved between S. elongatus and other organisms were indicated by black boxes. Asterisks on the top and at the end of sequences indicate the totally conserved residues and termination codons, respectively. Putative transit sequence is shown in italic letters. The line indicates a hydrophobic segment, which may span the thylakoids. S. 6803: Synechocystis sp. PCC 6803 (EMBL: D64001), A. 7120: Anabaena sp. PCC 7120, C. paradoxa: Cyanophora paradoxa (EMBL: U30821), P. purpurea: Porphyra purpurea (EMBL: U38804), O. sinensis: Odontella sinensis (EMBL: Z67753), G. theta: Guillardia theta (EMBL: AF041468), A. thaliana: Arabidopsis thaliana (MUID: 96305797), spinach and wheat (Ikeuchi et al. 1989c). Note that the partial sequences of spinach and wheat were obtained by protein sequencing.
Fig. 2 Construction for insertional disruption of psbX (panel A) and Southern blotting analysis of psbX-disrupted mutant of S. elongatus (panel B). Arrows indicate primers for disruption construct. Genomic DNA of wild type (WT) and the mutant (ΔpsbX) was digested by KpnI and ApaI.
Fig. 3 Growth curves of wild type (circles) and psbX-disrupted mutant (triangles) as measured with OD730 at 70 µE m–2 s–1 at 50°C. Panel A: bubbled with 1% (v/v) CO2, B: 0.3% CO2.
Fig. 4 Low-temperature (77 K) fluorescence emission spectra of thylakoid membranes from wild type (solid line) and psbX-disrupted mutant (dotted line). Chlorophyll was excited at 435 nm.
Fig. 5 Effects of 2,6DCBQ concentration on the oxygen evolution activity of cells from wild type (circles) and psbX-disrupted mutant (triangles).
Fig. 6 Effects of 2,6DCBQ (panel A) and 2,6DMBQ (panel B) on the oxygen evolution activity of thylakoid membranes isolated from wild type (circles) and psbX-disrupted mutant (triangles).
Fig. 7 Effects of 2,6DCBQ (panel A) and 2,6DMBQ (panel B) on the oxygen evolution activity of PSII particles isolated from wild type (circles) and psbX-disrupted mutant (triangles).
Fig. 8 Elution profile of PSII core complexes from wild type (panel A) and psbX-disrupted mutant (panel B) in HitrapQ ion-exchange chromatography. Portions collected as the PSII core complex are shown as thick bars. Absorbances at 280 nm and 680 nm are indicated with solid line and dotted line, respectively. NaCl concentration for elution is indicated with dashed line.
Fig. 9 SDS-urea-PAGE profile of PSII core complexes from wild type (WT) and psbX-disrupted mutant (ΔpsbX). The position of 4.1 kDa PSII-X protein is indicated with an arrow. Positions of molecular size markers are indicated on the left.
Fig. 10 Elution profile of PSII core complex from wild type (panel A) and psbX-disrupted mutant (panel B) in Superdex 200 gel filtration chromatography.
Abbreviations
References
Boekema, E.J., Hankamer, B., Bald, D., Kruip, J., Nield, J., Boonstra, A.F., Barber, J. and Rögner, M. (
Burnap, R., Koike, H., Sotiropoulou, G., Sherman, L.A. and Inoue, Y. (
Debus, R.J. (
Gant, E. (
Hankamer, B., Morris, E.P. and Barber, J. (
Ikeuchi, M. and Inoue, Y. (
Ikeuchi, M., Koike, H. and Inoue, Y. (
Ikeuchi, M., Koike, H. and Inoue, Y. (
Ikeuchi, M., Takio, K. and Inoue, Y. (
Kaneko, T., Sato, S., Kotani, H., Tanaka, A., Asamizu, E., Nakamura, Y., Miyajima, N., Hirosawa, M., Sugiura, M., Sasamoto, S., Kimura, T., Hosouchi, T., Matsuno, A., Muraki, A., Nakazaki, N., Naruo, K., Okumura, S., Shimpo, S., Takeuchi, C., Wada, T., Watanabe, A., Yamada, M., Yasuda, M. and Tabata. S. (
Kim, S.J., Robinson, D. and Robinson, C. (
Kirilovsky, D., Boussac, A.G.P., van Mieghem, F.J.E., Ducruet, J.M.R.C., Stief, P.R., Yu, J., Vermaas, W.F.J. and Rutherford, A.W. (
Koike, H. and Inoue, Y. (
Koike, H., Mamada, K., Ikeuchi, M. and Inoue, Y. (
Lorkovic, Z.J., Schroder, W.P., Pakrasi, H.B., Irrgang, K.D., Herrmann, R.G. and Oelmueller, R. (
Michl, D., Robinson, C., Shackleton, J.B., Herrmann, R.G. and Klosgen, R.B. (
Mühlenhoff, U., Haehnel, W., Witt, H. and Herrmann, R.G. (
Mühlenhoff, U. and Chauvat, F. (
Pakrasi, H.B. (
Reith, M.E. and Munholland, J. (
Rhee, K.H., Morris, E.P., Barber, J. and Kuhlbrandt, W. (
Rhee, K.H., Morris, E.P., Zheleva, D., Hankamer, B., Kuhlbrandt, W. and Barber, J. (
Satoh, K., Oh-hashi, M., Kashino, Y. and Koike, H. (
Shen, J.-R. and Inoue, Y. (
Shi, L.X., Kim, S.J., Marchhant, A., Robinson, C. and Schröder, W.P. (
Shimizu, T., Hiyama, T., Ikeuchi, M. and Inoue, Y. (
Sonoike, K., Ikeuchi, M. and Pakrasi, H.B. (
Stanier, R.Y., Kunisawa, R., Mandel, M. and Cohen-Bazire, G. (
Stirewalt, V.L., Michalowski, C.B., Loffelhardt, W., Bohnert, H.J. and Bryant, D.A. (
Sugiura, M. and Inoue, Y. (
Tang, X.S. and Diner, B.A. (
Tano, H., Ishizuka, M. and Sone, N. (
Thompson, S.J., Kim, S.J. and Robinson, C. (
Vermaas, W. (
Williams, J.G.K. (
Yamaoka, T., Satoh, K. and Katoh, S. (
Zheleva, D., Sharma, J., Panico, M., Morris, H.R. and Barber, J. (