-
PDF
- Split View
-
Views
-
Cite
Cite
Kosuke Hashimoto, Susumu Goto, Shin Kawano, Kiyoko F. Aoki-Kinoshita, Nobuhisa Ueda, Masami Hamajima, Toshisuke Kawasaki, Minoru Kanehisa, KEGG as a glycome informatics resource, Glycobiology, Volume 16, Issue 5, May 2006, Pages 63R–70R, https://doi.org/10.1093/glycob/cwj010
Close - Share Icon Share
Abstract
Bioinformatics approaches to carbohydrate research have recently begun using large amounts of protein and carbohydrate data. In this field called glycome informatics, the foremost necessity is a comprehensive resource for genome-scale bioinformatics analysis of glycan data. Although the accumulation of experimental data may be useful as a reference of biological and biochemical information on carbohydrates, this is insufficient for bioinformatics analysis. Thus, we have developed a glycome informatics resource (http://www.genome.jp/kegg/glycan/) in KEGG (Kyoto Encyclopedia of Genes and Genomes), an integrated knowledge base of protein networks, genomic information, and chemical information. This review describes three noteworthy features: (1) GLYCAN, a database of carbohydrate structures; (2) glycan-related pathways; and (3) Composite Structure Map (CSM), a map illustrating all possible variations of carbohydrate structures within organisms. GLYCAN includes two useful tools: an intuitive drawing tool called KegDraw, and an efficient glycan search and alignment tool called KEGG Carbohydrate Matcher (KCaM). KEGG’s glycan biosynthesis and metabolism pathways, integrating carbohydrate structures, proteins, and reactions, are also a pivotal resource. CSM is constructed as a bridge between carbohydrate functions and structures. CSM is able to display, for example, expression data of glycosyltransferases in a compact manner. In all the KEGG resources, various objects including KEGG pathways, chemical compounds, as well as carbohydrate structures are commonly represented as graphs, which are widely studied and utilized in the computer science field.
Accepted on July 8, 2005
Introduction
Sequences of genes and proteins have been determined on a grand scale by systematic experimental methods through genome projects. On the other hand, carbohydrate-structure determination has had many challenges and difficulties, mainly due to the complexes they form with other molecules and also to the structural variations resulting from biosynthetic pathways. This complex carbohydrate structure also requires the development of new computational methods for analysis (von der Lieth et al., 2004). Because of the recent improvement of experimental techniques such as mass spectrometry, nuclear magnetic resonance (NMR), and knockout mice analysis, much knowledge about carbohydrate structures and functions has been accumulated (Dell and Morris, 2001; Kogelberg et al., 2003; Miyakis et al., 2004). With this background, bioinformatics approaches to carbohydrate research have also recently begun using a large amount of protein and carbohydrate data. Some examples include studies on glycosylation sites on proteins (Ben-Dor et al., 2004; Petrescu et al., 2004; Julenius et al., 2005). In the dawn of full-scale bioinformatics research on glycans, which we call glycome informatics, the foremost necessity is a comprehensive resource for glycan data. To our knowledge, there has been no such glycan resource for genome-scale bioinformatics analysis.
Exactly what types of data resources are needed for glycome informatics? Although data of experimental results is useful as a reference of biological and biochemical information on carbohydrates, simply accumulating this data is insufficient for bioinformatics analysis. Within an organism, carbohydrates and other chemical compounds that are indirectly generated from the genome interact with genes and proteins, which are directly generated. It is thus necessary to amalgamate all of this information together for understanding a cell or an organism and for predicting complex cellular processes and organism behaviors at a higher level. From this perspective, we have been developing KEGG (Kyoto Encyclopedia of Genes and Genomes), which is an integrated knowledge base of protein networks, genomic information and chemical information (Kanehisa et al., 2004). Here, we present the glycome informatics resource recently constructed and developed in KEGG, including GLYCAN, a database of carbohydrate structures, and Composite Structure Map (CSM), a map illustrating the variations of carbohydrate structures.
