-
PDF
- Split View
-
Views
-
Cite
Cite
Karine Alix, Pierre R. Gérard, Trude Schwarzacher, J. S. (Pat) Heslop-Harrison, Polyploidy and interspecific hybridization: partners for adaptation, speciation and evolution in plants, Annals of Botany, Volume 120, Issue 2, August 2017, Pages 183–194, https://doi.org/10.1093/aob/mcx079
Close - Share Icon Share
Abstract
Background Polyploidy or whole-genome duplication is now recognized as being present in almost all lineages of higher plants, with multiple rounds of polyploidy occurring in most extant species. The ancient evolutionary events have been identified through genome sequence analysis, while recent hybridization events are found in about half of the world’s crops and wild species. Building from this new paradigm for understanding plant evolution, the papers in this Special Issue address questions about polyploidy in ecology, adaptation, reproduction and speciation of wild and cultivated plants from diverse ecosystems. Other papers, including this review, consider genomic aspects of polyploidy.
Approaches Discovery of the evolutionary consequences of new, evolutionarily recent and ancient polyploidy requires a range of approaches. Large-scale studies of both single species and whole ecosystems, with hundreds to tens of thousands of individuals, sometimes involving ‘garden’ or transplant experiments, are important for studying adaptation. Molecular studies of genomes are needed to measure diversity in genotypes, showing ancestors, the nature and number of polyploidy and backcross events that have occurred, and allowing analysis of gene expression and transposable element activation. Speciation events and the impact of reticulate evolution require comprehensive phylogenetic analyses and can be assisted by resynthesis of hybrids. In this Special Issue, we include studies ranging in scope from experimental and genomic, through ecological to more theoretical.
Conclusions The success of polyploidy, displacing the diploid ancestors of almost all plants, is well illustrated by the huge angiosperm diversity that is assumed to originate from recurrent polyploidization events. Strikingly, polyploidization often occurred prior to or simultaneously with major evolutionary transitions and adaptive radiation of species, supporting the concept that polyploidy plays a predominant role in bursts of adaptive speciation. Polyploidy results in immediate genetic redundancy and represents, with the emergence of new gene functions, an important source of novelty. Along with recombination, gene mutation, transposon activity and chromosomal rearrangement, polyploidy and whole-genome duplication act as drivers of evolution and divergence in plant behaviour and gene function, enabling diversification, speciation and hence plant evolution.
INTRODUCTION
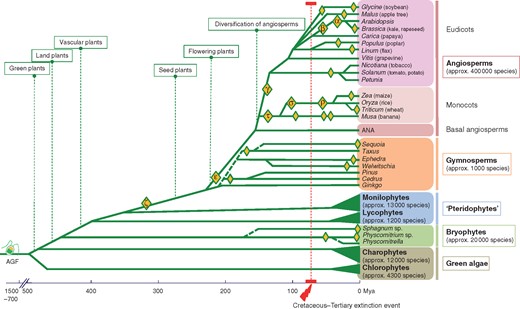
About half of all higher plant species are recognizable as evolutionarily recent polyploids, where multiple whole genomes or sets of chromosomes have come together from close ancestors (Soltis et al., 2015). Additionally, over evolutionary time, all flowering plants have at least one polyploidy event, also known as a whole-genome duplication (WGD), in their ancestry, from before the divergence of gymnosperms and angiosperms, the ζ (zeta) event (see Fig. 1, and references cited in the legend). Angiosperms, including Amborella and the basal angiosperms (i.e. ANA, for Amborellales, Nymphaeales, Austrobaileyales) that are sisters to all the other angiosperms, have a second polyploidy event in their lineage (ε, epsilon; Amborella Genome Project, 2013). Analyses of whole-genome sequences in the last decade have identified additional, and often multiple, polyploidy events in the ancestry of every eudicot and monocot where the genomes have been sequenced (summarized in Fig. 1). Notably, the near-universal occurrence of multiple polyploidy events (Wendel, 2015) during plant evolution is in contrast to most groups of animals in both recent and long-term evolutionary history (e.g. Hoffmann et al., 2012), marking a significant divergence between evolutionary mechanisms in the two kingdoms.
Simplified phylogeny of the green plant lineage focusing on the occurrence of WGD (whole-genome duplication) events. Polyploidy events (yellow diamonds) refer to either single or multiple rounds of WGD (i.e. duplication or triplication) and are labelled where applicable (Greek letters; see references below). Complete genome sequences have clearly established that WGD has remarkably shaped the evolutionary history of angiosperms compared with the other major clades of green plants. Estimates for the age of angiosperms have suggested the range of 167–199 million years ago (Mya) (Bell et al., 2010). Then rapid radiations responsible for the extant angiosperm diversity occurred after the early diversification of Mesangiospermae 139–156 Mya (Moore et al., 2007; Bell et al., 2010) with a burst of diversification specific for the Cretaceous, <125 Mya (age of the earliest angiosperm macrofossil; Cascales-Miñana et al., 2016). Early divergence times are from Bell et al. (2010) and Leliaert et al. (2012); for angiosperms from Fawcett et al. (2009), Jiao et al. (2011) and Li et al. (2016); and for gymnosperms from Lu et al. (2014). Dashed lines indicate imprecise timing or approximate representation of lineage divergence. WGD events are from Jiao et al. (2011); Leliart et al. (2011); D’Hont et al. (2012); Beike et al. (2014); Renny-Byfield and Wendel (2014); Li et al. (2015, 2016); Scott et al. (2016); Shaw et al. (2016); and Bombarely et al. (2016). See corresponding publications for precise estimates of time divergence and occurrence of WGD. AGF, hypothetical ancestral green flagellate; ANA, basal angiosperms including Amborellales, Nymphaeales, Austrobaileyales; following a standardized method, Greek letters are used to name polyploidy events along the phylogenetic tree, starting from the α (alpha) and β (beta) events that have been identified in the arabidopsis genome (Bowers et al., 2003).
