- Split View
-
Views
-
Cite
Cite
Carmen Palacios, Jennifer J. Wernegreen, A Strong Effect of AT Mutational Bias on Amino Acid Usage in Buchnera is Mitigated at High-Expression Genes, Molecular Biology and Evolution, Volume 19, Issue 9, September 2002, Pages 1575–1584, https://doi.org/10.1093/oxfordjournals.molbev.a004219
Close - Share Icon Share
Abstract
The advent of full genome sequences provides exceptionally rich data sets to explore molecular and evolutionary mechanisms that shape divergence among and within genomes. In this study, we use multivariate analysis to determine the processes driving genome-wide patterns of amino usage in the obligate endosymbiont Buchnera and its close free-living relative Escherichia coli. In the AT-rich Buchnera genome, the primary source of variation in amino acid usage differentiates high- and low-expression genes. Amino acids of high-expression Buchnera genes are generally less aromatic and use relatively GC-rich codons, suggesting that selection against aromatic amino acids and against amino acids with AT-rich codons is stronger in high-expression genes. Selection to maintain hydrophobic amino acids in integral membrane proteins is a primary factor driving protein evolution in E. coli but is a secondary factor in Buchnera. In E. coli, gene expression is a secondary force driving amino acid usage, and a correlation with tRNA abundance suggests that translational selection contributes to this effect. Although this and previous studies demonstrate that AT mutational bias and genetic drift influence amino acid usage in Buchnera, this genome-wide analysis argues that selection is sufficient to affect the amino acid content of proteins with different expression and hydropathy levels.
Introduction
The significance of symbiosis is now recognized for its abundance, wide distribution, and fundamental importance in many ecological processes (Douglas 1995 ). The advent of the molecular techniques has circumvented some of the initial difficulty in studying obligately intracellular, unculturable symbionts. Buchnera sp., the obligate endosymbiont located within specialized cells (bacteriocytes) in the body cavity of aphids (McLean and Houk 1973 ), is a Gram-negative γ-3 Proteobacteria that is closely related to Escherichia coli and other Enterobacteriaceae (Unterman, Baumann, and McLean 1989 ; Munson et al. 1991 ). Consistent with its strict maternal transmission (Buchner 1965 , pp. 297–332 and pp. 640–659), phylogenetic data support cospeciation between Buchnera and its aphid hosts dating back to 150–250 MYA, when this association is thought to have originated (Moran et al. 1993 ). Recently, the genome sequence of Buchnera strain APS (Shigenobu et al. 2000 ) demonstrated that long-term intracellular transmission has dramatically affected the content of this small endosymbiont genome (641 kb, compared with 4.6 Mb of E. coli K-12; Blattner et al. 1997 ).
Vertically transmitted, obligate endosymbionts may have relatively small effective population size (Ne) caused by recurrent bottlenecks upon transmission between host generations (Moran 1996 ) and limited genetic recombination between endosymbionts of different hosts (Moran and Baumann 1994 ; Funk et al. 2000 ; Wernegreen and Moran 2001 ). Therefore, the efficacy of selection may be reduced in intracellular replicating genomes (Muller 1964 ; Ohta 1973 ; Moran 1996 ) compared with free-living, recombining organisms such as E. coli, which are thought to have large long-term Ne (Selander, Caugant, and Whittam 1987 ).
Analyses of synonymous codon usage and amino acid composition are useful tools to explore shifts in the mutation-selection balance across bacterial species with different lifestyles. For example, in contrast to adaptive codon usage in E. coli and other free-living bacteria (Ikemura 1981 ; Bennetzen and Hall 1982 ), synonymous codon usage of intracellular pathogens such as Mycoplasma genitalium and Rickettsia prowazekii corresponds with local base compositional biases, and selection seems to have little effect (Andersson and Sharp 1996 ; McInerney 1997 ). Likewise, Buchnera shows an extreme AT bias at synonymous codon positions and spacer regions (Shigenobu et al. 2000 ) and lacks the adaptive codon bias shown by E. coli (Wernegreen and Moran 1999 ). Analysis of several protein-coding genes in this endosymbiont shows that mutational bias and drift drive not only codon usage but also amino acid changes (Ohtaka and Ishikawa 1993 ; Moran 1996 ; Brynnel et al. 1998 ; Clark, Baumann, and Baumann 1998 ; Clark, Moran, and Baumann 1999 ) and may contribute to gene loss (Mira, Ochman, and Moran 2001 ; Silva, Latorre, and Moya 2001 ).
Recently, full genome sequence data have strengthened multivariate analyses to explore factors that drive variation in amino acid and codon usage among genes within a given genome (e.g., de Miranda et al. 2000 ; Romero, Zavala, and Musto 2000 ). In this article, we build upon previous genome-level studies by employing multivariate analysis to identify major factors shaping amino acid usage in the full genomes of Buchnera APS and E. coli K-12. Our results argue that mutation and selection have strong but distinct effects on protein evolution in these two bacterial species.
