-
PDF
- Split View
-
Views
-
Cite
Cite
Andrew C. Clarke, Michael K. Burtenshaw, Patricia A. McLenachan, David L. Erickson, David Penny, Reconstructing the Origins and Dispersal of the Polynesian Bottle Gourd (Lagenaria siceraria), Molecular Biology and Evolution, Volume 23, Issue 5, May 2006, Pages 893–900, https://doi.org/10.1093/molbev/msj092
Close - Share Icon Share
Abstract
The origin of the Polynesian bottle gourd (Lagenaria siceraria), an important crop species in prehistoric Polynesia, has remained elusive. Most recently, a South American origin has been favored as the bottle gourd could have been introduced from this continent with the sweet potato by Polynesian voyagers around A.D. 1,000. To test the hypothesis of an American origin for the Polynesian bottle gourd, we developed seven markers specific to bottle gourd (two chloroplast and five nuclear). The nuclear markers were developed using a new technique where polymorphic inter simple sequence repeat (ISSR) markers are converted into single-locus polymerase chain reaction and sequencing markers—an approach that will be useful for developing markers in other taxa. All seven markers were sequenced in 36 cultivars of bottle gourd from Asia, the Americas, and Polynesia. The results support a dual origin for the Polynesian bottle gourd: the chloroplast markers are exclusively of Asian origin, but the nuclear markers show alleles originating in both the Americas and Asia. Because hybridization of Polynesian bottle gourds with post-European introductions cannot be excluded, ancient DNA from archaeological material will be useful for further elucidating the prehistoric movements of this species in Polynesia. This work has implications not only for the dispersal of the Polynesian bottle gourd but also for the domestication and dispersal of the species as a whole.
Introduction
Human settlement of the remote islands of the Pacific is a relatively recent event that began only ∼5,000 year B.P. with the Austronesian expansion out of Taiwan (Gray and Jordan 2000; Hurles et al. 2003) and is associated with the introduction of many crop species from Southeast Asia and Island Melanesia into Remote Oceania (east of the Solomon Islands) (Pawley 2002; Diamond and Bellwood 2003; Bellwood 2005). Under the standard model, Austronesian voyagers moved through the Philippines and coastal New Guinea and appeared in Island Melanesia and Western Polynesia (Samoa and Tonga) by 3,100 year B.P. where they are often associated with the appearance of the Lapita cultural complex (Kirch 2000). Descendents of the Lapita peoples, the Polynesians, settled in Central Polynesia (the Cook Islands and French Polynesia) by A.D. 800 and the distant apices of the Polynesian triangle by A.D. 600–800 for Hawai'i, A.D. 800–1000 for Easter Island, and A.D. 1000–1200 for New Zealand (Kirch 2000; Anderson and Sinoto 2002; Hurles et al. 2003; see fig. 1). The recent and rapid settlement of the Pacific makes it an interesting region in which to examine human dispersal and the dispersal of domesticated plants and animals.
Prehistoric distribution and dispersal of the bottle gourd (Lagenaria siceraria) in Asia, the Americas, and Oceania. The bottle gourd has been present in the Americas and East Asia since 10,000 and 7,000 year B.P., respectively (Chang 1986; Smith 2005). In the case of the East Asian bottle gourd, it is unclear how far south it spread in prehistory (indicated by dashed line). The Southeast Asian bottle gourd may in fact be a much more recent arrival from India 200 B.C. (Green 2000) and spread only as far east as Vanuatu in prehistory (Yen 1973). The bottle gourd was apparently not present in Western Polynesia (Whistler 1990) (the Bottle Gourd Gap), suggesting that it was not introduced from Asia into Polynesia via human-mediated dispersal (although natural dispersal is still possible). However, the bottle gourd may not have been required in the Gap region as Lapita pottery was widely available as an alternative for containers (distribution of Lapita sites from Kirch [2000]). The bottle gourd was also present in Eastern Polynesia since before A.D. 1,200 (Green 2000) and may have been introduced from the Americas by either natural (floating) or human-mediated dispersal. A human-mediated introduction from South America could have been effected by Polynesian voyagers who departed from Easter Island around A.D. 1,000, sailed to the Peruvian Coast, and returned probably to the Tuamotu Archipelago with the sweet potato (route based on that suggested for the sweet potato by Green [2005]). Similarly, Polynesian voyagers could have introduced the bottle gourd from North America via a return sailing trip from Hawai'i to the Californian Channel Islands around A.D. 400–800 (Jones and Klar 2005), although this hypothesis is yet to be tested.
The reconstruction of prehistoric (pre-European) movements of peoples in the Pacific has benefited significantly from molecular studies of human mitochondrial DNA (e.g., Redd et al. 1995; Lum and Cann 2000; Friedlaender et al. 2005) and Y chromosomes (e.g., Hage and Marck 2003). Prehistoric human mobility has been further resolved by molecular studies on the commensal plants (e.g., Zerega, Ragone, and Motley 2004), animals (e.g., Matisoo-Smith and Robins 2004; Larson et al. 2005), and pathogens (Yanagihara et al. 2002) that accompanied humans on their voyages. We are employing methods similar to these to reconstruct the origins and dispersal of the Polynesian bottle gourd.