With respect to its omnipresence during the evolutionary history of higher plants, polyploidy has been the subject of numerous reviews with emphasis on the genetic and genomic consequences of WGDs (Soltis et al., 2016). The present review has two main objectives: first, overviewing the papers in this Special Issue ‘Polyploidy in Ecology and Evolution’ and secondly to discuss complementary polyploidy-related topics covered. In the Special Issue, we made the choice not simply to consider the occurrence of polyploidy (or WGD) in plants (including the bryophyte Sphagnum), but also to provide an overview of the consequences of polyploidy in adaptation, speciation and evolution in plants: the relationships between polyploidy and stressful environmental conditions have suggested a major role for polyploidy in adaptation. This has been extensively analysed for cultivated plants, and we review this topic, in the context of concepts related to papers in the Special Issue, many of which present research in the novel area of polyploidy in natural plant populations. With the number of polyploidy events being revealed in plant evolution, the study of its evolutionary significance on wild plant species at the population scale, considering both evolutionarily ancient (deep) phylogenies and recent polyploids (some below the level of species), is now underway. The Special Issue articles consider the success and diversity found in polyploids from the ecological and evolutionary points of view, including developmental and genetic studies. Several papers deal with the relationships of polyploidy to plant reticulate evolution (i.e. natural hybridization), while others bear on the origin and formation of neopolyploids. Some papers discuss the relationships between allopolyploidy and reproductive systems, two major processes driving angiosperm diversification, and other papers highlight the link between polyploidy and adaptation, in a biogeographical context. Finally, a review dedicated to the impact of transposable elements (TEs) on polyploid plant genomes gives consideration to the molecular basis of genomic conflicts, particularly present in genome duplications with hybrid origins. We have specifically developed this renewed interest for the study of polyploidy in plants with new avenues of investigations dedicated to the epigenetic consequences of polyploidy and their role for plasticity and adaptation in plants.
OCCURRENCE AND DETECTION OF POLYPLOIDY IN PLANTS
Plant evolution and polyploidy
Since the historical Greek and Roman cultures, more than two millennia of research on plant diversity and relationships have established a robust phylogeny, placing all plants in a few monophyletic groups (as exemplified with angiosperms that now constitute a single monophyletic group), most recently based on evidence from DNA sequences (Angiosperm Phylogeny Group, 2016). Novel interpretations and new discoveries in the fossil record have been important for identifying innovations and divergences within plants (e.g. Matsunaga and Tomescu, 2016; Gandolfo and Hemsen, 2017), giving extra strength to modern phylogenies. Polyploidy is a confounding factor in phylogenetic analysis, in particular when it involves interspecific hybridization at the origins of the genome duplication. The resulting duplicated gene copies may lead to incongruence among multiple gene phylogenies, and the reticulate evolutionary patterns that mean lineages – typically at the level of genus or tribe and below – will be defined as multiple monophyletic groups (e.g. Fortune et al., 2007). Understanding the evolutionary and biological processes in their past diversification and considering future evolutionary pathways is a major research objective, notably addressed in the articles presented here.
Polyploidy and genome rearrangements, complementing mutation and recombination, are major evolutionary events, having the potential to lead to genetic and reproductive isolation. Most of the nodes within the phylogeny of plants (Fig. 1) represent disruptive changes in evolution, with vascularization, seeds and flowers representing successive evolutionary innovations. Previous Special Issues of Annals of Botany have stressed the major role of pollination biology and coevolution with pollinators in the ecological diversification of flowering plants (e.g. Friis et al., 2006; Van der Niet et al., 2014). Another important feature of angiosperms is their broad metabolic performance and their remarkable photosynthetic efficiencies based on carbon- and water-use capacities, thanks to morphological innovations in leaves, wood and roots, which have enabled colonization of a large diversity of habitats (Field and Arens, 2007). Angiosperm diversity can also be linked to the impact of the rounds of polyploidy (WGD) duplicating all the genes (Tank et al., 2015). As well as potentially giving reproductive isolation, polyploidy leads to multiplication of the number of genes and incorporating redundancy in function, enabling new genetic variability that may be acted upon by natural selection for evolution and adaptation. Indeed, allopolyploidy (associated with interspecific hybridization, see below) has long been considered as a source of the ‘most important amendment to Darwin and Wallace’s account of evolution’ (Haldane, 1959).
