- Split View
-
Views
-
Cite
Cite
Angela H. A. M. van Hoek, Theo A. van Alen, Vera S. I. Sprakel, Jack A. M. Leunissen, Theo Brigge, Godfried D. Vogels, Johannes H. P. Hackstein, Multiple Acquisition of Methanogenic Archaeal Symbionts by Anaerobic Ciliates, Molecular Biology and Evolution, Volume 17, Issue 2, February 2000, Pages 251–258, https://doi.org/10.1093/oxfordjournals.molbev.a026304
Close - Share Icon Share
Abstract
Anaerobic heterotrichous ciliates (Armophoridae and Clevelandellidae) possess hydrogenosomes that generate molecular hydrogen and ATP. This intracellular source of hydrogen provides the basis for a stable endosymbiotic association with methanogenic archaea. We analyzed the SSU rRNA genes of 18 heterotrichous anaerobic ciliates and their methanogenic endosymbionts in order to unravel the evolution of this mutualistic association. Here, we show that the anaerobic heterotrichous ciliates constitute at least three evolutionary lines. One group consists predominantly of gut-dwelling ciliates, and two to three, potentially four, additional clades comprise ciliates that thrive in freshwater sediments. Their methanogenic endosymbionts belong to only two different taxa that are closely related to free-living methanogenic archaea from the particular ecological niches. The close phylogenetic relationships between the endosymbionts and free-living methanogenic archaea argue for multiple acquisitions from environmental sources, notwithstanding the strictly vertical transmission of the endosymbionts. Since phylogenetic analysis of the small-subunit (SSU) rRNA genes of the hydrogenosomes of these ciliates indicates a descent from the mitochondria of aerobic ciliates, it is likely that anaerobic heterotrichous ciliates hosted endosymbiotic methanogens prior to their radiation. Therefore, our data strongly suggest multiple acquisitions and replacements of endosymbiotic methanogenic archaea during their host's adaptation to the various ecological niches.
Introduction
Three to four times in their evolution, ciliated protozoa adopted an anaerobic life style (Embley et al. 1995 ; Hirt, Wilkinson, and Embley 1998 ). Most of them evolved “hydrogenosomes,” which arose from mitochondria in the course of their adaptation to anaerobic environments (Akhmanova et al. 1998 ; van Hoek et al. 2000 ). Such hydrogenosomes produce ATP and hydrogen, and, as a consequence, hydrogenosome-bearing ciliates acquired methanogenic endosymbionts that use the intracellular hydrogen for the reduction of carbon dioxide (Embley et al. 1995 ; Fenchel and Finlay 1995 ; Hirt et al. 1995 ; Akhmanova et al. 1998 ; Hirt, Wilkinson, and Embley 1998 ; Martin and Müller 1998 ). Such an endosymbiotic association is obviously mutualistic, since the functioning of the hydrogenosomes requires a low partial pressure of hydrogen (Fenchel and Finlay 1995 ; Akhmanova et al. 1998 ; Embley and Martin 1998 ) that can be guaranteed by methanogenic archaea that use intracellular hydrogen as a substrate for methane formation.
Anaerobic heterotrichous ciliates are especially suited for analysis of the evolution of such endosymbiotic associations in the various environments: they are closely related to each other, but they thrive in the most divergent niches, such as marine and freshwater sediments and the intestinal tracts of animals (Hirt, Wilkinson, and Embley 1998 ; van Hoek et al. 1998 , 1999 ). All of the anaerobic heterotrichous ciliates studied to date possess hydrogenosomes that most likely evolved from the mitochondria of their aerobic ancestors (Embley, Horner, and Hirt 1997 ; Hackstein et al. 1999 ; van Hoek et al. 2000 ). Therefore, one might assume that these ciliates hosted methanogenic endosymbionts from the very beginnings of their evolution because, as mentioned before, hydrogenosomes require a low intracellular hydrogen partial pressure for proper functioning.
We sequenced the small-subunit (SSU) rDNA genes of 18 anaerobic heterotrichous ciliates and their methanogenic endosymbionts in order to analyze the evolutionary relationships between anaerobic heterotrichous ciliates and their methanogenic endosymbionts. Here, we will show that coevolution between hosts and their endosymbionts can be demonstrated in only a few cases, notwithstanding a strictly vertical transmission of the methanogenic endosymbiont. We will also demonstrate that the endosymbionts from ciliates living in a particular ecological niche, e.g., freshwater sediments or intestinal tracts, respectively, are monophyletic and related to their free-living relatives. These observations strongly suggest that methanogenic endosymbionts were acquired prior to radiation but that endosymbiont replacements most likely accompanied the evolution of the anaerobic heterotrichous ciliates.