The foundation of a glycan resource is its database of carbohydrate structures. Some currently and previously available carbohydrate databases include both public and commercial databases (Marchal et al., 2003). Some public databases include CarbBank (Doubet et al., 1989; Doubet and Albersheim, 1992) and SWEET-DB (Loss et al., 2002), and known commercial databases include GlycoSuiteDB (Cooper et al., 2003) and the Glycomics database by GlycoMinds (http://www.glycominds.com). CarbBank, containing published structures of oligosaccharides and glycoconjugates as a flat file, was developed during the 1980s and 1990s and had finally grown to over 45, 000 records. But the project was terminated due to lack of funding in 1999. SWEET-DB is a web-based database of the glycosciences.de resource, which includes the CarbBank structures and literature references, as well as NMR data. The commercial GlycoSuiteDB is a relational database, which mainly includes O-linked and N-linked oligosaccharides.
Although carbohydrate structures and functions are being determined individually, there is still the issue of connecting the structures to the functions. The reason for this difficulty in determining the relationship between carbohydrate structures and function is due to the diversity or microheterogeneity of different carbohydrate structures attached to the same position of the same protein (Hui et al., 2002). In fact, carbohydrate structures are assumed to potentially form extremely complicated structures due to the diversity of glycosidic linkages. In other words, although DNA and proteins have only one kind of linkage for connecting two elements, carbohydrate structures have eight kinds, comprising four positions of linkages, the 2nd, 3rd, 4th, and 6th hydroxyl groups, and two types of anomeric configurations, alpha and beta. When real carbohydrate structure data is investigated, the combinations of monosaccharides with glycosidic linkages were actually found to be limited to some extent. Thus, one role of our resource is to represent all possible variations of carbohydrate structures within organisms.
In this review, we present a comprehensive glycome informatics resource, which includes the newly developed GLYCAN database, pathways of glycan biosynthesis and metabolism, and a newly constructed structural variation map CSM. GLYCAN is an important component of KEGG for linking glycans in the chemical universe to the gene universe and protein networks. It also includes two useful tools: (1) a new intuitive drawing tool for branched carbohydrate structures called KegDraw and (2) an efficient tool to search substructures and similar structures, called KEGG Carbohydrate Matcher (KCaM) (Aoki et al., 2004a,b). The glycan biosynthesis and metabolism pathways, with the integration of carbohydrate structures, proteins, and reactions, are also a pivotal resource. CSM is a bridge between carbohydrate structures and relevant genes. It is able to display, for example, expression data of glycosyltransferases in a compact manner, illustrating its versatility as a new bioinformatics instrument capable of analyzing carbohydrate structures on a global scale. Finally, we note that carbohydrate structures are represented as graph objects, or sets of nodes (monosaccharides) and edges (glycosidic linkages), which is a representation also used for chemical compounds and pathways in KEGG, and which enables the application of known computational algorithms for analysis such as structure comparison and motif detection.
KEGG GLYCAN
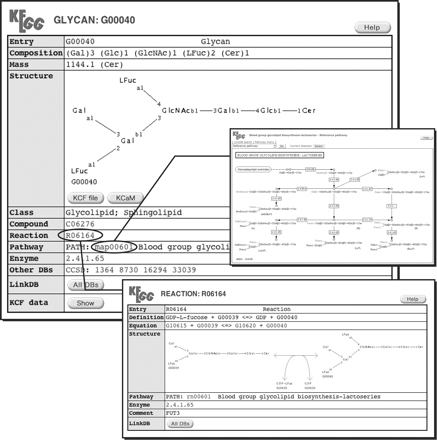
GLYCAN, which contains 11 066 unique entries (as of 1 July, 2005), is a publicly accessible glycan database at http://www.genome.jp/kegg/glycan/. Most of the GLYCAN entries are derived from ∼45,000 structures in CarbBank, including redundant ones. Over 150 structures were derived from KEGG PATHWAY. About 750 new carbohydrate structures determined after the termination of the CarbBank project have also been assiduously collected with the help of professional glycobiologists, mainly consisting of researchers in academia. In the GLYCAN database, a unique structure corresponds to an entry, which is identified by an accession number starting with the letter “G.” Each entry is annotated with various information for the structure, including composition, class, and mass, as well as a separate GIF image file for display. Additionally, entries may contain links to other databases in KEGG; such as COMPOUND, REACTION, PATHWAY, ENZYME, and KO (KEGG Orthology), summarized in Figure 1 and Table I.
Overview of a GLYCAN entry. The entry includes the structure, some fundamental information, and links to its entries in other databases, such as REACTION, and PATHWAY. See Table I for details regarding the content of these entries.