The bottle gourd (Lagenaria siceraria [Mol.] Standl.) is a diploid, self-compatible monoecious annual in the Cucurbitaceae. It is known almost exclusively in cultivation but is probably native to Africa (Heiser 1979), where a wild Zimbabwean population has recently been discovered (Decker-Walters et al. 2004). Two morphologically distinct subspecies of bottle gourd are recognized: L. siceraria ssp. siceraria (the African and American/New World gourds) and L. siceraria ssp. asiatica (the Asian gourds) (Kobiakova 1930; Heiser 1973a). The bottle gourd is predominantly grown for its fruit which, when dry, form a woody rind (exocarp) that is used mostly for the manufacture of containers (for water and food) and also musical instruments (drums and flutes), fishing floats, and apparel such as penis sheaths/phallocrypts (Heiser 1979).
Archaeological evidence shows that domesticated bottle gourds were present in the Americas (New World) by at least 9,900 year B.P. (Piperno, Andres, and Stothert 2000; Smith 2005; fig. 1), making it one of the first species domesticated by humans. Based on morphological analyses, the American gourds have been classified as L. siceraria ssp. siceraria (Heiser 1973a), consistent with an African origin, and may have arrived by floating; domesticated bottle gourds contain still-viable seeds even after floating in sea water for more than 7 months (Whitaker and Carter 1954). The Asian bottle gourd, L. siceraria ssp. asiatica, appears as a domesticated species in China and Japan from 7,000 year B.P. (Chang 1986; Imamura 1996; Habu et al. 2001; fig. 1).
Linguistic evidence and dated archaeological specimens suggest that the bottle gourd has been present in Eastern Polynesia since before A.D. 1,200 (see Leach 1984; Green 1998, 2000; for Hawai'i see Burney et al. 2001; for New Zealand see Horrocks et al. 2002). The dispersal of the bottle gourd within Polynesia, where it was widely distributed, was certainly human mediated—consistent with linguistic evidence (Green 2000), Polynesian oral history, and the considerable importance of the bottle gourd as a crop species (Best 1976 [1925]).
Morphological analyses of both contemporary and archaeological (prehistoric) New Zealand Māori bottle gourd cultivars have produced conflicting results regarding their origins, with some samples exhibiting features typical of the Asian subspecies (Heiser 1973a, 1973b) and others with features intermediate between the Asian and American subspecies (Maingay 1985; Burtenshaw 1999), suggesting at least some American contribution to the Polynesian bottle gourds.
The Polynesian bottle gourd may have been dispersed from either or both continents, and this dispersal may have been effected naturally (by floating) or by humans. If the Polynesian bottle gourd is Asian in origin, it may have arrived by natural dispersal, as linguistic evidence (Ross 1996; Green 2000) suggests it did not reach Island Southeast Asia and Melanesia until after the settlement of Remote Oceania and therefore could not have been carried by Lapita peoples into Polynesia. This is further supported by the Bottle Gourd Gap—an area covering Fiji and Western Polynesia (Samoa, Tonga, and Niue) in which it appears there was no prehistoric presence of the bottle gourd (Whistler 1990; fig. 1).
Both natural and human-mediated dispersals of the bottle gourd from South America into Polynesia are possible, although the latter hypothesis is favored by some authors (Green 2000, 2005) and is consistent with the evidence which indicates at least two episodes of Polynesian contact with the Americas. Most famously is the presence of the South American sweet potato (Ipomoea batatas) in prehistoric Polynesia (Yen 1974; Hather and Kirch 1991; Hurles et al. 2003) which, supported by linguistic and other evidence (Green 2005), may have been introduced into Polynesia by a return sailing trip from Easter Island to South America around A.D. 1,000–1,100 (Green 2005; fig. 1). Very recently, a prehistoric Polynesian voyage to the Channel Islands off the southern Californian coast has also been proposed after distinctive Polynesian-style sewn-plank canoes and two-piece bone fishhooks were identified in the archaeological record of these islands, along with an apparent Polynesian borrowing of the word for the sewn-plank canoes (Jones and Klar 2005). The bottle gourd, as a species distributed throughout the Americas during prehistory, could have been collected from either North or South America on such a voyage (fig. 1).
In this paper, we employ a set of molecular markers to explicitly test the hypothesis of an American origin for the Polynesian bottle gourd. The morphological evidence suggests at least some contribution from Asia—consequently we will examine the extent to which the Polynesian accessions may represent a polyphyletic assemblage which has arisen from multiple introductions of the bottle gourd from both continents combined with postestablishment gene flow.