Detection of polyploidy
Extant polyploid plants can be detected by counting chromosome numbers in cytological metaphase preparations made from a group of related accessions (or species, as shown in Fig. 2). Where chromosome numbers in related plants are multiples (or sums) of others, the plant is likely to be a recent polyploid, and many species have a ploidy series with a base chromosome number x, including diploids (2x) and polyploids such as triploids (3x) and tetraploids (4x), and polyploids with even higher ploidies (see Heslop-Harrison, 2017). Equally, DNA content of the accessions can be measured by flow cytometry or Feulgen densitometry, and will show an additive series from the ancestral diploids to multiples in the polyploids (Kron et al., 2007). Nevertheless, DNA content and chromosome number are not always directly correlated to ploidy levels. For instance, karyotype evolution by chromosome fission and fusion has significantly marked the evolutionary history of diverse plant lineages, thus making it difficult to infer ploidy levels from cytogenetic data (e.g. in Crocus; Harpke et al., 2015). Where species have diverged before coming together in a polyploid, the use of total genomic DNA as a probe for in situ hybridization has been valuable both to identify the occurrence of a polyploid and to assist with identification of the ancestral species (e.g. Patel et al., 2011; Hunt et al., 2014). Within many species and species complexes, polyploid series can be present and are described as cytotypes (in this Special Issue, e.g. Čertner et al., 2017; Paule et al., 2017; or see Taketa et al., 1999) or, when polyploidy events are older, as different species (Hordeum; Cuadrado et al., 2017).
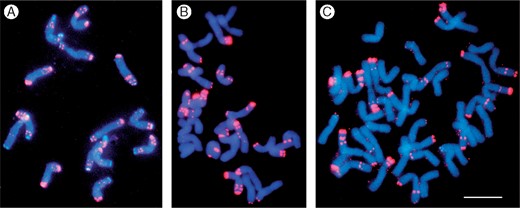
Metaphase chromosomes of diploid, tetraploid and hexaploid wheats stained with the DNA stain 4',6-diamidino-2-phenylindole (DAPI; cyan) and showing fluorescent in situ hybridization signal (magenta) from the 120 bp tandemly repeated (pSc119.2) DNA family common to many Triticeae species (see Contento et al., 2005). This repeat family originated before the split of rye, barley, wheat and other grasses in the tribe, but has been amplified differentially in the different species. It forms large blocks at sub-telomeric and intercalary chromosomal regions in the B genome wheats, both seen in the seven chromosome pairs in the diploid (A), tetraploid (B) and hexaploid (C), but has only few sites in about half of the A and D genome chromosomes with weak single sub-telomeric foci (B, C). (A) Aegilops speltoides (2n = 2x = 14, genome constitution B'B'); (B) Triticum durum (2n = 4x = 28, AABB); (C) T. aestivum (2n = 6x = 28, AABBDD). Scale bar = 10 μm.
Two main types of recent polyploidy can be distinguished. Autopolyploidy defines duplication of one genome within one species, which results in homologous chromosome sets in the cell (diploid AA doubles to become the autotetraploid AAAA). Allopolyploidy defines WGD associated with the merger of two or more divergent genomes in a single nucleus following interspecific hybridization, resulting in homoeologous chromosome sets in the cell (AA × BB → AABB). In practice, auto- and allopolyploidy are not entirely separate, since this terminology depends on the taxonomic definition of a species and the scope of a designated ‘genome’. Higher polyploids may be combinations as a result of their evolutionary histories that have been marked by recurrent WGDs including both auto- and allopolyploidy events, thus giving the ‘auto-allo-polyploid’ designation (usually summarized as polyploid). For ancient WGD events, where the ancestral diploid species (singular or plural) are unknown and DNA sequences are diverged, auto- or allopolyploidy, as depicted in Fig. 1, cannot be distinguished.
For the detection of ancient polyploid events, whole-genome sequencing coupled with comparative genomics has proved to be remarkably efficient; this is particularly well illustrated by the Amborella Genome Project (2013). Typically, following assembly of the DNA sequence, all-frame protein translations are made to produce a full set of gene predictions. Structural comparisons of the set of gene predictions then allow the identification of paralogous gene pairs used to reconstruct ancient WGDs. Indeed, even over very long evolutionary periods, regions of synteny – the same list and order of genes – and of similar, paralogous genes (thus corresponding to syntenic blocks) are readily identified on different chromosomes, enabling the detection of multiple ancient WGD events (see Fig. 1, and the references cited in the legend).
HOW SUCCESSFUL ARE POLYPLOIDS?
Evolutionary success of polyploidy
Before the recognition of the ubiquitous ancient polyploidy or WGD events in plants, polyploids were widely considered as being present in many lineages. The long-term compared with short-term evolutionary success of polyploids, whether in crops or wild species, was a significant question (Arrigo and Barker, 2012; Soltis et al., 2014). Mayrose et al. (2011) consider that ‘polyploidy is most often an evolutionary dead end’ with respect to neopolyploids and their associated low diversification, and thus speciation rates, in comparison with diploids. Nevertheless, polyploids do represent longer term evolutionary successes; the WGD events at their origins may have generated the necessary genetic diversity that has been under positive selection for long periods, and then associated with phenotypic novelty, adaptability and higher fitness. Of course, the role of subsequent genome evolution (mutation, chromosome rearrangement and TE mobilization) in contributing to this genetic diversity has also to be considered. Interestingly, the polyploidy events that have marked the evolutionary history of the green plant lineage occurred prior to or simultaneously with major evolutionary transitions and adaptive radiation of species, suggesting a role for polyploidy in adaptive speciation (De Bodt et al., 2005). This is particularly well illustrated with the survival and proliferation of polyploid plant lineages during the Cretaceous–Tertiary mass extinction event (Fawcett et al., 2009; Fig. 1). The occurrence of polyploidy in response to stressful conditions has been notably analysed in the context of plant domestication that represents conditions of strong selection pressure for adaptation to human cultivation.