Annotation of entries in GLYCAN
| Attribute . | Description . |
|---|---|
| Name | Common name(s) for this entry, if any |
| Composition | Textual description of the monosaccharide composition |
| Mass | Molecular mass, calculated by summing the mass of the monosaccharides minus the number of bonds times the mass of water |
| Class | Classification of glycans, as in (glycoprotein; N-glycan) |
| Remark | Any comments, such as on lectins and other interacting molecules |
| Compound | Corresponding chemical compound entry in COMPOUND database. Some relatively smaller carbohydrates are also registered as compound entries |
| Reaction | Corresponding REACTION database entry if this entry is a reactant |
| Pathway | Corresponding PATHWAY database entry for the biological process in which this entry is involved |
| Enzyme | Corresponding ENZYME database entry for catalytic actions on this entry |
| Ortholog | Corresponding KO (KEGG Orthology) database entry to an ortholog group, which is an identifier for the orthologous genes from different organisms |
| Reference | Literature citation for this entry with links to PubMed |
| DBLINKS (other databases) | Links to corresponding entries in other external databases, such as CarbBank |
| Attribute . | Description . |
|---|---|
| Name | Common name(s) for this entry, if any |
| Composition | Textual description of the monosaccharide composition |
| Mass | Molecular mass, calculated by summing the mass of the monosaccharides minus the number of bonds times the mass of water |
| Class | Classification of glycans, as in (glycoprotein; N-glycan) |
| Remark | Any comments, such as on lectins and other interacting molecules |
| Compound | Corresponding chemical compound entry in COMPOUND database. Some relatively smaller carbohydrates are also registered as compound entries |
| Reaction | Corresponding REACTION database entry if this entry is a reactant |
| Pathway | Corresponding PATHWAY database entry for the biological process in which this entry is involved |
| Enzyme | Corresponding ENZYME database entry for catalytic actions on this entry |
| Ortholog | Corresponding KO (KEGG Orthology) database entry to an ortholog group, which is an identifier for the orthologous genes from different organisms |
| Reference | Literature citation for this entry with links to PubMed |
| DBLINKS (other databases) | Links to corresponding entries in other external databases, such as CarbBank |
Annotation of entries in GLYCAN
| Attribute . | Description . |
|---|---|
| Name | Common name(s) for this entry, if any |
| Composition | Textual description of the monosaccharide composition |
| Mass | Molecular mass, calculated by summing the mass of the monosaccharides minus the number of bonds times the mass of water |
| Class | Classification of glycans, as in (glycoprotein; N-glycan) |
| Remark | Any comments, such as on lectins and other interacting molecules |
| Compound | Corresponding chemical compound entry in COMPOUND database. Some relatively smaller carbohydrates are also registered as compound entries |
| Reaction | Corresponding REACTION database entry if this entry is a reactant |
| Pathway | Corresponding PATHWAY database entry for the biological process in which this entry is involved |
| Enzyme | Corresponding ENZYME database entry for catalytic actions on this entry |
| Ortholog | Corresponding KO (KEGG Orthology) database entry to an ortholog group, which is an identifier for the orthologous genes from different organisms |
| Reference | Literature citation for this entry with links to PubMed |
| DBLINKS (other databases) | Links to corresponding entries in other external databases, such as CarbBank |
| Attribute . | Description . |
|---|---|
| Name | Common name(s) for this entry, if any |
| Composition | Textual description of the monosaccharide composition |
| Mass | Molecular mass, calculated by summing the mass of the monosaccharides minus the number of bonds times the mass of water |
| Class | Classification of glycans, as in (glycoprotein; N-glycan) |
| Remark | Any comments, such as on lectins and other interacting molecules |
| Compound | Corresponding chemical compound entry in COMPOUND database. Some relatively smaller carbohydrates are also registered as compound entries |
| Reaction | Corresponding REACTION database entry if this entry is a reactant |
| Pathway | Corresponding PATHWAY database entry for the biological process in which this entry is involved |
| Enzyme | Corresponding ENZYME database entry for catalytic actions on this entry |
| Ortholog | Corresponding KO (KEGG Orthology) database entry to an ortholog group, which is an identifier for the orthologous genes from different organisms |
| Reference | Literature citation for this entry with links to PubMed |
| DBLINKS (other databases) | Links to corresponding entries in other external databases, such as CarbBank |
The size of the GLYCAN database continues to increase daily using information from the literature in which carbohydrate structures are newly determined. Furthermore, an Application Programming Interface (API) is available to allow programmers to access all data in KEGG including glycan information from software programs over the Internet.