Materials and Methods
A total of 38 cultivars of bottle gourd were obtained: 13 from Asia, 15 from the Americas/New World, 8 from New Zealand (Polynesia), and 2 from Africa. See table 1 for the locations of the cultivars and Table S1 (Supplementary Material online) for further details. The historic period has seen the all but complete loss of pre-European gourds in Polynesia (Dodge 1943), making it difficult to obtain samples to test hypotheses about the dispersal of the species. The eight New Zealand Māori accessions are believed to be genuine pre-European introductions, with provenance based on the areas from which these samples have been collected (isolated Māori communities) or discussions with the people from whom seed was obtained. The New Zealand accessions are currently a proxy for Eastern Polynesia as a whole, but future research, including ancient DNA (aDNA) analysis, must be undertaken to obtain samples from other Polynesian islands. Of the remaining 30 accessions, 27 were obtained from Charles Heiser (Indiana University, Bloomington) and are described in Heiser (1973a). None of the Heiser accessions are directly derived from commercial seed companies, and most should represent landraces of the region from which they were obtained.
Bottle Gourd (Lagenaria siceraria) Sample Details
Region . | Location (cultivar) . | Cultivar Code . |
|---|---|---|
| Asia | Indonesia | 020(1)Ind |
| Asia | Indonesia | 020(2)Ind |
| Asia | India | 061Ind |
| Asia | India | 101Ind |
| Asia | Malaysia | 111Mal |
| Asia | Philippines | 149Phi |
| Asia | Philippines | 157Phi |
| Asia | India | 159Ind |
| Asia | Malaysia | 161Mal |
| Asia | Philippines | 174Phi |
| Asia | Thailand | 188Tha |
| Asia | Yuwa-machi, Japan | 195 |
| Asia | Akita, Japan | AK |
| Polynesia | Bay of Plenty, New Zealand | 183 |
| Polynesia | New Zealand (Bottle Ruku) | BR |
| Polynesia | New Zealand (Gourd “A”) | GA |
| Polynesia | New Zealand (Gourd “D”) | GD |
| Polynesia | New Zealand (Māori Gourd [1973–1974]) | MA |
| Polynesia | New Zealand (Māori Gourd) | MG |
| Polynesia | New Zealand (New Zealand Bottle) | NB |
| Polynesia | New Zealand (Nga Puhi) | NP |
| Americas | Costa Rica | 006Cos |
| Americas | Ecuador | 027Ecu |
| Americas | Argentina | 035BArg |
| Americas | Peru | 036(1)Per |
| Americas | Peru | 036(2)Per |
| Americas | Peru | 037Per |
| Americas | Brazil | 051Bra |
| Americas | Costa Rica | 059Cos |
| Americas | Mexico | 079Mex |
| Americas | Hopi, Arizona, United States | 093Hop |
| Americas | Mexico | 152Mex |
| Americas | Mexico | 153Mex |
| Americas | Peru | 195Per |
| Americas | Brazil | 315Bra |
| Americas | Brazil | 407Bra |
| Africa | Madagascar | 291Mad |
| Africa | Africa (Maranka) | MR |
Region . | Location (cultivar) . | Cultivar Code . |
|---|---|---|
| Asia | Indonesia | 020(1)Ind |
| Asia | Indonesia | 020(2)Ind |
| Asia | India | 061Ind |
| Asia | India | 101Ind |
| Asia | Malaysia | 111Mal |
| Asia | Philippines | 149Phi |
| Asia | Philippines | 157Phi |
| Asia | India | 159Ind |
| Asia | Malaysia | 161Mal |
| Asia | Philippines | 174Phi |
| Asia | Thailand | 188Tha |
| Asia | Yuwa-machi, Japan | 195 |
| Asia | Akita, Japan | AK |
| Polynesia | Bay of Plenty, New Zealand | 183 |
| Polynesia | New Zealand (Bottle Ruku) | BR |
| Polynesia | New Zealand (Gourd “A”) | GA |
| Polynesia | New Zealand (Gourd “D”) | GD |
| Polynesia | New Zealand (Māori Gourd [1973–1974]) | MA |
| Polynesia | New Zealand (Māori Gourd) | MG |
| Polynesia | New Zealand (New Zealand Bottle) | NB |
| Polynesia | New Zealand (Nga Puhi) | NP |
| Americas | Costa Rica | 006Cos |
| Americas | Ecuador | 027Ecu |
| Americas | Argentina | 035BArg |
| Americas | Peru | 036(1)Per |
| Americas | Peru | 036(2)Per |
| Americas | Peru | 037Per |
| Americas | Brazil | 051Bra |
| Americas | Costa Rica | 059Cos |
| Americas | Mexico | 079Mex |
| Americas | Hopi, Arizona, United States | 093Hop |
| Americas | Mexico | 152Mex |
| Americas | Mexico | 153Mex |
| Americas | Peru | 195Per |
| Americas | Brazil | 315Bra |
| Americas | Brazil | 407Bra |
| Africa | Madagascar | 291Mad |
| Africa | Africa (Maranka) | MR |
NOTE.—Full details (including provenance) of bottle gourd samples are provided in Table S1 (Supplementary Material online).