Material and Methods
Genome Sequence Data
Coding sequences were extracted from the complete genome sequences of Buchnera sp. APS (chromosome and two plasmids; Shigenobu et al. 2000 ) and E. coli K-12 (Blattner et al. 1997 ) available at GeneBank (June 2001). Hypothetical proteins and annotated genes with less than 50 codons were excluded from the analysis to reduce stochastic variation (as recommended by CodonW tutorial; see below), resulting in a final sample of 479 loci from Buchnera and 2,919 loci from E. coli.
Multivariate Analysis of Amino Acid Composition
We used correspondence analysis (COA, Greenacre 1984 ) to identify the major factors that shape variation in amino acid usage among proteins of Buchnera and E. coli, as implemented by CodonW v. 1.4.2 for UNIX (available with John Peden at http://www.molbiol.ox.ac.uk/cu/). Because COA vectors may be affected by unusual amino acid usage of Buchnera plasmid-encoded proteins, plasmid genes were added after COA, and their positions were calculated based on vectors obtained from nuclear genes only. Major axes did not distinguish plasmid from nuclear genes of Buchnera.
Identifying Sources of Trends in Amino Acid Usage
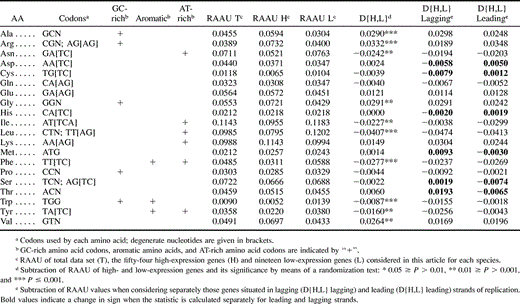
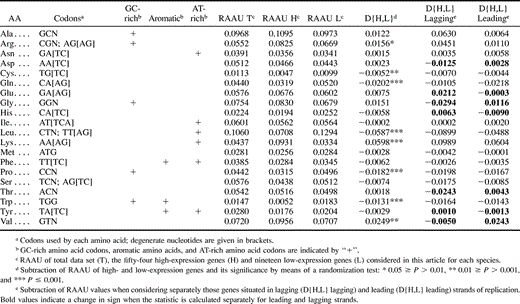
We used nonparametric tests of association to test the significance of associations between the position of loci (or amino acids) on principal axes of COA and 39 parameters relating to properties of loci or amino acids (JMP v. 4; SAS Institute). Given that multiple tests were performed, we adjusted values of type I error (α) by means of the Bonferroni correction (Sokal and Rohlf 1995 , pp. 236–240). Parameters of loci included several measures of nucleotide composition (e.g., AT skew defined by {A − T/A + T}, base composition at each codon position, etc.), gene length, relative frequency of aromatic amino acids, and the overall hydropathicity score of a protein (GRAVY; Kyte and Doolittle 1982 ). Properties of amino acids included molecular weight, hydropathy level (i.e., degree of hydrophilicity or hydrophobicity), and AT-richness of codons scored as in Clark, Moran, and Baumann (1999) . In this study, we selected the four amino acids at each extreme of that scale to define “GC-rich” and “AT-rich” categories (tables 1 and 2 ). We also included Leu in the AT-rich category, as this amino acid is encoded by TT[AG] and CTN.
Comparing High- Versus Low-Expression Genes
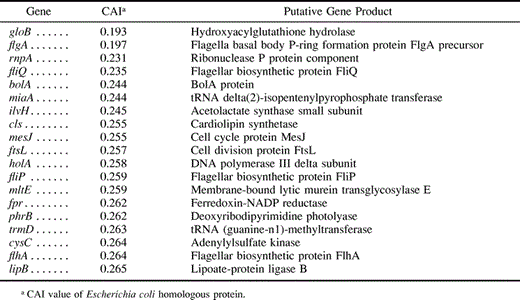
In E. coli, gene expression levels correlate closely with the codon adaptation index (CAI) (Sharp and Li 1987 ). But gene expression data in Buchnera are scarce. For high-expression genes, we selected the fifty-two ribosomal proteins, which are highly expressed across diverse taxa (Srivastava and Schlessinger 1990 ), mopA (groEL), which is highly expressed in Buchnera (Ishikawa 1984 ) and several other intracellular bacteria (e.g., the tsetse fly endosymbiont Wigglesworthia; Aksoy 1995 ) and its cotranscribed product mopB (groES;Sato and Ishikawa 1997 ). For low-expression genes, we selected 20 Buchnera loci that are homologous and named identically to the 639 E. coli genes with CAI values lower than 0.269 (table 3 ). From this pool of low-expression genes we omitted ilvH (acetolactate synthase involved in isoleucine and valine biosynthesis), a biosynthetic gene that may be highly expressed in this nutritional symbiont.