Materials and Methods
Isolation of Ciliates
Intestinal ciliates from two anuran species, one millipede, and seven different strains of cockroaches were isolated as described earlier (van Hoek et al. 1998 ). Free-living anaerobic ciliates from three freshwater sediments near Nijmegen, the Netherlands (Bruuk, Dekkerswald, and Plasmolen), were collected by hand and identified as described (van Hoek et al. 1999 ).
Microscopical Procedures
Endosymbiotic methanogenic bacteria were studied with a Leitz epifluorescence microscope (Doddema and Vogels 1978 ). The number of methanogenic endosymbionts was counted from color slides made from squash preparations of unfixed ciliates. The methanogenic endosymbionts were identified in situ by hybridization with a fluorescent archaea-specific probe (ArcR915, see below) according to Fokin et al. (1996) .
Gas Chromatography
Approximately 200–1,000 intestinal ciliates were isolated by electromigration and incubated in an anaerobic culture medium (Yamin 1978, modified). Methane production was measured with a Pye Unicam gas chromatograph (FID; Porapak Q column; 80/100 mesh). Ethane was used as an internal standard.
DNA Isolation, PCR Amplification, Sequencing, and Amplified Ribosomal DNA Restriction Analysis
DNA was isolated from single ciliates (van Hoek et al. 1998 ). The complete 16S rDNA sequence of the methanogenic endosymbiont of Nyctotherus ovalis from Blaberus sp. var. Nijmegen was determined after PCR amplification with the primers ArcF7 (5′-GTTGATCCTGCCAGAGG-3′) and ArcR1326 (5′-TGTGTGCAAGGAGCAGGGAC-3′). For all other methanogenic endosymbionts, partial 16S rDNA sequences (approximately 900 nt) were obtained with the primers ArcF26 (5′-TTGGGTTCGATTAAGCCAT-3′) and ArcR915 (5′-GTGCTCCCCCGCCAATTCCT-3′). PCR amplification of the 18S rDNA genes of the ciliates was performed using the primers euk-forward and euk-1200R or euk-300F and euk-reverse (van Hoek et al. 1998 , 1999 ).
PCR products of the 16S and 18S rRNA genes of the ciliates and their methanogenic endosymbionts were sequenced directly in both directions using an ABI Prism 310 Automated DNA sequencer (Perkin Elmer; van Hoek et al. 1998 ). For the determination of the complete rDNA sequence of the above-mentioned methanogenic endosymbiont, the PCR product was cloned using the TA Cloning Kit (Invitrogen) and INVαF One Shot competent Escherichia coli. Two clones were sequenced using six different methanogenic-archaea-specific and two plasmid primers. The DNA sequences were deposited in the EMBL database (accession numbers for 16S rRNA: AJ132639–AJ132655, AJ238131).
Amplified 16S rDNA of one to three individuals of all freshwater ciliates and two to nine individuals of the intestinal ciliates was subjected to amplified ribosomal DNA restriction analysis (ARDRA; van Hoek et al. 1998 ) with the enzymes HaeIII, MspI, and NciI (GibcoBRL), respectively, in order to confirm the validity of DNA sequence analysis. Also, the homogeneity of the endosymbiont populations was tested by ARDRA. For each freshwater sampling place, individual representatives of the various ciliate genera were subjected to ARDRA analysis of both 18S rDNA (of the host) and 16S rDNA (of the methanogenic endosymbionts) to validate the endosymbiont-host relationships. PCR was also used to control contamination of the washing buffers by environmental methanogenic archaea. Contaminated samples were excluded from further analysis.