Bottle Gourd (Lagenaria siceraria) Sample Details
Region . | Location (cultivar) . | Cultivar Code . |
|---|---|---|
| Asia | Indonesia | 020(1)Ind |
| Asia | Indonesia | 020(2)Ind |
| Asia | India | 061Ind |
| Asia | India | 101Ind |
| Asia | Malaysia | 111Mal |
| Asia | Philippines | 149Phi |
| Asia | Philippines | 157Phi |
| Asia | India | 159Ind |
| Asia | Malaysia | 161Mal |
| Asia | Philippines | 174Phi |
| Asia | Thailand | 188Tha |
| Asia | Yuwa-machi, Japan | 195 |
| Asia | Akita, Japan | AK |
| Polynesia | Bay of Plenty, New Zealand | 183 |
| Polynesia | New Zealand (Bottle Ruku) | BR |
| Polynesia | New Zealand (Gourd “A”) | GA |
| Polynesia | New Zealand (Gourd “D”) | GD |
| Polynesia | New Zealand (Māori Gourd [1973–1974]) | MA |
| Polynesia | New Zealand (Māori Gourd) | MG |
| Polynesia | New Zealand (New Zealand Bottle) | NB |
| Polynesia | New Zealand (Nga Puhi) | NP |
| Americas | Costa Rica | 006Cos |
| Americas | Ecuador | 027Ecu |
| Americas | Argentina | 035BArg |
| Americas | Peru | 036(1)Per |
| Americas | Peru | 036(2)Per |
| Americas | Peru | 037Per |
| Americas | Brazil | 051Bra |
| Americas | Costa Rica | 059Cos |
| Americas | Mexico | 079Mex |
| Americas | Hopi, Arizona, United States | 093Hop |
| Americas | Mexico | 152Mex |
| Americas | Mexico | 153Mex |
| Americas | Peru | 195Per |
| Americas | Brazil | 315Bra |
| Americas | Brazil | 407Bra |
| Africa | Madagascar | 291Mad |
| Africa | Africa (Maranka) | MR |
Region . | Location (cultivar) . | Cultivar Code . |
|---|---|---|
| Asia | Indonesia | 020(1)Ind |
| Asia | Indonesia | 020(2)Ind |
| Asia | India | 061Ind |
| Asia | India | 101Ind |
| Asia | Malaysia | 111Mal |
| Asia | Philippines | 149Phi |
| Asia | Philippines | 157Phi |
| Asia | India | 159Ind |
| Asia | Malaysia | 161Mal |
| Asia | Philippines | 174Phi |
| Asia | Thailand | 188Tha |
| Asia | Yuwa-machi, Japan | 195 |
| Asia | Akita, Japan | AK |
| Polynesia | Bay of Plenty, New Zealand | 183 |
| Polynesia | New Zealand (Bottle Ruku) | BR |
| Polynesia | New Zealand (Gourd “A”) | GA |
| Polynesia | New Zealand (Gourd “D”) | GD |
| Polynesia | New Zealand (Māori Gourd [1973–1974]) | MA |
| Polynesia | New Zealand (Māori Gourd) | MG |
| Polynesia | New Zealand (New Zealand Bottle) | NB |
| Polynesia | New Zealand (Nga Puhi) | NP |
| Americas | Costa Rica | 006Cos |
| Americas | Ecuador | 027Ecu |
| Americas | Argentina | 035BArg |
| Americas | Peru | 036(1)Per |
| Americas | Peru | 036(2)Per |
| Americas | Peru | 037Per |
| Americas | Brazil | 051Bra |
| Americas | Costa Rica | 059Cos |
| Americas | Mexico | 079Mex |
| Americas | Hopi, Arizona, United States | 093Hop |
| Americas | Mexico | 152Mex |
| Americas | Mexico | 153Mex |
| Americas | Peru | 195Per |
| Americas | Brazil | 315Bra |
| Americas | Brazil | 407Bra |
| Africa | Madagascar | 291Mad |
| Africa | Africa (Maranka) | MR |
NOTE.—Full details (including provenance) of bottle gourd samples are provided in Table S1 (Supplementary Material online).
Viable seeds were germinated and young leaf tissue was used for extracting DNA. For nonviable seeds, DNA was extracted directly from the dead embryo. DNA extractions were performed using a hexadecyltrimethylammonium bromide (CTAB) method (J. J. Doyle and J. L. Doyle 1990) or the DNeasy Plant Mini Kit (Qiagen, Valencia, Calif.).