Comparisons of amino acid usage at high- and low-expression genes in Buchnera and E. coli were limited to the same set of homologous loci in both genomes. Homologs were identified conservatively as proteins with the highest degree of similarity and named identically in the two genomes. We can assume that pairs of homologs in Buchnera and E. coli are also orthologous because Buchnera lacks any species-specific duplicated proteins (with the exception of grpE, which was not included in comparisons of high- and low-expression genes) (Shigenobu et al. 2001 ). We quantified differences in relative amino acid usage (RAAU) between high- and low-expression genes by developing a new statistic, D{H,L} (the difference in RAAU of an amino acid at high- and low-expression genes). This statistic is defined as the RAAU at high-expression genes in a given genome minus the RAAU at low-expression genes in that genome and was calculated separately for each amino acid. We tested the significance of D{H,L} values using a sampled permutation (randomization) test (Sokal and Rohlf 1995 , pp. 803–819) and performed 1,000 permutations using a perl program kindly provided by E. T. Harley (http://www.cs.hmc.edu/∼eharley/research/tools/).
Locating Genes Situated on Leading Versus Lagging Strands of Replication
Asymmetrical mutational bias between the two complementary DNA strands may contribute to variation in both codon and amino acid usage (Karlin, Campbell, and Mrazek 1998 ; McInerney 1998 ; Lafay et al. 1999 ). To consider effects of leading versus lagging strands of replication, we determined strand orientation of genes on the basis of their position relative to the origin and terminus of replication. The presence of the DnaA-box of the Buchnera genome is thought to mark the origin of replication of the Buchnera chromosome (Shigenobu et al. 2000 ). But a shift of the GC skew in noncoding and synonymous third codon positions 13 kb upstream of this DnaA-box (Shigenobu et al. 2000 ) may correlate with the origin, as shown for other bacterial genomes (Lobry 1996 ; Blattner et al. 1997 ). To account for ambiguity in the location of the Buchnera origin, we considered this 13-kb region (from 627,681 to position 1 of the sequenced genome) as an “origin window.” Likewise, we defined the “terminus window” as the 13-kb region immediately opposite (180 degrees) the origin region (307,340 to 320,340). We excluded genes in these windows from comparisons of leading versus lagging strands. In contrast, the origin and terminus of E. coli are well defined experimentally (e.g., Yoshikawa and Ogasawara 1991 ) so that all genes can be assigned to the leading or lagging strand.
Results
Multivariate Analysis of Amino Acid Usage in Buchnera
Four of the nineteen axes generated by COA of Buchnera account for more than 50% of the total variance in amino acid composition of proteins.
Axis 1
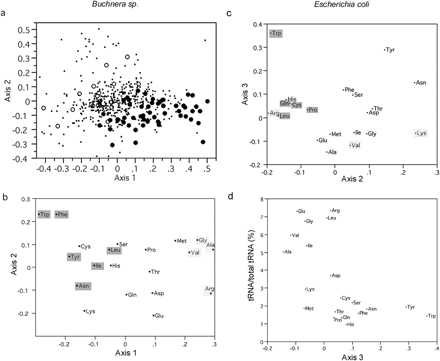
The first axis accounts for 23.0% of the total variation of the data. This axis correlates positively with GC content at first and second codon positions (rs (Spearman's Rho coefficient) = 0.89, P < 0.0001; 0.78, P < 0.0001; respectively) but notably, not third codon positions. Axis 1 also correlates negatively with aromaticity levels of each protein (rs = −0.67, P < 0.0001) and differentiates putative high- and low-expression genes in Buchnera (fig. 1a ).
Axis 2
The second axis of COA accounts for 13.3% of the variance. This axis correlates with the global levels of hydropathy of each Buchnera protein (rs = 0.70, P < 0.0001) and separates a group of presumed membrane proteins (with high GRAVY scores) from all other loci (data not shown). Axis 2 correlates negatively with AT skew (rs = −0.81, P < 0.0001) because of the nucleotide composition of codons for amino acids situated at the extreme of this axis (fig. 1b, e.g., the hydrophobic Phe TT[T or C] vs. the hydrophilic Lys AA[A or G]).
Other Axes
The third and fourth axes of COA in Buchnera account for 8.1% and 6.3% of the variation in the data, respectively. Axis 3 does not correlate significantly with any parameter considered. The fourth axis separates proteins that are rich in Cys, the most rare amino acid in Buchnera (table 1 ). The low dispersion observed in these and subsequent axes did not warrant further consideration.