Polyploidy and plant domestication
Many crop species have been identified as polyploids through the presence of polyploid series in chromosome number (e.g. Salman-Minkov et al., 2016) and through generation of hybrids (e.g. Renny-Byfield and Wendel, 2014), and many crops are relatively recent polyploids (Leitch and Leitch, 2008). This has led to the hypothesis of some relationship between intensive selection, typically accompanying plant domestication, and polyploidy. Salman-Minkov et al. (2016) considered that polyploidy followed by domestication was a key feature in early selection of crops, and showed that domesticated plants have gone through more polyploidy events than their wild relatives. Nevertheless, it can be noted that the advantages of polyploidy under domestication are not overwhelming: the top three crops in the world, all cereal grains, are wheat (a modern hexaploid 2n = 6x = 42; Fig. 2C), rice (diploid, 2n = 2x = 14) and maize (a paleotetraploid, 2n = 4x = 20). With the strict polyploid selection criteria of Salman-Minkov et al. (2016), 30 % of crops are polyploid compared with 24 % of wild species.
Induced or post-domestication polyploids represent a small proportion of crops (reviewed by Sattler et al., 2016). As well as the genetic buffering and increased number of gene alleles associated with the increase in cell size, a major consequence of polyploidy may be the global increase of organ size (including leaves, stems, flowers, stigmas and tubercles) of the polyploid compared with its diploid counterpart; it is thus generally considered that polyploidy may increase fruit size as well as seed size, valuable traits for crop breeding. Such a consequence also illustrates one of the major advantages of allopolyploids, i.e. their ‘doubled interspecific hybrid’ composition, which directly results in fixing heterozygosity at the origin of heterosis, or hybrid vigour (Comai, 2005). Wu et al. (2012) stated that there has been little systematic evaluation of fruit from induced polyploids in comparison with diploids, and that there was only little evidence for larger fruits. However, their detailed work in kiwifruit (Actinidia chinensis) showed that colchicine-induced autotretraploids were some 50 % larger than their diploid progenitors, and that the increase was stable during vegetative propagation. In addition, specific work on wheat polyploids clearly demonstrated that seeds from synthetic hexaploid wheat were significantly larger than seeds from either progenitor, tetraploid or diploid (Kenan-Eichler et al., 2011). Thus, one can suggest that the consequences of polyploidy extend to developmental effects (see also Münzbergová, 2017).
Further investigations of the consequences of genome duplication in relation to adaptation abilities in natural populations from various plant lineages should provide a valuable insight into the important evolutionary process of polyploidy, as reported by the articles presented in this Special Issue.
POLYPLOIDY IN ECOLOGY AND EVOLUTION
Polyploidy, diversity and reticulate evolution
Understanding the components of the evolutionary success of polyploidy has generated important research activity in the last 20 years. Autopolyploidy has received relatively little attention by evolutionary biologists, not least since diploids and autopolyploids can be considered as one species (sometimes subject to redefinition such as, for example, in Brachypodium by Catalan et al., 2012); indeed, it has been estimated that the evolutionary advantages of allopolyploidy are largely superior (Parisod et al., 2010b). Evolutionary advantages of allopolyploidy can be associated with merger of differentiated genomes (i.e. via interspecific hybridization) with consequent genomic restructuring and rebalancing of gene expression, leading to phenotypic changes (Hegarty and Hiscock, 2008; Parisod et al., 2010b; Tayalé and Parisod, 2013; Mutti et al., 2017). How large is the role of reticulate evolution in the evolutionary success of flowering plants (reviewed by Soltis and Soltis, 2009), and should polyploidy be considered, following interspecific hybridization, as the major driver of plant evolution and speciation? Hybridization has been proposed as a direct driver of WGD, but the causal relationships between both processes are far from clear, and autopolyploidization has certainly contributed to plant diversification (Soltis et al., 2007; Buggs et al., 2009; Barker et al., 2016).
Diploid plants, including hybrids, produce unreduced gametes at relatively high rates, which participate in the formation of many polyploid lineages (Ramsey and Schemske, 2002; Otto, 2007). These lineages are often reproductively isolated from their progenitors, giving rise to new species or cultivars (Čertner et al., 2017, see below; and, for example, in Crocus, Orgaard et al., 1995; Alsayeid et al., 2015; in Mimulus, Sweigart et al., 2008; Vallejo-Marín et al., 2015). Interploid fitness loss might not be universal though, because polyploid plant species with odd chromosome numbers are widespread (Otto, 2007) and triploids can often have a role as bridges between diploid and tetraploid species (Ramsey and Schemske, 1998) often exploited in breeding (Heslop-Harrison and Schwarzacher, 2007). The role of hybridization in creating high genetic variability by producing new genotypes at the onset of potential speciation events is a question that has been recurrently addressed, notably by two papers in this Special Issue.