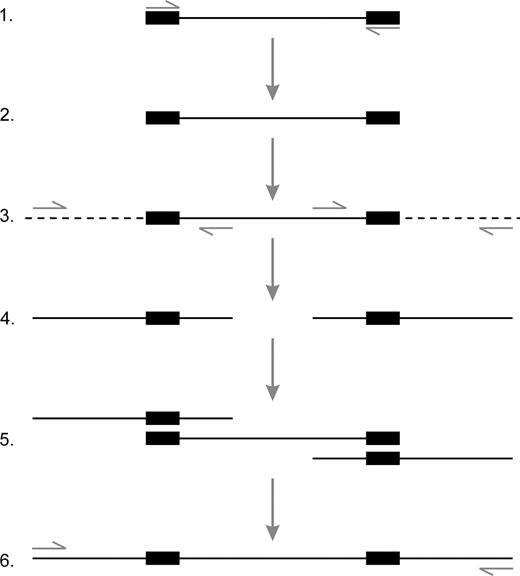
Detecting genetic variation in very recently diverged plants, especially domesticated species where genetic bottlenecks have further reduced variation (see Salamini et al. 2002), is extremely difficult and requires the development of species-specific microsatellite or sequence-characterized amplified region (SCAR) markers. In this project, nuclear DNA (nDNA) SCAR markers were developed from inter simple sequence repeat (ISSR) fingerprints using a novel method outlined in figure 2 (and which will be described in detail in a separate publication). In addition, two polymorphic chloroplast DNA (cpDNA) markers were used: trnC-trnD from Lee and Wen (2004) and trnS-trnG. The latter was discovered by using different combinations of primers described in Chung and Staub (2003).
Development of ISSR-derived SCAR markers. Boxed areas indicate the terminal simple sequence repeats (SSRs) of the inter-SSR (ISSR) product. Small arrows indicate PCR primers. 1. ISSR fingerprints are generated using PCR (Ziętkiewicz, Rafalski, and Labuda 1994) and separated by electrophoresis on a 5% acrylamide gel. 2. Polymorphic ISSR bands are excised from the gel, reamplified by PCR, cloned, and sequenced. 3. In a novel approach to increase the amount of useful sequence data, regions flanking each terminal SSR are amplified from genomic DNA using thermal asymmetric interlaced (TAIL) PCR (Liu and Whittier 1995) (primer sequences are provided in Table S2, Supplementary Material online). 4. TAIL PCR products are then direct sequenced. 5. Sequence data from the ISSR and its TAILed flanking regions are used to construct a contig spanning the entire locus. 6. SCAR marker primers are designed in the flanking regions and are used to amplify the polymorphic locus, including the SSRs, in any cultivar of bottle gourd.
The ISSR-derived SCAR markers and chloroplast markers (Table S3, Supplementary Material online) were successfully amplified in all 38 cultivars of bottle gourd. Each polymerase chain reaction (PCR) consisted of 1× PCR buffer, 250 μM of each deoxynucleoside triphosphate 1 M betaine, 0.5 μM L primer, 0.5 μM R primer (see Table S3 for primer sequences, Supplementary Material online), 1 U Taq DNA polymerase, and ∼1 ng of genomic DNA in a total volume of 20 μl. PCR for the ISSR SCAR markers was carried out using the following program: denaturation at 94°C for 2 min; 35 cycles of denaturation at 94°C for 30 s, annealing for 1 min, extension at 72°C for 1 min; and final extension at 72°C for 5 min. Chloroplast marker PCRs were identical except that the extension step was 68°C for 3 min for trnC-trnD, 68°C for 1 min 30 s for trnS-trnG, and the final extension steps were 68°C for 5 min for both cpDNA markers. The annealing temperatures for all markers are shown in Table S3 (Supplementary Material online).
PCR products were sequenced using the BigDye Terminator v3.1 Cycle Sequencing Kit and separated on an ABI3730 Genetic Analyzer (both Applied Biosystems, Foster City, Calif.) according to the manufacturer's protocols by the Allan Wilson Centre Genome Service, Massey University. The sequencing primers are described in Table S3 (Supplementary Material online). Due to the low level of variation, known positions of useful polymorphisms and high-quality sequence data, PCR products were sequenced in one direction only (electropherograms available upon request). Individuals which were heterozygous (either for single-nucleotide polymorphisms [SNPs] or length polymorphisms) were cloned and sequenced.
Results
A total of 5.7 kb of sequence data were generated across seven markers for each of the 38 accessions of bottle gourd. All sequence data have been deposited in GenBank (accession numbers DQ281822–DQ282115). Data were aligned and all variable sites, of which there are 49, are shown in Table S4 (Supplementary Material online).
A wide variety of analysis methods were explored in a comprehensive search to obtain the best method for analyzing the data (e.g., various tree-building methods, network building, and statistical analysis), and it was decided that the results were most clearly displayed as genotype frequencies presented as simple pie charts, supplemented with a Spectronet network diagram constructed from all informative sites (Huber et al. 2002; Fig. S1, Supplementary Material online).
Pie charts were constructed to show haplotype frequencies (cpDNA) or genotype frequencies (nDNA) for each marker in each of the three geographical regions (Asia, Polynesia and the Americas) (fig. 3). Data for the two chloroplast markers were concatenated (see fig. 3 legend). For the nDNA markers, the method of displaying genotype frequencies, rather than haplotype (allele) frequencies, allows figure 3 to retain information about each individual (i.e., whether it is a homozygote or heterozygote).