Multivariate Analysis of Amino Acid Usage in E. coli
The first four axes of COA of the complete genome sequence of E. coli K-12 explain 49.2% of the total variation of the data (distributed along the 19 total axes). Axis 1 (19.1% of the total variation) correlates positively with the GRAVY score of proteins (rs = 0.83, P < 0.0001) and negatively with AT skew (rs = −0.67, P < 0.0001). This result agrees with a previous multivariate analysis of 999 E. coli genes, in which integral membrane proteins (IMP's) form a distinct group along Axis 1 (Lobry and Gautier 1994 ). Axis 2 (12.4% of the total variation) correlates with the gene expression (approximated using CAI) significantly (rs = 0.36, P < 0.0001) but not as strongly as the correlation previously reported (rs = 0.55, P < 0.0001; Lobry and Gautier 1994 ). A high correlation with C and A content at first codon positions (rs = −0.79, P < 0.0001; and 0.68, P < 0.0001, respectively) does not coincide with that expected from the correlation with gene expression, i.e., excess of guanine at first codon position (Gutierrez, Marquez, and Marin 1996 ). Axis 3 (9.4% of the variation) correlates positively with aromaticity and correlates with CAI almost as well as does Axis 2 (rs = −0.31, P < 0.0001). Both Axes 2 and 3 differentiate high- and low-expression genes considered in this study for E. coli (data not shown) as predicted by their correlations with CAI. As in Buchnera, the distinction of proteins rich in Cys (also the most rare amino acid in E. coli; table 2 ) along Axis 4 (8.3% of the variation) suggests that the frequency of Cys is highly variable among loci.
Comparative Analysis of Major Trends Shaping Amino Acid Usage in Buchnera and E. coli
Gene Expression
To test for differences in amino acid usage of high- and low-expression genes, we calculated the D{H,L} for each amino acid and determined the significance of observed values on the basis of the simulated null distribution of this statistic (tables 1 and 2 ). Amino acids that are significantly overrepresented in putative high-expression Buchnera genes are generally encoded by GC-rich codons (Arg, Ala, and Gly, with the exception of Val) and are not aromatic. Comparisons of E. coli and Buchnera indicate that most amino acids show similar trends in both species (i.e., are either over- or underrepresented in high-expression genes of both genomes) but to different degrees (fig. 2 ). No amino acid is significantly overrepresented in highly expressed genes of one species but significantly underrepresented in the other. Only two amino acids, Met and Asn, show opposite (but not significant) trends in Buchnera and E. coli. Notably, the aromatic amino acid Trp is severely reduced in high-expression genes of both species. The two other aromatic amino acids, Tyr and Phe (both of which are encoded by AT-rich codons), are significantly underrepresented in putative high-expression genes of Buchnera but not significantly in E. coli. Interestingly, Asn and Ile are relatively rare in the high-expression genes of Buchnera but show no strong bias in E. coli. Both Asn and Ile are encoded by AT-rich codons and are not aromatic. This pattern suggests selection against AT-rich amino acids at high-expression genes of Buchnera that is independent of selection against aromatic amino acids.
Because Axis 1 of COA in Buchnera distinguishes putative high- and low-expression genes, it is not surprising that D{H,L} values coincide with the positions of amino acids along Axis 1 (fig. 1b ). That is, those amino acids that are overrepresented in putative high-expression Buchnera genes (i.e., D{H,L} > 0; Val, Arg, Gly, and Ala; fig. 2 ) are positioned at high values along Axis 1 (fig. 1b ) (as are the high-expression genes; fig. 1a ). Likewise, amino acids overrepresented in low-expression Buchnera genes (i.e., with D{H,L} < 0; Trp, Leu, Phe, Tyr, Ile, and Asn) are positioned at low values of Axis 1 (as are the low-expression genes).
In E. coli, D{H,L} values only partially account for variation at Axes 2 and 3, both of which correlate with gene expression. Amino acids that are underrepresented in high-expression genes (Trp, Leu, Cys, Gln, and Pro; fig. 2 ) appear at the extreme of Axis 2, but only Trp is extreme in Axis 3 (fig. 1c ). Amino acids that are overrepresented in high-expression E. coli genes (Lys, Val, and Arg) are situated at the extreme of Axis 3, but only Lys is extreme in Axis 2.
Strand of Replication
Our COA of amino acid usage across the full genomes of Buchnera and E. coli did not clearly distinguish genes on the leading and lagging strands of replication in either species. But strand orientation and gene expression levels are not independent. As noted previously for E. coli and several other bacterial species (Francino and Ochman 1999 ), we found that putative high-expression genes tend to occur on the leading strand (78% of Buchnera and 96% of E. coli genes sampled here), perhaps because of selection to avoid collision between DNA and RNA polymerases (Brewer 1988 ).
Thus, we tested whether prevalence of high-expression genes on the leading strand, coupled with strand-specific mutational asymmetries, could account for the distinct amino acid profiles we observed at high- and low-expression genes. To distinguish the effects of strand orientation and gene expression level, we compared D{H,L} values calculated separately for genes on leading and lagging strands with D{H,L} values obtained when strand orientation was not considered (tables 1 and 2 ). In general, strand position did not affect the sign of significant D{H,L} values (i.e., had no effect on whether an amino acid is over- or underrepresented in high-expression genes). In Buchnera, switches in the sign of D{H,L} occurred only when this value was not significant; therefore, these sign changes may be attributed to random variation. The same result was found in E. coli, with the exception of Val (GTN) which is more abundant on the leading strand (see Discussion).