In the first paper from Yan et al. (2017), the role of hybridization in diversifying and structuring natural populations is particularly well illustrated within the genus Rhododendron, where interspecific hybridization is frequent. By analysing 24 Rhododendron populations including 15 independent hybrid swarms, the authors have addressed the question of the extent of hybridization in population structure and differentiation (Yan et al., 2017). Using microsatellite and chloroplast DNA sequencing, the authors demonstrate that hybridization events are complex and highly variable from one population to another, and result mainly in introgressions. The recurrent backcrosses lead to new allele combinations, generating genetic diversity for new variants among populations and potentially driving speciation. Secondly, Landrein et al. (2017) investigated the genetic diversity among five Abelia species (Caprifoliaceae) originating from China, by analysing wild taxa and horticultural varieties, to identify the origins of the genetic diversity, including that exploited in breeding programmes. Their study was based on phylogenetic reconstruction using nuclear and chloroplast markers, and molecular genotyping for genetic diversity and population structure analysis. They demonstrate that the Abelia genus has been subjected to recurrent hybridization and introgression, at the origins of allopatric speciation events. Their results demonstrate the value of exploiting hybridization, which, in association with random recombination, may produce interesting allele combinations for ornamental breeding purposes.
A third paper of the Special Issue also reports the study of the relationships between interspecific hybridization and genetic diversity, but includes the genome duplication process by analysing and comparing the allopolyploids. Cuadrado et al. (2017) demonstrate the role of allopolyploidy in diversifying Hordeum (a genus in which nearly half of the taxa are polyploids, and where some species such as Hordeum murinum are represented by morphologically indistinguishable diploid and autopolyploid cytotypes; Taketa et al., 1999) by analysing the hybrid genome composition of two closely related allotetraploid species using molecular cytogenetics. Even closely related genomes in the allopolyploid harbour high genomic diversity, enabling colonization of different geographical areas. The evolutionary divergence of H. secalinum and H. capense and their current intragenome diversity may thus be related to the differential genomic and chromosomal modifications encountered by the two parental genomes that merged during the allopolyploidization event.
Outside the flowering plants, polyploidy has been important in the evolution of mosses (Shaw et al., 2016; Fig. 1), where it is associated with hermaphroditism (Crawford et al., 2009) and more generally with changes in mating systems (Jesson et al., 2011). With respect to the high level of fixed heterozygosity observed for numerous moss species, a preponderant role for allopolyploidy in bryophyte diversification has been suggested. This allopolyploidy-based evolutionary process is particularly exemplified in Sphagnum (Ricca and Shaw, 2010), which is the model of the study reported in this Special Issue. Karlin and Smouse (2017) investigated the allelic diversity of the peat moss Sphagnum × falcatulum, a double allopolyploid species originating from the combination of three monoploid genomes from three ancestral species (i.e. it is an allo-allo-triploid). They used simple sequence repeat (SSR) genotyping on a large sample of S. × falcatulum gametophytes (in comparison with the immediate progenitors, i.e. the monoploid genomes) from different populations widespread in the Holantarctic. Most of the genetic diversity is captured directly from the three ancestral monoploid genomes, and multiple origins of S. × falcatulum are likely (i.e. recurrent allopolyploidization). Allopolyploidy may thus be the single most important factor in generating the genetic diversity allowing this peat moss to colonize the Holantarctic region so successfully.
Neopolyploid formation: a chromosomal point of view
Polyploids are usually formed from hybridization of taxonomically close species, where the gene content, sequences and often chromosomal organization, if not the repetitive DNA (e.g. Fig. 2), are similar. New hybrids in the first or early generations, including both spontaneous hybrids and those made in a research or plant breeding context, are classified as neopolyploids. In nature, some species complexes with high phenotypic diversity and many microspecies of various ploidies give complex taxonomic problems (e.g. Rhododendron discussed above; Taraxacum, Majeský et al., 2012; Rubus, Heslop-Harrison, 1968). The taxonomic challenges and diversity in populations mean that studies with large numbers of individuals and molecular markers are required.
The first advantage of allopolyploids (AABB) in comparison with homoploid hybrids (AB) is the immediate recovery of homologous chromosome pairs as a prerequisite to regular meiosis, and therefore potentially restoring fertility (at least partial) in the resulting hybrid genotypes, thanks to chromosome set doubling. Nevertheless, polyploids usually also require a tight genetic control of crossover formation and distribution to minimize aneuploidy and to ensure further fertility and genome stability, and thus the establishment and success of neopolyploids (Jenczewski and Alix, 2004; Grandont et al., 2013). Three papers in this Special Issue focus on neopolyploids, dealing with the extent of the formation of polyploid cytotypes in natural conditions, which mainly involve the production of unreduced gametes from diploids (as mentioned above) and the immediate consequences of polyploidization that can be identified using synthetic polyploids.