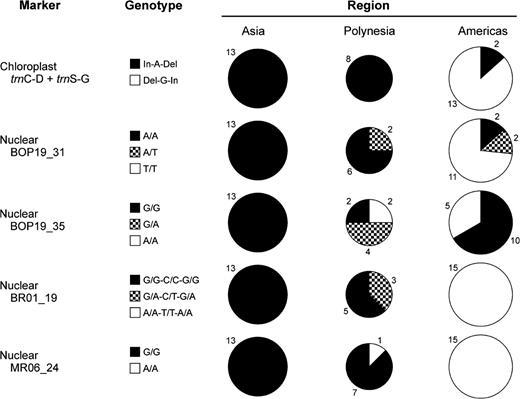
Genotype frequencies for 36 cultivars of bottle gourd (Lagenaria siceraria). Genotype frequencies for 1 chloroplast locus and 4 nuclear markers in 13 Asian, 8 Polynesian, and 15 American bottle gourd individuals. Total area of each pie is proportional to the number of individuals sampled from that region. Data from the chloroplast markers trnC-D and trnS-G are concatenated as the polymorphisms are congruent between markers, and the chloroplast represents a single “locus.” The number adjacent to each slice indicates the number of individuals comprising that slice. For the chloroplast, solid (black or white) pie slices indicate different haplotypes. For the nDNA markers, solid pie slices indicate individuals which are homozygous for a haplotype (allele), and checkered slices indicate individuals which are heterozygous for both haplotypes (alleles). See the main text for an explanation of the genotype notation.
cpDNA (trnC-D and trnS-G)
Three variable sites are observed in the chloroplast data: two indels and a G/A SNP. The polymorphisms are always concordant so that an individual is always of the In-A-Del haplotype or the Del-G-In haplotype. The In-A-Del (Asian) and Del-G-In (American) haplotypes are fixed on either side of the Pacific except for two American individuals (one each from Brazil and Argentina, see Table S4, Supplementary Material online) that possess the Asian haplotype (fig. 3). These gourds may represent a prehistoric introduction from Asia (or even Africa), but a modern introduction is also quite likely, especially given the ∼500 year presence of the Spanish and Portuguese in both Southeast Asia (where gourds could have been collected) and South America (where gourds could have been dispersed). In any case, these anomalous Asian haplotypes in South America were not observed on the west coast (from where the Polynesian bottle gourd would most likely originate) and therefore are probably not implicated in the origin of the Polynesian bottle gourd. All eight Polynesian individuals possess the Asian chloroplast haplotype. The trnC-D and trnS-G markers were also sequenced in two outgroups, wax gourd (Benincasa hispida) and watermelon (Citrullus lanatus), which both possess the “Asian” haplotype, consistent with the Asian haplotype being ancestral in bottle gourd and the African derived (Table S4, Supplementary Material online).
BOP19_31 and BOP19_35
Both markers possess an informative SNP where one allele is present in Asia, Polynesia, and the Americas and the other allele in Polynesia and the Americas only.
BR01_19
Three informative SNPs are observed and are concordant so that an individual is always of the G-C-G haplotype and/or the A-T-A haplotype. Asia is fixed for the G-C-G haplotype so that all individuals possess two copies of this allele (denoted as the G/G-C/C-G/G genotype). The Americas, however, are fixed for the A-T-A haplotype so that all individuals are of the A/A-T/T-A/A genotype. The Polynesian individuals possess either the Asian genotype or are heterozygous for both the Asian G-C-G haplotype and the American A-T-A haplotype; this heterozygous genotype is denoted as G/A-C/T-G/A. Cloning of heterozygous individuals showed there was no recombination between these two haplotypes. BR01_19 was the only marker that had a significant match to any sequence in GenBank (a putative protein kinase in Arabidopsis thaliana).
MR06_24
One informative SNP is observed that again is fixed in Asia (G/G genotype) and in the Americas (A/A genotype) but has both the G and A alleles present in Polynesia.
BOP19_27
The marker BOP19_27 was excluded from the analysis as although it possesses a 14-bp indel and a variable dinucleotide microsatellite that will be useful for studying diversity within Asia, it is not informative in terms of the origin of the Polynesian bottle gourd (see Table S4, Supplementary Material online).
The cpDNA markers and the BR01_19 and MR06_24 nuclear markers are fixed for different alleles on either side of the Pacific (fig. 3), clearly separating the Asian and American bottle gourds (congruent with the subspecies taxonomy), and allowing us to infer from which side of the Pacific the Polynesian bottle gourds originate.
Discussion
An Asian origin for the Polynesian bottle gourd is supported by the presence of the Asian chloroplast haplotype in all New Zealand bottle gourds. Nuclear markers, however, indicate that there is also a significant genetic contribution from the Americas and that the New Zealand bottle gourds are the result of hybridizations between cultivars from both continents (also supported by network analysis, see Fig. S1, Supplementary Material online). New Zealand individuals which possess American alleles do not possess them at all markers (see Table S4, Supplementary Material online), suggesting that although some of the New Zealand accessions are hybrids between Asian and American cultivars, these individuals are not the first generation (F1) of such a cross.