Hydrophobicity
We further explored the effect of hydrophobicity on amino acid usage in Buchnera and E. coli by comparing the inferred functions of genes at extreme positions of the axes that correlate with the hydropathy of proteins (i.e., loci positioned at >0.20 on Axis 2 of Buchnera and loci at >0.20 on Axis 1 for E. coli). In both genomes, these hydrophobic proteins tend to function as IMPs, with functions such as transport and anchoring of dehydrogenases. All but two Buchnera genes positioned >0.20 on Axis 2 were homologous to E. coli genes at extreme of Axis 1. These two exceptions, znuB and secY, also encode IMPs involved in transport. Moreover, secY is listed among E. coli genes at extreme of Axis 1 in a previous COA of this species (Lobry and Gautier 1994 ) but is absent from E. coli K-12.
Discussion
Shift in the Mutation-Selection Balance Contributes to Distinct Amino Acid Usage in Buchnera Versus E. coli
Previous studies demonstrate a strong effect of AT mutational bias on amino acid changes along Buchnera lineages, especially early in the symbiosis with aphids (Clark, Moran, and Baumann 1999 ). Virtually all Buchnera proteins are strongly influenced by directional mutational bias, as demonstrated for several other bacterial genomes (Singer and Hickey 2000 ). But we have addressed a distinct question: what processes drive variation in amino acid usage among loci within the Buchnera genome? The results of this study argue that selection relating to gene expression contributes to intragenomic variation in amino acid usage in this endosymbiont.
The first axis of the COA clearly distinguishes putative high- and low-expression Buchnera genes. To confirm this strong effect of gene expression on amino acid usage, we compared the RAAU of each amino acid at high- and low-expression genes using the new statistic D{H,L}. Several amino acids show significant differences in their abundance at putative high- and low-expression Buchnera genes. Amino acids significantly overrepresented at high-expression Buchnera genes include Ala, Gly, Arg, and Val, whereas those significantly underrepresented include Trp, Leu, Phe, Tyr, Ile, and Asn (table 1 and fig. 2 ).
This distinct amino acid profile of high-expression Buchnera loci may be shaped by selection against aromatic amino acids (Trp, Phe, and Tyr), which are expensive to biosynthesize (Craig and Weber 1998 ; Akashi and Gojobori 2002 ). In addition, amino acids that are more abundant at high-expression Buchnera genes tend to use codons that are relatively GC-rich at first and second positions (but not at third positions). This relative GC-richness suggests that selection counteracting a genome-wide AT mutational pressure in Buchnera is stronger at high-expression genes compared with low-expression genes.
Because many aromatic amino acids are encoded by AT-rich codons, selection against aromatic residues and selection against AT-rich codons are complementary and overlapping. But results of this study show their independent effects. For example, the amino acids Asn and Ile are encoded by AT-rich codons but are not aromatic. Thus, the low frequencies of Asn and Ile in high-expression Buchnera genes argues for selection against AT-rich codons at high-expression genes that cannot be explained by selection against aromaticity. Likewise, the low frequency of the very aromatic Trp (encoded by TGG) in high-expression genes suggests selection against aromatic amino acids that cannot be explained by selection against AT-rich codons.
The specific function of Buchnera as a nutritional endosymbiont may influence certain patterns of amino acid usage in this genome. Interestingly, the essential amino acids Trp and Leu are generally overproduced by Buchnera to supplement its host diet (Douglas and Prosser 1992 ; Bracho et al. 1995 ; Baumann et al. 1998 ) and might be relatively abundant amino acids in the endosymbiont cell. Therefore, the paucity of Trp and Leu in high-expression Buchnera genes may be shaped by host-level selection for energetic efficiency (see Rispe and Moran 2000 for models of host and symbiont-level selection). In addition, host-level selection may also influence the usage of several nonessential amino acids that Buchnera cannot synthesize but must acquire from the aphid host (Shigenobu et al. 2000 ).
Although strong AT bias in Buchnera may drive distinct amino acid usage at high- and low-expression genes, this mutational bias may actually narrow that difference in some cases. Notably, Lys, encoded by the AT-rich codons AA[AG], is significantly overrepresented in ribosomal proteins of E. coli (RAAU of 0.0948 in ribosomal proteins vs. 0.0437 genome-wide). But Lys is the most common amino acid across the Buchnera genome (RAAU of 0.0988 genome-wide), consistent with the strong AT mutational bias. The slightly higher frequency of Lys in high-expression Buchnera genes is not significant given the genome-wide abundance of this amino acid.