KEGG pathways for glycan biosynthesis and metabolism
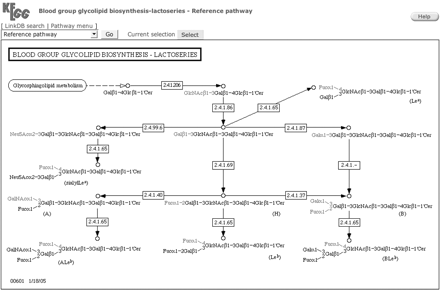
KEGG PATHWAY is a database that represents molecular interaction networks, including metabolic pathways, regulatory pathways, and molecular complexes. A unique feature of KEGG PATHWAY is its role in associating the gene universe with the chemical universe (and vice versa), consequently illustrating biological processes. Unlike other glycan databases, these known biosynthesis, metabolism, and degradation pathways integrate the structures, reactions, and enzymes related to glycans. For example, the pathway map of lactoseries glycolipid biosynthesis is composed of 13 glycosyltransferase reactions including nine glycosyltransferases and 13 glycans (Figure 2). Boxes represent enzymes that correspond to glycosyltransferases, which are hyperlinked to KEGG ENZYME, and circles represent glycans (or chemical compounds), which are hyperlinked to KEGG GLYCAN (or COMPOUND). In any pathway map, selecting an organism from the pull-down menu at the top colors the boxes of enzymes (or proteins) and changes the hyperlinks. In the case of Figure 2, after selecting a particular organism such as “Homo sapiens” and pushing the “Go” button, enzymes that have known genes in the human genome will be colored green and hyperlinked to their corresponding GENES entries. Furthermore, if “all organisms in KEGG” are selected under the same pull-down menu, the boxes of enzymes will be colored blue and hyperlinked to the KO entries representing ortholog groups across all organisms. Figure 3 illustrates the overall relationship of the 15 KEGG pathway maps currently available for glycan biosynthesis and metabolism as well as other metabolic pathway maps.
The pathway of the glycolipid lactoseries biosynthesis. Circles, rectangles, and arrows represent carbohydrate structures, enzymes, and reactions, respectively.
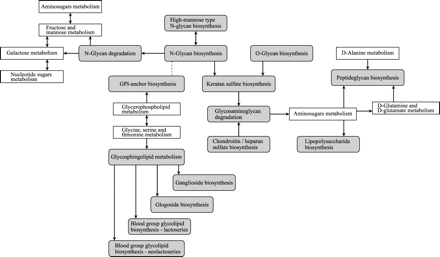
The overall relationship of the 15 KEGG pathway maps for glycan biosynthesis and metabolism (shaded) and some other metabolic pathway maps.
Composite structure map
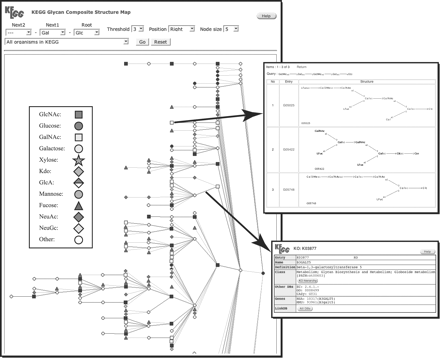
KEGG PATHWAY represents the step-by-step process of biochemical reactions involving genes and carbohydrate structures. On the other hand, the CSM is a static representation of all possible variations of carbohydrate structures in a tree format (Hashimoto et al., in press). Starting with the entire KEGG GLYCAN database as our data set, a CSM tree was constructed by taking all the carbohydrate structures containing a particular monosaccharide at its root and superimposing them into one unified structure by a tree structure alignment program (a modified version of Aoki et al., 2003). For example, CSM having “Glc” as the root is the tree containing all variations of carbohydrate structures whose roots are Glc (Figure 4). A different tree can also be displayed for any combination of up to three monosaccharides at the root. Each monosaccharide is represented by a symbol according to the standards set forth by the Consortium for Functional Glycomics (http://www.functionalglycomics.org/). Because every type of glycosidic linkage is distinguished, different glycosidic linkages are represented by different edges even if the attached monosaccharide is the same. For example, when a Gal can be attached to Glc by a b1–3 and a b1–4 glycosidic linkage, CSM will contain different edges “Gal b1–3 Glc” and “Gal b1–4 Glc.”