In their analysis of a collection of > 800 samples of dog roses (Rosa sect. Caninae), Herklotz and Ritz (2017) addressed the question of the extent of natural interspecific hybridization between the sub-sections Caninae and Rubigineae, finding that populations had a mixture of hybridogenic and non-hybridogenic individuals. Dog roses comprise many allopolyploid species (4x, 5x and 6x) notably thanks to a unique asymmetric meiosis process with only two sets of chromosomes forming bivalents that results in haploid pollen grain but a polyploid egg cell (Ritz and Wissemann, 2011). The authors demonstrate that polyploids are more frequent in Rubigineae hybrids in relation to their higher capacity in producing unreduced gametes than in Caninae, representing a major bias between reciprocal crosses. However, despite their viability and abundance, it seems that neopolyploid hybrids, in contrast to non-hybrids, do not spread between localities. In a large-scale spatio-temporal study in central Europe, Čertner et al. (2017) used flow cytometry data on > 11 000 individuals, as well as ex situ germination data, to study the cytotypic structure and dynamics of a contact zone between diploid and tetraploid populations of the annual herb Tripleurospermum inodorum (Asteraceae). In spite of an apparent substantial amount of gene flow between the two cytotypes, and the presence of fertile triploids that could serve as bridges to new polyploids (see above), alternative cytotypes are rare. Newly formed tetraploids were extremely rare and were thus not able to establish, suggesting that successful tetraploids probably result from one or a few rare and ancient polyploidization events – echoing the theory developed by Mayrose et al. (2011). Together, these studies show the importance of mixed-ploidy populations in the generation of cytotype variation, with population structure (in sympatry) that may help maintain a resource of genetic diversity conferring at least long-term adaptive advantages.
Neopolyploids, and in particular synthetic polyploids, are the most appropriate plant material to analyse and further understand the direct consequences of polyploidization during speciation. Münzbergová (2017) addresses this question on the model Vicia cracca that comprises both diploid and autotetraploid cytotypes. By analysing and comparing natural diploids, natural tetraploids and synthetic tetraploids from four different populations, the author demonstrates the direct impact of polyploidy on morphological traits (e.g. seed weight, plant height and stomatal size). This study also highlights the need to analyse multiple populations of polyploids to provide robust conclusions about the impact of polyploidization that may be variable with respect to the diversity of the populations at the origins of the new polyploids. Finally, the author performed a comparative study between the synthetic tetraploids and their diploid offspring to evaluate the impact of the colchicine treatment at the origin of the synthetic polyploids: only the analysis of the third and subsequent generations of synthetic polyploids may allow the consequences of polyploidization per se to be revealed. Such conclusions should be beneficial to the study of ‘old’ polyploid species (e.g. oilseed rape, cotton, wheat and tobacco) for which the true diploid progenitors are no longer available and the synthetic polyploids are the unique plant material useful to model and understand their formation.
Polyploidy and reproductive systems
Reproduction, whether sexual, apomictic or vegetative, is central to the success of polyploids. Notably, the near-universal occurrence of multiple polyploidy events in plant evolution is in complete contrast to the animal kingdom in both recent and long-term evolutionary history, where most analyses have found only one or two early WGD events, such as the R1 and R2 WGDs in the vertebrate stem lineage followed by another WGD in early fish (e.g. Hoffmann et al., 2012); these WGDs are generally viewed as having provided the genetic diversity for many innovative vertebrate- or fish-specific characteristics and fuelled their burst-like evolution. Recent polyploids are unusual, but have been found in individual species or lineages, e.g. fish, amphibians, crabs or insects (Mable et al., 2011; Kenny et al., 2016). Differences between plant and animal reproduction have been proposed for the differential polyploid formation rate (Otto and Whitton, 2000; Coyne and Orr, 2004; Husband et al., 2013). Few animals, with the exception of protozoa and a few insect species, have the possibility of apomictic or the equivalent of vegetative reproduction. Even with mechanisms to ensure correct pairing of chromosomes in a polyploid (e.g. Sepsi et al., 2017), the widespread occurrence of sex chromosomes in animals may also limit fertility in polyploid animals (Collares-Pereira et al., 2013). An evolutionary association between reproductive modes and polyploidy has been studied for decades (e.g. Stebbins, 1950) through two main aspects: the occurrence and rate of asexual vs. sexual reproduction on one hand, and the transitions in mating systems (i.e. self-compatibility or sexual dimorphism) on the other hand. When correlations have been detected, it has usually proven difficult to disentangle the causes and consequences of the different processes. As Ashman et al. (2013) showed, the observed association between sex dimorphism (e.g. dioecy) and polyploidy can be explained by direct or indirect effects of one process on the other, since evolutionary transitions usually occur simultaneously, and different factors can affect these transitions in different clades (Glick et al., 2016). The same questions hold for the association between self-fertilization and ploidy, since polyploidy usually correlates with self-compatibility (Barringer, 2007; Alix et al., 2008; Husband et al., 2008; Robertson et al., 2011) even if exceptions have been reported (e.g. in the Fragaria genus; Liston et al., 2014). Thus, the evolutionary link between mating system and ploidy has been fairly well documented, but the association of polyploidy with asexual reproduction (beyond the odd ploidies such as 3x or 5x), and particularly clonal reproduction (i.e. vegetative growth in plants), has been less studied, particularly in wild species. Three articles of this Special Issue deal with this question.
Herben et al. (2017) report association of vegetative reproduction with polyploidy in a large, phylogenetically broad, sample of 900 European angiosperm species to infer macroevolutionary patterns of both traits, and examine their temporal trends. As hypothesized, they detect a signal of correlated evolution between polyploidy and vegetative growth, mostly mediated by increased distance of spread. This is nicely confirmed by experimental ‘garden’ data, which show that diploids rely more on seed reproduction whereas polyploids rely more on vegetative spread. They also show that vegetative reproduction may often evolve before polyploidization and could then enhance the rates of polyploid speciation.