Despite a strong effect of gene expression level on amino acid usage in Buchnera, we found no effect of gene expression on patterns of relative synonymous codon usage (RSCU) in this species. That is, in a multivariate analysis of RSCU of the 479 Buchnera loci included in this study, no major axis distinguished putative high- and low-expression genes (data not shown). This genome-wide analysis supports previous evidence that mutational bias and drift shape codon usage in Buchnera (e.g., Brynnel et al. 1998 ; Wernegreen and Moran 1999 ) and adds to the accumulating evidence that translational selection is not sufficiently strong or effective (or both) to counter the effects of genetic drift and mutational bias on synonymous codon usage. Therefore, gene expression levels influence amino acid usage, where selection may act more strongly, but not synonymous codon usage, where weak selection is apparently ineffective in small Buchnera populations. Another plausible explanation for an absence of translational selection on codon usage in Buchnera could be the equal abundance of tRNAs in this genome, which contains only one or a few copies of each tRNA. The reduced tRNA populations in the AT-rich genome of the parasite R. prowazekii was also postulated as a major factor in the absence of codon usage biases (Andersson and Sharp 1996 ). Interestingly, a more detailed analysis of codon bias in Buchnera suggests that different mutational biases on leading and lagging strands affects synonymous codon usage (Claude Rispe, personal communication).
In E. coli, hydrophobicity is the primary factor shaping variation in amino acid usage among proteins, but the effects of gene expression (although secondary) are nonetheless apparent. Comparisons of putative high- and low-expression genes show many similarities between Buchnera and E. coli because no amino acid shows significantly different trends in the two genomes (fig. 2 ). The significant underrepresentation of Trp in high-expression genes of both genomes suggests selection against the use of this aromatic amino acid. On the basis of similar results for E. coli, Lobry and Gautier (1994) suggested that the energetic costs of aromatic amino acids may account for their low abundance in high-expression genes. Recently, a more detailed study of metabolic efficiency in bacteria analyzed the cost of each amino acid in terms of high-energy phosphate bonds (the most expensive amino acids being the aromatic Trp, Phe, and Tyr) (Akashi and Gojobori 2002 ). The authors found decreased abundance of costly amino acids in high-expression E. coli loci regardless of gene function. Interestingly, our comparison of high- and low-expression E. coli genes is based on a limited gene sample but yielded results entirely consistent with this previous genome-wide analysis. Only Phe, Pro, Ser, and Arg differ in whether they show significant changes with gene expression, but this could be attributed to our necessarily smaller gene sample. Consistent with our results, Akashi and Gojobori (2002) also found no relationship between gene expression and the abundance of amino acids encoded by AT-rich or GC-rich codons in E. coli. This pattern in E. coli contrasts with the striking correlation in Buchnera between GC-richness of amino acid codons and gene expression (as reflected in significant correlations between Axis 1 and GC content at first and second codon positions and the biased amino acid profiles at high- vs. low-expression genes [fig. 1b,fig. 2] ).
Previous work shows that tRNA pools match overall amino acid usage of proteins in several genomes (Yamao et al. 1991 ). In E. coli, the amino acid composition of high-expression genes correlates more strongly with tRNA abundances than do low-expression genes (Lobry and Gautier 1994 ). This trend has been interpreted as coadaptation between amino acid composition of proteins and tRNA-pools to enhance translational efficiency (Lobry and Gautier 1994 ; Akashi and Eyre-Walker 1998 ). In this study, we plot the two axes that correlated with gene expression in the E. coli COA against major tRNA abundances (Dong, Nilsson and Kurland 1996 ). Axis 2 is not correlated with tRNA abundance of the corresponding amino acid, whereas Axis 3 correlates significantly with tRNA abundance (but only before applying the Bonferroni correction; rs = −0.66, P < 0.001; fig. 1d ). This data supports previous evidence that translational selection may shape amino acid usage in E. coli (Lobry and Gautier 1994 ). This pattern contrasts with Buchnera, in which equal abundances of tRNA molecules or reduced efficacy of selection may limit effects of translational selection on both codon usage (see above) and amino acid usage.
Other Processes that may Drive Intragenomic Variation in Amino Acid Composition of Buchnera and E. coli
We calculated D{H,L} on the leading and lagging strands separately to distinguish the effects of strand orientation and gene expression level. In Buchnera, strand orientation does not account for the observed differences between high- and low-expression genes (table 1 ). But in E. coli, switches in the sign of D{H,L} depending on strand orientation suggest that strand-specific mutational biases may affect amino acid usage (table 2 ). Namely, high-expression genes of E. coli may experience distinct mutational pressures by virtue of their prevalence on the leading strand of replication. For example, the strong bias of Val (encoded by GTN) on the leading strand in E. coli and other species, independent of gene expression level, is consistent with a G > C and T > A skew on the leading strand resulting from strand mutational asymmetries (Mackiewicz et al. 1999 ; Rocha, Danchin, and Viari 1999 ). Likewise, strand position apparently contributes to the overrepresentation of Val in high-expression E. coli genes in our study.