Phylogenetic Analysis
DNA sequences were aligned using the PileUp program of the Wisconsin package, version 8.1 (Genetics Computer Group [GCG], Madison, Wis.; see also van Hoek et al. 1998 ). For distance analyses, alignments of 1,243 nt of the 18S rRNA genes and 770 nt of the 16S rRNA genes were used. In order to test the reliability of the different radiations of anaerobic ciliates, phylogenetic analysis was also performed with a data set that contained the concatenated partial 18S and complete ITS1, ITS2, and 5.8S rDNAs from at least one representative of the potential branches. EDNADIST (Rice et al. 1995 ) was used to calculate the sequence similarity and evolutionary distances using the Jukes and Cantor (1969) nucleotide substitution model. Phylogenetic trees were constructed using the neighbor-joining method (Saitou and Nei 1987 ). The NUCML program from the MOLPHY package (version 2.2; Adachi and Hasegawa 1996 ), PUZZLE (Strimmer and von Haeseler 1996 ), and likelihood mapping (Strimmer and von Haeseler 1997 ) were used to perform maximum-likelihood (ML) analysis. The TreeMap program (Page 1995 ), obtained from the Internet site http://taxonomy.zoology.gla.ac.uk/rod/treemap, was used to test for cospeciation of the hosts and their symbionts.
Results
The Endosymbionts of Anaerobic Heterotrichous Ciliates Are Methanogenic Archaea
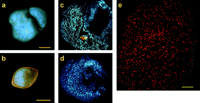
All anaerobic heterotrichous ciliates that have been described in this study host endosymbiotic methanogenic archaea (table 1 and fig. 1 ). These endosymbionts were tentatively identified by their characteristic blue (F420) autofluorescence (see Doddema and Vogels 1978 ) and by measuring the methane production of isolated ciliates. In addition, the endosymbionts of N. ovalis were identified by in situ hybridization with a fluorescent rDNA probe that is specific for methanogens (fig. 1e ).
Division of the ciliate cells causes a random distribution of the methanogenic endosymbionts over the two daughter cells. Also, the resting stages of the heterotrichous ciliates (cysts) contain endosymbiotic methanogens (fig. 1b ). Since transfaunation experiments with N. ovalis from the hindgut of cockroaches also could not provide evidence for an environmental transmission of the endosymbionts (van Hoek et al. 1999 ), we have to conclude that the endosymbionts of heterotrichous ciliates are vertically transmitted.
Molecular Phylogenies of Anaerobic Heterotrichous Ciliates and Their Methanogenic Endosymbionts
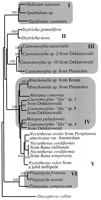
Phylogenetic analysis of the 18S rRNA genes of 18 anaerobic heterotrichous ciliates with the neighbor-joining method revealed monophyly for the intestinal ciliates and the existence of two to five paraphyletic clusters of the anaerobic, heterotrichous ciliates from freshwater sediments and intestinal tracts (fig. 2 ). The Plagiopyla and Trimyema species form a separate branch. This conclusion is also supported by ML analysis using NUCML (star-decomposition and quick-add-OTU options), quartet-puzzling, and likelihood mapping (see Materials and Methods). Moreover, using concatenated partial 18S, 5.8S, ITS1, and ITS2 rDNA sequences from at least one representative for each branch of the phylogenetic tree, the paraphly of the Caenomorpha species, certain Caenomorpha-like species, Brachonella, and Nyctotherus species could be confirmed. Only the branching order of the Brachonella and Caenomorpha-like species could not be resolved completely when using ML analysis. Consistently, this analysis supported clustering of the Nyctotherus species from frogs and millipedes on the one side and the N. ovalis species from the cockroach guts on the other side.
The 16S rRNA genes of the methanogenic endosymbionts of the above-mentioned anaerobic, heterotrichous ciliates were amplified by PCR on DNA from single ciliates using primers that are specific for the ribosomal repeat of methanogenic archaea. Since every ciliate cell hosts several thousands of endosymbionts but only a few phagocytized or adhering methanogens, the accidental PCR amplification of “environmental” methanogens is unlikely under these conditions. Moreover, PCR was performed on the washing buffers and the supernatants that were obtained during the isolation of the DNA in order to detect potential contamination by environmental methanogens.
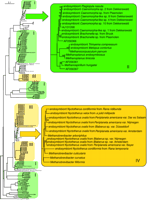
DNA sequence analysis of the 16S rRNA genes consistently revealed that only one rDNA of methanogenic origin could be amplified from a single ciliate. Also, ARDRA analysis of the amplified DNA confirmed the homogeneity of the rDNA. As a rule, ciliates from one and the same isolate hosted identical endosymbionts. Also, ciliates that had identical 18S rRNA genes but were from different environments (i.e., from Blaberus sp. var. Amsterdam and Periplaneta americana var. Bayer) possessed identical methanogens. However, DNA sequence analysis of the 16S rRNA genes of the endosymbionts of the majority of intestinal and free-living ciliates studied here revealed a significant degree of divergence between the endosymbionts. Notably, phylogenetic analysis of the 16S rRNA genes of these endosymbionts, together with 163 sequences from environmental methanogens, revealed that the endosymbionts of the intestinal ciliates cluster among the Methanobacteriales, whereas those of the free-living ciliates cluster among the Methanomicrobiales (fig. 3 ). Also, three DNA sequences from endosymbionts from Plagiopyla nasuta, Metopus contortus, and Trimyema compressum cluster with Methanomicrobiales. Three more DNA sequences from Plagiopyla frontata, Metopus striatus, and Metopus palaeformis cluster with different methanogens from environmental sources.