Both the chloroplast and nuclear data from the New Zealand accessions strongly support a partly Asian origin for the Polynesian bottle gourd. Although human-mediated dispersal is not concordant with both the apparently late introduction of the bottle gourd into Southeast Asia (∼200 B.C.) and the Bottle Gourd Gap, it is premature to exclude human-mediated dispersal of the Polynesian bottle gourd from Asia (fig. 1). Firstly, the bottle gourd was present in Taiwan during the Austronesian expansion out of this area ∼5,000 year B.P. (Bellwood 1997), so it was available to be taken further south as new islands were settled. Secondly, the “negative evidence” of the lack of bottle gourd in archaeological sites from Melanesia does not prove its absence as there is no body of wetland archaeobotanical research from this area (the excavations from highland New Guinea are exceptional). Thirdly, the apparent Bottle Gourd Gap region is nested within the distribution of known Lapita sites (fig. 1), where pottery was abundant and the bottle gourd may not have been required as a container. Further east however, in Eastern Polynesia where pottery was not manufactured (Kirch 2000), the bottle gourd would have been an indispensable vessel for water and food.
The nuclear data from the New Zealand accessions are consistent with a partly American origin for the Polynesian bottle gourd although, as for the Asian contribution, it cannot be resolved whether this dispersal was natural or human mediated. The latter is certainly possible if it is established that Polynesians sailed to South America (and collected the sweet potato [Green 2005]) or the Channel Islands (Jones and Klar 2005) as the bottle gourd could have been collected on such a voyage.
In conclusion, the Polynesian bottle gourd may have a dual origin, with the chloroplast data from the New Zealand accessions indicating a partly Asian origin and the nuclear data supporting genetic contributions from both Asia and the Americas (consistent with the morphological data). It is possible that the New Zealand bottle gourds used for this study have been contaminated by gene flow (especially pollen-mediated) from post-European contact introductions. Therefore, our tentative conclusion of a dual origin should be further tested by aDNA analysis of protohistoric herbarium and anthropological bottle gourd material, as well as prehistoric archaeological material from sites throughout Polynesia, Southeast Asia, and the Americas. The high-copy number trnC-D and trnS-G chloroplast markers will be most amenable to this but, as evidenced by this study, the nuclear DNA is also required. Additional sampling within Asia may also resolve whether the Asian alleles in the Polynesian bottle gourd are derived from an old ∼5,000 year B.P. Austronesian (Chinese-Taiwanese) lineage or the more recent 200 B.C. Southeast Asian (Indian-Indonesian) lineage.
The new markers described here will be also useful for addressing other outstanding questions regarding the domestication and dispersal of the bottle gourd, including the origin of the New Guinea phallocrypt gourds (which are morphologically closer to the African subspecies [Heiser 1973b]), levels of genetic variation in Africa, and the relationship between the wild Zimbabwean bottle gourd and the domesticated forms.
Data from the chloroplast and two of the nuclear markers (BR01_19 and MR06_24) support the Asian subspecies and African/American subspecies each comprising a monophyletic group. However, the large number of genetic differences between the subspecies are inconsistent with their recent (i.e., postdomestication) divergence, and the cpDNA data from the outgroup species are inconsistent with the prevailing hypothesis that Asian cultivars are derived from African cultivars (instead the Asian chloroplast haplotype appears basal). Although a rapid “subspecies” divergence may postdate a single domestication event (followed by a change in allele frequencies due to migration, genetic drift, and selection events), a perhaps simpler explanation is divergence of the subspecies predating independent domestication of the bottle gourd in Asia and Africa. It is now clear that a number of crop and livestock species have been domesticated more than once (Diamond 2002), and the hypothesis of multiple domestications of the bottle gourd should be tested by analyzing variation between domesticated and supposed wild plants—the latter have only been documented in Zimbabwe (Decker-Walters et al. 2004), and more effort should be focused on locating wild plants in Asia.
Yoko Satta, Associate Editor
For samples we thank Charles Heiser, John Palmer, Richard Cross, Peter Matthews, and Steve Lewthwaite. For assistance with growing the bottle gourd plants we thank Ruth Morrison. Roger Green helped to develop the hypotheses and provided general guidance. Barbara Holland, Peter Lockhart, and Tim White assisted with the analysis. The authors would like to thank Jonathan Procter and Rangitaane O Manawatu for their support. D.L.E. and A.C.C. would like to thank Bruce Smith and Noreen Tuross for supporting work completed at the Laboratories of Analytical Biology, Smithsonian Institution, Washington, D.C. For insightful comments on the manuscript we are grateful to Peter Matthews and two anonymous reviewers. Funding was provided by the Allan Wilson Centre for Molecular Ecology and Evolution, New Zealand. A.C.C. was supported by a Massey University Doctoral Scholarship.
References
Anderson, A., and Y. Sinoto.