Any node on the CSM tree corresponds to a list of carbohydrate structures that are located on the single path from the root to that node. Thus, each node is hyperlinked to its corresponding list. Each structure in the resulting list is also hyperlinked to their GLYCAN entries. The top right of Figure 4 illustrates such a selected path. Because glycosyltransferases catalyze the biosynthesis of carbohydrate structures by the addition of individual linkages, each edge in the CSM is hyperlinked to its corresponding glycosyltransferase-related information, if known (see bottom right of Figure 4). When a specific organism is selected, each edge is colored and hyperlinked to its glycosyltransferase GENES entry. On the other hand, when “all organisms in KEGG” are selected, each edge is hyperlinked to the KO entry (the ortholog group) of the glycosyltransferase to which it corresponds (across all organisms).
An example of CSM (Composite Structure Map), which contains Glc as root, and Gal at the 2nd level. Each monosaccharide is represented by symbols according to the standards set forth by the Consortium for Functional Glycomics. A monosaccharide and a glycosidic linkage are represented as a node and an edge in this figure, respectively. The top right figure is the structure list corresponding to a node, and the bottom right figure is the KO (KEGG Orthology) list corresponding to the edge.
It is also possible to color the edges corresponding to a specification of colors and genes stored in a local file. Thus, a variety of analyses can be performed using this tool. For example, microarray gene expression data can be displayed in this tree by color coding the up- or down-regulation of genes involved in glycan synthesis, thereby, linking gene expression to glycan structures. Furthermore, CSM may also allow the prediction of structures based on other types of data, such as mass spectroscopy data.
KegDraw: glycan structure drawing tool
KegDraw is a freely available software tool for drawing chemical structures (Figure 5). Although there are several applications already available for drawing chemical compounds, few are available for drawing glycans. KegDraw is a Java application, so it runs locally in a platform-independent manner, and it allows the drawing of not only simple chemical compounds but also of glycan structures. KegDraw consists of two drawing modes: compound mode for drawing chemical compounds in a similar way as ChemDraw, and glycan mode for drawing glycans with monosaccharide units. In glycan mode, glycan structures can be drawn in a variety of ways. The simplest method is selecting monosaccharides and linkage conformations from popup menus one by one. However, convenient functionalities such as cut-and-paste and predefined template structures are also available. KegDraw currently handles files for input and output in the KCF (KEGG Chemical Function) format, which was originally developed for representing chemical compounds (Hattori et al., 2003) and extended to carbohydrate structures. KCF defines graph objects of carbohydrate structures as sets of nodes and edges, consistent with other KEGG graph objects including KEGG pathways. This allows the application of known efficient algorithms to the computational analysis of carbohydrate structures, as in KCaM described below. Glycan structures drawn in KegDraw can be used as queries to search against KEGG GLYCAN and other databases by KCaM. In the near future, other data formats for carbohydrates such as LInear Notation for Unique description of Carbohydrate Sequences (Bohne-Lang et al., 2001) will also be supported.
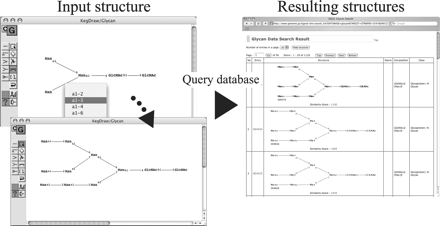
An example of inputting a carbohydrate structure using KegDraw. The inputted structure can then be sent to the database server as a query, and then similar structures are searched. Results are listed with a similarity score by KEGG Carbohydrate Matcher (KCaM). Three resulting structures in the figure are very close but have some different glycosidic linkages.
KCaM: structure search tool
The structures in the GLYCAN database can be accessed by queries using keywords such as the accession number or commonly used names (i.e., keyword search) or by using a structure search. The structure search tool employed by KEGG GLYCAN is called KCaM (Aoki et al., 2004b), which utilizes a dynamic programming technique and a theoretically proven efficient algorithm for finding the maximum common subtree between two trees (Aoki et al., 2003). This tool is available both at the KEGG website and through KegDraw.