Comparative Analysis of Host and Endosymbiont Phylogenies
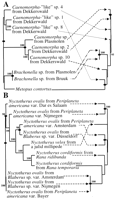
Because the endosymbionts of the ciliates from the intestinal tracts and freshwater sediments are substantially different (fig. 3 ), the potential coevolution of the ciliates and their endosymbionts was analyzed separately for the two different ecosystems. Figure 4 shows that there is only a poor match between host and endosymbiont phylogenies. Only rarely are parts of host and symbiont trees congruent. Different tree-building algorithms (parsimony and ML) and user-defined trees did not permit us to create trees that allowed a better match between host and endosymbiont phylogenies (not shown). Also, TreeMap analysis failed to reveal a substantial coevolution between the ciliates and their hosts. At best, three cospeciation events could be detected with this algorithm in each of the two ecosystems (fig. 4 ).
Discussion
The association between anaerobic heterotrichous ciliates and endosymbiotic methanogenic archaea depends on the presence of hydrogenosomes that provide intracellular hydrogen as a substrate for methane formation. Phylogenetic analysis of the hydrogenosomal SSU rRNA genes of different isolates of N. ovalis strongly suggests a common descent of the hydrogenosomes of anaerobic heterotrichous ciliates from the mitochondria of their aerobic relatives (Akhmanova et al. 1998 ; van Hoek et al. 2000 ). Therefore, it is likely that anaerobic heterotrichous ciliates hosted methanogenic endosymbionts from the beginning of their evolution, because the proper functioning of the hydrogenosomes relies on a low intracellular partial pressure of hydrogen. Low intracellular hydrogen levels can be maintained by the presence of methanogenic endosymbionts or, alternatively, by a small ciliate size (Fenchel and Finlay 1995 ; Biagini, Finlay, and Lloyd 1997 ; Embley, Horner, and Hirt 1997 ; Akhmanova et al. 1998 ).
The persistence of the endosymbionts through cell divisions and cyst (resting) stages permits a vertical transmission of the methanogenic endosymbionts. There is no evidence for an environmental transmission of the endosymbionts, since transfaunation experiments with ciliates possessing morphologically distinct endosymbionts (cf. fig. 1a, c, and d ) failed to reveal an uptake of endosymbiotic methanogens from the environment (van Hoek et al. 1999 ). Also, the observation that closely related ciliate species from the same sampling place (Dekkerswald; fig. 4 ) possess clearly divergent endosymbionts suggests a strictly vertical transmission of the endosymbionts and a high specificity of the association. Also, the fact that identical ciliates from different environments host identical endosymbionts (fig. 4 ) argues for a vertical transmission. Moreover, the endosymbionts of ciliates living in the intestinal tracts of frogs (Rana ridibunda and Rana temporaria; figs. 3 and 4 ) belong to the “intestinal” type of endosymbionts, notwithstanding that a wealth of free-living methanogens populates the particular habitat of the frogs and their larvae. Thus, all available evidence argues for a high specificity of the association and a long-lasting genetic isolation of the ciliates and their endosymbionts (fig. 4 ).
Comparable symbiotic associations exist between prokaryotes and insects, e.g., between the eubacterium Buchnera and many aphids (Baumann et al. 1995 ; Baumann, Moran, and Baumann 1997 ), and between flavobacteria and their termite and cockroach hosts (Bandi et al. 1994 , 1997 ). Phylogenetic analysis of these associations revealed a complete congruency between host and symbiont phylogenies. Therefore, it was very surprising that the phylogenetic analysis of the SSU rRNA genes of 18 anaerobic heterotrichous ciliates and their methanogenic endosymbionts did not provide any evidence for a significant match between the phylogenies of the ciliate hosts and their symbionts (fig. 4 ). Such an incongruence between host and symbiont phylogenies can have many reasons, for example, insufficient phylogenetic information (Distel 1998 ; Moran and Telang 1998 ). With our data, it is unlikely that the tree topology is biased by the use of partial SSU rDNA sequences: a phylogenetic analysis that includes the internal transcribed spacer (ITS-1 and ITS-2) DNA sequences and the 5.8S rDNA genes of the ciliates generates trees with an identical topology. We also have no indication that the topology of the phylogenetic trees is biased by the alignments or the tree-building algorithms (fig. 4 ).