The evolutionary association between asexual production of seeds (i.e. apomixis) and polyploidy has been studied frequently, not least because gametophytic apomixis and polyploidization share the same characteristic feature, the production of unreduced gametes (Ramsey and Schemske, 1998; Whitton et al., 2008; Majeský et al., 2012; Husband et al., 2013). Here, two papers study the dynamics of apomixis in a context of hybridization and polyploidy. In the first paper, Uhrinová et al. (2017) assess the genetic relationships between parental and hybrid species in the genus Sorbus, which is known for exhibiting apomictic microspecies produced by hybridization (Robertson et al., 2010). Three parental species, one a tetraploid species (S. chamaemespilus) and two mainly diploid cytotypes (S. aria and S. aucuparia), seem to have produced a set of genetically distinct polyploid microspecies through interspecific hybridization. These polyploid species seem at least partially reproductively isolated and exhibit a low population genetic diversity compared with the parental species populations as well as a high rate of clonality. If this low diversity results from predominant apomixis, this would be another confirmation of the evolutionary link between apomixis and polyploid speciation through hybridization, at least in the genus Sorbus. Similarly, ancient polyploid apomicts of the subgenus Rubus studied by Šarhanová et al. (2017) seem to have been maintained in central Europe without relying on their facultative sexuality but rather on hybridization with a sexual species (mainly ser. Glandulosi). This process may regularly produce successful apomictic polyploid lineages but with low genetic polymorphism.
Polyploidy and ecological divergence
Over long evolutionary times, at least among angiosperms, new polyploid species have driven their diploid ancestors to extinction at several points (Fig. 1). However, open questions remain. Are most polyploid lineages evolutionary dead ends? Have particular events, such as changing climate, enabled polyploids to outcompete their diploid ancestors? Are any evolutionarily recent (perhaps the last 5 million years) polyploids more successful than the equivalent diploids? Several authors have considered related questions about plant genome evolution (Mayrose et al., 2011, 2015; Arrigo and Barker, 2012; Soltis et al., 2014; Vanneste et al., 2014), some reflecting the biogeographical patterns of polyploid species and their relationship to ecological adaptation (Madlung, 2013; e.g. Herklotz and Ritz, 2017). It has been postulated for a long time that polyploidization might be associated with particular and large species ranges and/or extreme habitats (Stebbins, 1950; Otto and Whitton, 2000; Levin, 2002; Husband et al., 2013; Weiss-Schneeweiss et al., 2013). Potentially contrasting results have been obtained from biogeographical surveys of polyploid and diploid species (e.g. Martin and Husband, 2009; Pandit et al., 2011). However, even environmental correlations (e.g. polyploidy associated with higher latitudes or particular climatic niches) do not necessarily imply ecological adaptation as an explanatory factor, since other processes such as demography (i.e. dispersal variance and genetic drift) can produce the same patterns.
Experimental designs such as reciprocal transplants can provide compelling evidence of adaptation of polyploids (e.g. Ramsey, 2011; Herben et al., 2017), although local biogeographical studies of sister or closely related species can provide useful insights into the patterns of ecological divergence of polyploid and diploid species (e.g. Laport et al., 2016). In this Special Issue, Paule et al. (2017) assess the degree of ecological divergence of polyploid species from the bromeliad Fosterella in Andean mountains, in relation to historical biogeographical processes. Polyploids seem to occupy slightly divergent climatic niches, but patterns of cytotype geographical distribution suggest a historical parapatric differentiation of polyploid species that has not necessarily been caused by ecological adaptation. Both processes are potentially at the origin of polyploid range shifts in this southern American species complex. This is also probably the case for the Mediterranean grass Anthoxanthum studied by Chumová et al. (2017). They show that several diversification events might have taken place since the Miocene from a diploid ancestor, by recurrent range expansions possibly linked to climatic niche differentiation.
THE GENOMIC CONFLICTS AWAKEN: POLYPLOIDY AS A PEACEKEEPER?
Apart from the presence of ancient or recent polyploidy, and the wide variation in chromosome number, the large variation in size of plant genomes is an enigma. Measured as the 1C or unreplicated haploid genome size, the range known is from 63 Mb for Genlisea margaretae to 149 000 Mb for Paris japonica (Bennett and Leitch, 2011). The ‘C-value paradox’ after work of Swift (1950) notes that there are few correlations between apparent organism characteristics and genome size (e.g. Freeling et al., 2015). For example, Krahulcová et al. (2017) show no significant correlation between DNA content and seed size in nine Aesculus species. However, a correlation between plant genome size and content of TEs has been observed (Pearce et al., 1996; Ågren and Wright, 2011; Negi et al., 2016), with long terminal repeat (LTR) retrotransposons being the most abundant source of genome size variation in many lineages (Bennetzen and Wang, 2014; Biscotti et al., 2015). In addition to polyploidy, TE activity (proliferation and/or elimination; Vitte and Panaud, 2005) is associated with genomic recombination and contributes to shape plant genomes by copy number increase (e.g. Santos et al., 2015), gene disruption (e.g. Tam et al., 2007) or co-mobilizing of other sequences such as those associated with reproductive incompatibility (Alix et al., 2008).