Strand-specific biases may also drive asymmetries between the coding and noncoding DNA strands, as a result of transcription-associated mutation or DNA repair (or both) (e.g., Francino and Ochman 1999 ). If this bias increases with transcription levels, then transcription-associated asymmetries may contribute to differences between high- and low-expression genes. For example, it is possible that C→T mutational bias on the coding strand (Beletskii and Bhagwat 1996 ) may contribute to the observed high frequencies of certain GT-rich amino acids (e.g., Gly [GGN] and Val [GTN]) at high-expression genes in Buchnera, and low frequencies of CA-rich amino acid codons ([Pro (CCN)], Gln [CA(AG)]) at these genes in E. coli. But transcription-associated biases alone cannot account for distinct amino acid profiles in high- and low-expression genes. For example, several amino acids that use GT-rich codons are significantly underrepresented at high-expression genes of one or both species (Trp [TGG], Phe [TT(TC)] Cys [TG(TC)]). Nor can transcription-associated biases account for the observed reduction in the AT content of amino acid codons at high-expression genes because a C→T mutational bias is expected to increase T-richness of high-expression genes.
In analyses of single genomes, apparent differences in amino acid usage at high- and low-expression genes may partially reflect distinct structural or functional requirements of the proteins selected. But gene-specific structural or functional constraints have a minimal effect on the conclusions of our study, which is based on a comparison of an identical set of genes in Buchnera and E. coli. Moreover, the consistency of our results with a previous genome-wide study of E. coli (Akashi and Gojobori 2002 ) suggests that, for the purposes of this study, our limited gene sample is largely characteristic of high- and low-expression genes. Paralogous genes also pose complications for analyses of individual genomes because amino acid profiles of paralogs may be similar because of common ancestry. But in this study the observed differences between E. coli and Buchnera cannot be explained by gene duplication in Buchnera, which basically represents a subset of the E. coli genome (Shigenobu et al. 2000 ).
Conclusions
In summary, this comparative genome analysis of Buchnera and E. coli K-12 highlights important differences in the effects of mutation and selection on amino acid usage in these species. Mutational bias and genetic drift likely explain trends toward genome reduction and AT richness in Buchnera, as well as other bacterial endosymbionts, intracellular parasites, and organelles (Andersson and Kurland 1998 ; Mira, Ochman, and Moran 2001 ; Selosse, Albert, and Godelle 2001 ). This analysis of the Buchnera genome suggests a shift in the mutation-selection balance across loci that depends on gene expression level. Strong AT mutational bias impacts all Buchnera loci, but its effect on amino acid usage is apparently greater at low-expression genes than at high-expression genes. Tighter selective constraints at high-expression Buchnera genes may limit changes toward AT-rich or aromatic amino acids (or both), and may act either at the level of the bacterium or the aphid host (Rispe and Moran 2000 ). In addition to selection associated with gene expression levels, selection to maintain hydrophobic amino acids in IMPs also shapes global amino acid composition of Buchnera as a secondary force. This and previous analyses of E. coli show that hydrophobicity of IMPs is the primary source of variation in amino acid usage among loci, whereas factors relating to gene expression (e.g., selection for amino acids with high tRNA abundances or biased strand orientation of high-expression genes) are secondary. These distinct patterns of protein evolution in Buchnera and E. coli may result from different magnitudes and effects of mutational pressure versus selection because of the obligate host association of the endosymbiont. Further genome-level studies of other endosymbionts will provide a comparative framework to determine whether bacterial species with similar lifestyles show parallel modes of evolution and whether processes shaping amino acid usage in Buchnera extend to other endosymbiont species.
Adam Eyre-Walker, Reviewing Editor
Abbreviations: Ne, effective population size; COA, correspondence analysis; RAAU, relative amino acid usage; D{H,L}, difference in RAAU of an amino acid at high- and low-expression genes; rs, Spearman's Rho coefficient; RSCU, relative synonymous codon usage.