We cannot exclude the possibility that the molecular clocks of the eukaryotic hosts and their prokaryotic endosymbionts are different and responsible for the low degree of congruence of host and symbiont trees (Hafner et al. 1994 ; Page and Hafner 1996 ). However, the observed relationship between the endosymbionts and their free-living relatives strongly suggests that host switches or endosymbiont replacements occurred, albeit rarely, during the evolution of these symbiotic associations. The main argument for this hypothesis stems from the observation that the endosymbionts of ciliates from freshwater sediment and those from the intestinal tracts of animals are monophyletic but clearly different in the two ecosystems. Also, the endosymbionts of Plagiopyla and Trimyema are related to free-living methanogens from the particular environments. The observed vertical transmission, the degree of DNA sequence divergence of the methanogenic endosymbionts, and the clustering of the endosymbionts in the phylogenetic tree (fig. 4 ) exclude a frequent uptake of methanogens from the environment. Notably, there is also no evidence for the presence of substantial numbers of identical or closely related free-living methanogens in the freshwater sediments studied. In the intestinal tract of the cockroach P. americana as well, there is no evidence for the presence of a Methanobrevibacter-like methanogen (W. Sprenger, personal communication; PCR and ribotype analysis, not shown). Such a distribution resembles the situation in certain bivalves that host chemoautotrophic eubacterial endosymbionts (Distel 1998 ; Krueger and Cavanaugh 1997 ). Also for these symbioses, a symbiont replacement has been postulated (Krueger and Cavanaugh 1997 ). In the case of the anaerobic, heterotrichous ciliates and their endosymbiotic methanogens, physiology and phylogeny clearly support the assumption of endosymbiont replacements. Figure 5 shows that at least two (but potentially as many as four) different clades of anaerobic heterotrichous ciliates evolved in the freshwater sediments, where they acquired closely related methanogenic endosymbionts. These endosymbionts cluster among the free-living “environmental” Methanomicrobiales. The Nyctotherus species that inhabit the intestinal tracts of the various animals are monophyletic and host endosymbionts that are related to intestinal Methanobacteriales. The phylogeny of the hydrogenosomes of N. ovalis suggests an origin of these organelles from the mitochondria of their aerobic ancestors (van Hoek et al. 2000 ).
Since the intestinal anaerobic heterotrichous ciliates and most of the free-living anaerobic heterotrichous ciliates are monophyletic, one might assume a common ancestry of their hydrogenosomes (fig. 2 ). Notwithstanding the obvious paraphyly of the Caenomorpha species, the hydrogenosomes of these ciliates might share a common ancestry with the hydrogenosomes of the Nyctotherus, Metopus, Brachonella, and Caenomorpha-like cluster too. This observation suggests that the common ancestor of all anaerobic heterotrichs already possessed methanogenic endosymbionts. The scenario shown here requires at least one endosymbiont replacement, but the lack of congruence between host and endosymbiont phylogenies within both clusters of intestinal and freshwater ciliates suggests that several endosymbiont replacements occurred. The reasons for these replacements are unclear, but one might assume that endosymbiont populations must suffer from the accumulation of deleterious mutations unless special mechanisms allow them to cope with this challenge (Hurst and McVean 1996 ; Moran 1996 ; Brynnel et al. 1998 ). In the absence of such mechanisms, dietary methanogens from the environment might eventually be able to replace an endosymbiont population that suffers from its genetic load (cf. Doolittle 1998 ).