KCaM consists of two main variations, an approximate matching algorithm and an exact matching algorithm. The former aligns monosaccharides allowing gaps in the alignment, whereas the latter aligns linkages and disallows any gaps, resulting in a stricter criterion for alignment. Both variations provide local and global options. The local approximate matching algorithm does not penalize the gaps for unaligned regions, whereas the global version does. Thus only conserved regions can be found using the local approximate matching algorithm. The local exact matching algorithm simply finds the first largest matching subtree to the query, although the global attempts to find as many matching subtrees as possible. As a guideline, local exact matching should be sufficient for queries using specific structures. In contrast, local approximate matching can be used for more general queries.
A genomic perspective of carbohydrate structures
There seems to be a wide variety of carbohydrate structures considering the thousands of unique structures already in the database. This complexity is caused by the combination of different monosaccharides with different glycosidic linkages, which are also related to the complexity of biosynthetic pathways and responsible genes. However, we find that most of the structures include core structures and common structures such as the root of N-glycan, O-glycan, and glycolipid. That is, the structures are not constructed randomly, and their variety is rather limited. This reflects the architecture of biosynthetic pathways, consisting of conserved portions and terminal variations as shown, for example, in the KEGG pathway maps of “N-glycan biosynthesis” and “High-mannose type N-glycan biosynthesis” (Figure 3). This, in turn, reflects the inventory of genes in the genome, which contains orthologous genes and paralogous genes, respectively responsible for the conservation and variation of such pathways.
The CSM is a tool for integrated analysis of both structural information and genomic information. Merging the common root structures, all currently available carbohydrate structures are represented compactly in CSM. Because a carbohydrate structure is synthesized by glycosyltransferases that add monosaccharides one by one, if the entire set of glycosyltransferases can be elucidated, the CSM can display all possible carbohydrate structures. In addition to structural variations, CSM also illustrates the relationship between a glycosidic linkage and the glycosyltransferase that catalyzes it. Currently, 70 known ortholog groups in KEGG, which include 176 organisms, are assigned to the edges in CSM (see supplemental material and also the latest list at http://www.genome.jp/kegg/glycan/GT.html). However, this corresponds to only 24% of the edges that are present in CSM, suggesting that there is still a large number of unknown genes or genes whose functions have not been fully characterized yet.
Conclusion
Toward the aim of constructing a comprehensive glycome informatics resource, we have built the carbohydrate structure database GLYCAN and represented the variations of carbohydrate structures as in CSM. GLYCAN contributes to functional glycomics in the sense that the glycan information in the database and the genome information of the other KEGG resources join together and form pathways, which relate to functions in organisms. This is in contrast to other carbohydrate databases, which are generally independent of other such data resources. Some bioinformatics research using GLYCAN has recently gotten underway, such as the analysis of bond patterns in the database for the prediction of glycan structures from mRNA expression data (Kawano et al., submitted for publication), the generation of a score matrix for carbohydrate structure alignments (Aoki et al., 2003, 2005), the biological classification of glycan structures using a machine learning method (Hizukuri et al., 2004), and the extraction of glycan structure motifs in leukemia cells (Hizukuri et al., in press). Such research is dependent on the fact that large quantities of carbohydrate structure data are computationally available. GLYCAN and our glycomics tools will surely contribute to glycome informatics and especially to carbohydrate structural analysis, as well as assist biologists to obtain the related information they want in experiments.
Supplementary Data
Supplementary data are available at Glycobiology online (http://glycob.oxfordjournals.org.).
Acknowledgments
We thank Dr. Yasushi Okuno and the Kansai Glycoinformatics Research Group for providing their specialized knowledge of glycans. We also thank Professors Peter Albersheim and Akira Kobata for allowing and arranging the use of CarbBank, Satoshi Miyazaki for developing KegDraw and Masayuki Kawasima for helping in the construction of CSM. The computational resource was provided by the Bioinformatics Center, Institute for Chemical Research, Kyoto University. This work was supported by the grants from the Ministry of Education, Culture, Sports, Science and Technology of Japan, Japan Society for the Promotion of Science, and Japan Science and Technology Corporation.
References
Author notes
2Bioinformatics Center, Institute for Chemical Research, Kyoto University, Uji, Kyoto 611-0011, Japan; 3Department of Biological Chemistry, Graduate School of Pharmaceutical Sciences, Kyoto University, Sakyo-ku, Kyoto 606-8501, Japan; 4Research Center for Glycobiotechnology, Ritsumeikan University, Kusatsu, Shiga 525-8577, Japan; and 5Human Genome Center, Institute of Medical Science, University of Tokyo, Minato-ku, Tokyo 108-8639, Japan