Bellwood, P. S.
Best, E.
Burney, D. A., H. F. James, L. Pigott Burney et al. (12 co-authors).
Chung, S.-M., and J. E. Staub.
Decker-Walters, D. S., M. Wilkins-Ellert, S.-M. Chung, and J. E. Staub.
Diamond, J.
Diamond, J., and P. Bellwood.
Dodge, E. S.
Friedlaender, J., T. Schurr, F. Gentz et al. (12 co-authors).
Gray, R. D., and F. M. Jordan.
Green, R. C.
———.
———.
Habu, J., M. Kim, M. Katayama, and H. Komiya.
Hage, P., and J. Marck.
Hather, J., and P. V. Kirch.
Heiser, C. B.
Horrocks, M., M. D. Jones, R. E. Beever, and D. G. Sutton.
Huber, K. T., M. Langton, D. Penny, V. Moulton, and M. Hendy.
Hurles, M. E., E. Matisoo-Smith, R. D. Gray, and D. Penny.
Jones, T. L., and K. A. Klar.
Kirch, P. V.
Larson, G., K. Dobney, U. Albarella et al. (13 co-authors).
Leach, H.
Lee, C., and J. Wen.
Liu, Y.-G., and R. F. Whittier.
Lum, J. K., and R. L. Cann.
Maingay, J.
Matisoo-Smith, E., and J. H. Robins.
Pawley, A.
Piperno, D. R., T. C. Andres, and K. E. Stothert.
Redd, A. J., N. Takezaki, S. T. Sherry, S. T. McGarvey, A. S. M. Sofro, and M. Stoneking.
Ross, M.
Salamini, F., H. Özkan, A. Brandolini, R. Schäfer-Pregl, and W. Martin.
Smith, B. D.
Whitaker, T. W., and G. F. Carter.
Yanagihara, R., V. R. Nerurkar, I. Scheirich et al. (12 co-authors).
Yen, D. E.
———.
Zerega, N. J. C., D. Ragone, and T. J. Motley.
Author notes
*Allan Wilson Centre for Molecular Ecology and Evolution, Massey University, Palmerston North, New Zealand; †Institute of Molecular BioSciences, Massey University, Palmerston North, New Zealand; ‡Natural Resources Centre, The Open Polytechnic of New Zealand, Lower Hutt, New Zealand; and §Laboratories of Analytical Biology, National Museum of Natural History, Smithsonian Institution, Washington, DC



![Prehistoric distribution and dispersal of the bottle gourd (Lagenaria siceraria) in Asia, the Americas, and Oceania. The bottle gourd has been present in the Americas and East Asia since 10,000 and 7,000 year B.P., respectively (Chang 1986; Smith 2005). In the case of the East Asian bottle gourd, it is unclear how far south it spread in prehistory (indicated by dashed line). The Southeast Asian bottle gourd may in fact be a much more recent arrival from India 200 B.C. (Green 2000) and spread only as far east as Vanuatu in prehistory (Yen 1973). The bottle gourd was apparently not present in Western Polynesia (Whistler 1990) (the Bottle Gourd Gap), suggesting that it was not introduced from Asia into Polynesia via human-mediated dispersal (although natural dispersal is still possible). However, the bottle gourd may not have been required in the Gap region as Lapita pottery was widely available as an alternative for containers (distribution of Lapita sites from Kirch [2000]). The bottle gourd was also present in Eastern Polynesia since before A.D. 1,200 (Green 2000) and may have been introduced from the Americas by either natural (floating) or human-mediated dispersal. A human-mediated introduction from South America could have been effected by Polynesian voyagers who departed from Easter Island around A.D. 1,000, sailed to the Peruvian Coast, and returned probably to the Tuamotu Archipelago with the sweet potato (route based on that suggested for the sweet potato by Green [2005]). Similarly, Polynesian voyagers could have introduced the bottle gourd from North America via a return sailing trip from Hawai'i to the Californian Channel Islands around A.D. 400–800 (Jones and Klar 2005), although this hypothesis is yet to be tested.](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/mbe/23/5/10.1093_molbev_msj092/1/m_molbiolevolmsj092f01_4c.jpeg?Expires=1716326241&Signature=MoyBU-0~LHV1XetbtrotrT8FWvqjSFkZ20KBcoI58C-XW819fRsN~aQgAZDNS~7NWChSWjF5Vb1q7AZt6xrXJ4xKpVYQHnkw2SEFbkrfck9qjuyzE16clNGTvzwBEkf7jbnbGCW9N51TlikDfTHNvpzSj1gL0f9ESTah9WbdKiwIF2dpkZEoyer837h0CriUEexEkizkSXIUJseZtoOLmP-849Q8YHk9cl71tkVBX2FiXd12U01QN9hlS8SKo7EjIZyuAtzaza~uGRpf~Kpa7XULcAa5kS0S0L0qNBq4qfScYA0~JSVO8Y2KGSxbvW29l1RbtuoOM1nHpguK0Ucx0g__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)