Sequence Availability
The sequence data from the 16S rDNA genes from the methanogenic endosymbionts of the various anaerobic ciliates described here have been deposited in the EMBL database with the following accession numbers: methanogenic endosymbiont of N. ovalis from P. americana var. Amsterdam, AJ132639; methanogenic endosymbiont of N. ovalis from P. americana var. Bayer, AJ132640; methanogenic endosymbiont of N. ovalis from P. americana var. Dar es Salaam, AJ132641; methanogenic endosymbiont of N. ovalis from P. americana var. Nijmegen, AJ132642; methanogenic endosymbiont of N. ovalis from Blaberus sp. var. Amsterdam, AJ132643; methanogenic endosymbiont of N. ovalis from Blaberus sp. var. D;auusseldorf, AJ132644; methanogenic endosymbiont of N. ovalis from Blaberus sp. var. Nijmegen, AJ132645; methanogenic endosymbiont of N. cordiformis from R. temporaria, AJ238131; methanogenic endosymbiont of N. cordiformis from R. ridibunda, AJ132646; methanogenic endosymbiont of N. velox from a julid milliped, AJ132647; methanogenic endosymbiont of Caenomorpha-like sp. 1 from Dekkerswald, AJ132648; methanogenic endosymbiont of Caenomorpha-like sp. 4 from Dekkerswald, AJ132649; methanogenic endosymbiont of Caenomorpha-like sp. 8 from Dekkerswald, AJ132650; methanogenic endosymbiont of Caenomorpha sp. 10 from Dekkerswald, AJ132651; methanogenic endosymbiont of Caenomorpha sp. 2 from Dekkerswald, AJ132652; methanogenic endosymbiont of Caenomorpha sp. from Plasmolen, AJ132653; methanogenic endosymbiont of Brachonella sp. from Bruuk, AJ132654; methanogenic endosymbiont of Brachonella sp. from Plasmolen, AJ132655.
Pekka Pamilo, Reviewing Editor
Abbreviations: ARDRA, amplified ribosomal DNA restriction analysis; ITS, internal transcribed spacer; LSM, laser scanning microscope.
Keywords: anaerobic ciliates methanogenic archaea symbiosis endosymbiont replacement hydrogenosomes coevolution
Address for correspondence and reprints: J. Hackstein, Department of Microbiology and Evolutionary Biology, Toernooiveld 1, NL-6525ED Nijmegen, the Netherlands. E-mail: hack@sci.kun.nl.
Table 1 Numbers and Sizes of Methanogenic Endosymbionts and Methane Production Rates of Nyctotherus Ciliates

Table 1 Numbers and Sizes of Methanogenic Endosymbionts and Methane Production Rates of Nyctotherus Ciliates

Fig. 1.—Endosymbiotic methanogenic archaea of Nyctotherus ovalis. a–d, F420 autofluorescence. e, In situ hybridization. a, Nyctotherus ovalis from Blaberus sp. var. Amsterdam. b, Cyst of N. ovalis from the same isolate. c, Squash preparation of N. ovalis from Blaberus sp. var. Amsterdam. d, Squash preparation of N. ovalis from Periplaneta americana var. Amsterdam; note the rod shape of the methanogens. e, part of N. ovalis from Blaberus sp. var. Nijmegen; in situ hybridization with a probe specific for methanogenic archaea, labeled with Cy5. Confocal LSM. Bars indicate 25 ;gmm in a, c, and d, 20 ;gmm in b, and 10 ;gmm in e.
Fig. 2.—Neighbor-joining tree (Saitou and Nei 1987) inferred from approximately 1,243 positions of the 18S rDNA sequences of ciliates. I, rumen ciliates; II, aerobic, hypotrichous ciliates; III and IV, free-living, anaerobic, heterotrichous ciliates; V, intestinal, anaerobic, heterotrichous ciliates; VI, anaerobic, litostomatid ciliates. The distance data were bootstrap resampled 100 times (Felsenstein 1985 ). Only bootstrap values above 90% are displayed. Accession numbers are found in EMBL
Fig. 3.—Neighbor-joining tree (Saitou and Nei 1987) inferred from approximately 770 positions of the 16S rDNA of methanogenic archaea. The clades with the endosymbionts from freshwater (green, II) and intestinal (yellow, IV) ciliate sources are highlighted. The light-green boxes (I) indicate predominantly free-living methanogens from environmental sources such as sediments and rice fields. The light-yellow boxes (III) mark predominantly uncultured intestinal methanogens. The distance data were bootstrap resampled 100 times (Felsenstein 1985). Only bootstrap values above 90% are displayed in the highlighted area