Keywords: amino acid usage selection endosymbiosis mutational bias multivariate analysis gene expression Escherichia coli
Address for correspondence and reprints: Jennifer J. Wernegreen, Josephine Bay Paul Center for Comparative Molecular Biology and Evolution, Marine Biological Laboratory, 7 MBL Street, Woods Hole, Massachusetts 02543. E-mail: jwernegreen@mbl.edu
Fig. 1.—a, Positions of putative high- and low-expression genes (closed and open circles, respectively) considered in this study according to their location in the two main axes of COA of amino acid usage in Buchnera. b, Position of amino acids along the two main axes of COA of amino acid usage in Buchnera. Shaded amino acids have significantly different RAAU at high- and low-expression genes (refer to table 1 for significance values). Amino acids that are significantly overrepresented in high-expression genes (marked with light shadow) use relatively GC-rich codons with the exception of Val. Amino acids that are significantly underrepresented in high-expression Buchnera genes (dark shadow) tend to be aromatic and use relatively AT-rich codons. c, Position of amino acids along Axes 2 and 3 of COA of amino acid usage in E. coli. Amino acids that are significantly overrepresented in high-expression genes (see table 2 ) are marked with light shadows. Those significantly underrepresented in high-expression genes are marked with dark shadows. d, Position of amino acids along Axis 3 of COA of E. coli, plotted against frequencies of major tRNA molecules per cell according to Dong, Nilsson and Kurland (1996)
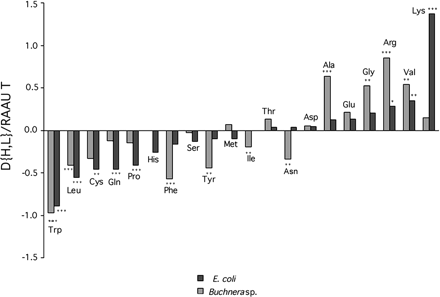
Fig. 2.—Difference in RAAU at high- and low-expression genes (D{H,L}) for Buchnera and E. coli. For comparative purposes, differences were divided by RAAU of all genes considered for each genome, respectively (“RAAU T,” see values in tables 1 and 2 ). Amino acids are ordered by E. coli values for the normalized statistic. Asterisks indicate significant values of D{H,L} according to a permutation test (tables 1 and 2 )
We are grateful to Claude Rispe for helpful discussion about Buchnera genome evolution and Maile C. Neel for statistical advice. We also thank Hilary G. Morrison, Laila Nahum, Michael P. Cummings, Adam Eyre-Walker, and two anonymous reviewers for helpful comments on the manuscript. This work is supported by an award from the NASA Astrobiology Institute (NCC2-1054) to C.P. and awards from the NIH (R01 GM62626-01), NSF (DEB 0089455), and the Josephine Bay Paul and C. Michael Paul Foundation to J.J.W.
References
Akashi H., A. Eyre-Walker,
Akashi H., T. Gojobori,
Aksoy S.,
Andersson S. G. E., C. G. Kurland,
Andersson S. G. E., P. M. Sharp,
Baumann P., L. Baumann, M. A. Clark, M. L. Thao,
Beletskii A., A. S. Bhagwat,
Blattner F. R., G. Plunkett III,, C. A. Bloch, et al. (17 co-authors)
Bracho A. M., D. Martinez-Torres, A. Moya, A. Latorre,
Brewer B.,
Brynnel E. U., C. G. Kurland, N. A. Moran, S. G. E. Andersson,
Clark M. A., L. Baumann, P. Baumann,
Clark M. A., N. A. Moran, P. Baumann,
Craig C. L., R. S. Weber,
de Miranda A. B., F. Alvarez-Valin, K. Jabbari, W. M. Degrave, G. Bernardi,
Dong H., L. Nilsson, C. G. Kurland,
Douglas A. E., W. A. Prosser,
Francino M. P., H. Ochman,
Funk D. J., L. Helbling, J. J. Wernegreen, N. A. Moran,
Gutierrez G., L. Marquez, A. Marin,
Ikemura T.,
Ishikawa H.,
Karlin S., A. M. Campbell, J. Mrazek,
Kyte J., R. F. Doolittle,
Lafay B., A. T. Lloyd, M. J. McLean, K. M. Devine, P. M. Sharp, K. H. Wolfe,
Lobry J. R., C. Gautier,
Mackiewicz P., A. Gierlik, M. Kowalczuk, M. Dudek, S. Cebrat,
McInerney J. O.,
———.
McLean D. L., E. J. Houk,
Mira A., H. Ochman, N. A. Moran,
Moran N.,
Moran N., P. Baumann,
Moran N. A., M. A. Munson, P. Baumann, H. Ishikawa,
Munson M. A., P. Baumann, M. A. Clark, L. Baumann, N. A. Moran, D. J. Voegtlin, B. C. Campbell,
Ohtaka C., H. Ishikawa,
Rispe C., N. A. Moran,
Romero H., A. Zavala, H. Musto,
Sato S., H. Ishikawa,
Selander R. K., D. A. Caugant, T. S. Whittam,
Selosse M., B. Albert, B. Godelle,
Sharp P. M., W. H. Li,
Shigenobu S., H. Watanabe, M. Hattori, Y. Sakaki, H. Ishikawa,
Shigenobu S., H. Watanabe, Y. Sakaki, H. Ishikawa,
Silva F. J., A. Latorre, A. Moya,
Singer G. A. C., D. A. Hickey,
Srivastava A. K., D. Schlessinger,
Unterman B. M., P. Baumann, D. L. McLean,
Wernegreen J. J., N. A. Moran,
———.
Yamao F., Y. Andachi, A. Muto, T. Ikemura, S. Osawa,