Fig. 4.—TreeMap trees of hosts and symbionts based on 460 positions of the 18S rDNA sequences of the hosts and 770 positions of the 16S rDNA sequences of the endosymbionts. A, Freshwater ciliates (left tree) and their methanogenic endosymbionts (right tree). B, Intestinal ciliates (left tree) and their methanogenic endosymbionts (right tree). Corresponding pairs of ciliates and their endosymbionts are indicated by arrows. Only bootstrap values above 90% are displayed. Presumed cospeciation events are indicated by bullets
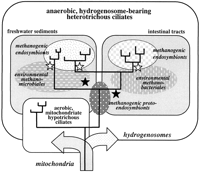
Fig. 5.—Illustration that summarizes crucial events in the evolution of anaerobic heterotrichous ciliates and their endosymbiotic methanogens. The metabolic basis for the adaptation of the ciliates to anaerobic niches such as freshwater sediments and intestinal tracts was the evolution of hydrogenosomes, most likely from the mitochondria of ciliates that were the common ancestors of both anaerobic heterotrichous and aerobic hypotrichous ciliates (arrows). The proper function of the hydrogenosomes requires low intracellular hydrogen concentrations that are most easily maintained by the acquisition of methanogenic protoendosymbionts. The radiation of the anaerobic ciliates in the various environments (solid black lines) was most likely accompanied by endosymbiont replacements (black stars). Also, the subsequent radiations (two in the intestinal tracts and two to four in the sediments) must have involved endosymbiont replacements (white stars). Apparently, the endosymbionts were acquired from different groups of free-living methanogens that were present in the various ecological niches
We thank Mr. G. H. M. Dekkers and Mr. J. Slippens (Photography and Illustrations) for their support with digitalization of the slides and preparation of the illustrations. We owe thanks to Mr. J. Eygensteyn (General Instrumentation) for his help with DNA sequence analysis. We gratefully acknowledge the advice of Mr. Adam McCoy (Biological Laboratories, Harvard University) on using TreeMap. This work was supported by a grant from the Netherlands Organization for Scientific Research (NWO) to A.H.A.M.v.H.
literature cited
Adachi, K., and M. Hasegawa.
Akhmanova, A., F. Voncken, T. van Alen, A. van Hoek, B. Boxma, G. Vogels, M. Veenhuis, and J. H. P. Hackstein.
Bandi, C., G. Damiani, L. Magrassi, A. Grigolo, R. Fani, and L. Sacchi.
Bandi, C., M. Sironi, C. A. Nalepa, S. Corona, and L. Sacchi.
Baumann, P., C.-Y. Lai, L. Baumann, D. Rouhbakhsh, N. A. Moran, and M. A. Clark.
Baumann, P., N. A. Moran, and L. Baumann.
Biagini, G. A., B. J. Finlay, and D. Lloyd.
Brynnel, E. U., C. G. Kurland, N. A. Moran, and S. G. E. Andersson.
Distel, D. L.
Doddema, H. J., and G. D. Vogels.
Doolittle, W. F.
Embley, T. M., B. J. Finlay, P. L. Dyal, R. P. Hirt, M. Wilkinson, and A. G. Williams.
Embley, T. M., D. A. Horner, and R. P. Hirt.
Felsenstein, J.
Fenchel, T., and B. J. Finlay.
Fokin, S. I., T. Brigge, J. Brenner, and H.-D. G;auortz.
Hackstein, J. H. P., A. S. Akhmanova, B. Boxma, H. R. Harhangi, and F. Voncken.
Hafner, M. S., P. D. Sudman, F. X. Villablanca, T. A. Spradling, J. W. Demastes, and S. A. Nadler.
Hirt, R. P., P. L. Dyal, M. Wilkinson, B. J. Finlay, D. M. Roberts, and T. M. Embley.
Hirt, R. P., M. Wilkinson, and T. M. Embley.
Jukes, T. H., and C. R. Cantor.
Krueger, D. M., and C. M. Cavanaugh.
Martin, W., and M. M;auuller.
Moran, N. A.
Moran, N. A., and A. Telang.
Page, R. D. M.
Page, R. D. M., and M. S. Hafner.
Saitou, N., and M. Nei.
Strimmer, K., and A. von Haeseler.
———.
van Hoek, A. H. A. M., A. S. Akhmanova, M. A. Huynen, and J. H. P. Hackstein.
van Hoek, A. H. A. M., V. S. I. Sprakel, T. A. van Alen, A. P. R. Theuvenet, G. D. Vogels, and J. H. P. Hackstein.
van Hoek, A. H. A. M., T. A. van Alen, V. S. I. Sprakel, J. H. P. Hackstein, and G. D. Vogels.