Here, Vicient and Casacuberta (2017) review the major contribution of TEs to plant genome diversity and evolution as well as to gene expression variation, focusing mainly on LTR retrotransposons. In relation to their capacity to move across the genome and their close association with unequal and illegitimate recombination, TEs (notably LTR retrotransposons) may generate a large variety of structural mutations that can be beneficial – even if they are more commonly detrimental or at least neutral. Such positive mutational TE insertions have represented interesting genomic targets for selection during plant domestication and crop breeding (for reviews, see Lisch, 2013; Vitte et al., 2014). Transposable elements are usually targeted efficiently by epigenetic marks (including DNA methylation and post-translational modifications of histones) to ensure their tight control in order to avoid any anarchical activation and transposition across the genome (Mirouze and Vitte, 2014). Stress conditions are assumed to reactivate TEs because of the extensive epigenetic remodelling of the genome they might trigger; interspecific hybridization and polyploidy can act as such stresses (an implication of McClintock, 1984; reviewed by Parisod et al., 2010a). Vicient and Casacuberta (2017) thus emphasize the close relationships between polyploidy and TE activity and the involvement of TEs in gene regulation in response to the epigenetic modifications associated with polyploidization. The authors highlight the impact of mediated DNA methylation changes at TE insertion sites on the functional regulation of neighbouring genes, as found, for instance, in arabidopsis in response to biotic stress (Dowen et al., 2012) and in rice following polyploidization (Zhang et al., 2015).
The major TE silencing mechanism in plants is the biogenesis of small non-coding RNAs, known as small (or short) interfering RNAs (siRNAs), with populations of 21 nucleotide long (21-nt) siRNAs and mainly 24-nt siRNAs that control TEs by TGS (i.e. transcriptional gene silencing) through RNA-directed DNA methylation, RdDM (Lewsey et al., 2016). Only a few studies have dealt with the immediate impact of polyploidy on the biogenesis of small RNA populations (Ha et al., 2009; Kenan-Eichler et al., 2011; Martinez Palacios, 2014; Zhang et al., 2015), but they all reported major changes in expression profiles of 24-nt siRNAs. The responses of small RNAs to (allo)polyploidy, which are also reminiscent of the more general dynamics of small RNAs depicted in response to hybridization, have progressively led to the idea of a major role for TE-derived siRNAs in the process of hybrid incompatibility (reviewed by Ng et al., 2012). At the onset of formation of the hybrid or the neoallopolyploid, there is a need to overcome hybrid failure, which ranges from early seed inviability to hybrid sterility. Similar to parental imprinting in animals, genome reprogramming occurs in plants, but specifically in vegetative cells, with the loss of DNA methylation at the origin of TE reactivation, resulting in the production of novel TE-derived siRNAs (Slotkin et al., 2009). If parental TE sequences differ substantially, often the case during interspecific hybridization, TE-derived siRNAs inherited from each parent represent key factors to control the reactivated TEs, and their efficiency in repressing TEs may determine the outcome of the interspecific cross (Martienssen, 2010).
Reproductive isolation, and particularly post-zygotic isolation, between plant and animal species is thought often to result from genomic conflicts involving selfish genetic elements (Presgraves, 2010; Rieseberg and Blackman, 2010; Ågren, 2013). The view of polyploidy, and especially allopolyploidy, as a ‘peacekeeper’ in genomic conflicts when two divergent genomes are brought together in hybrids is not new (Rieseberg, 2001; Tayalé and Parisod, 2013), but it can also be viewed as generating conflicts that will have to be resolved in a more complex way (Comai, 2005; Jones and Pašakinskienė, 2005). These genomic conflicts can have multiple origins such as TE-mediated silencing misregulation or segregation distorters, e.g. female meiotic centromere drive or supernumerary chromosome drive (Jones and Pašakinskienė, 2005), and there are different ways in which allopolyploidization can buffer their effects. First, when they are at the origin of epistatic autosomal Bateson–Dobzhansky–Muller (BDM) hybrid incompatibilities, they will be on average recessive and expressed mainly in F2 and subsequent generations. A direct consequence of polyploidization will be to avoid expression of this kind of hybrid defect because recessive BDM factors will almost never be homozygous, provided that the recombination rate between homoeologous chromosomes is low. A related effect of this lack of recombination will be the proper segregation of chromosomes in allopolyploids compared with homoploid hybrids (Tayalé and Parisod, 2013). Secondly, hybrid incompatibilities due to divergent gene expression (e.g. for parentally imprinted genes) or sub-functionalization of duplicate genes will potentially have no effect in polyploids. Thirdly, polyploidization could facilitate the occurrence of apomixis by involving low recombination, heterochromatic regions behaving as selfish genetic elements, such as supernumerary chromosomes (Roche et al., 2001; Comai, 2005), and providing in turn an advantage for propagating the new species in the first generations. Finally, diploid hybrid defects as well as dosage-related problems specific to polyploidization will be overcome, usually in later generations, by gene expression remodelling and/or chromosomal rearrangements and gene loss (Chen and Yu, 2013). The modulation of gene regulation, including genome dominance in polyploids (Woodhouse et al., 2014), is probably important for the success or evolutionary failure of a new hybrid.
The epigenetic control of genomic conflicts determining the viability of hybrids opens up new avenues of investigation, which include that of the evolutionary significance of epigenetic regulation in the success of interspecific hybridization and allopolyploidy, and the differential control of multiple genomes.
CONCLUSION AND PERSPECTIVES
Polyploidy is arguably the major feature of plant genome evolution, doubling the number of copies of each gene with each WGD event. Over evolutionary time, polyploids have displaced all their diploid ancestors several times in multiple independent lineages, suggesting a strong selective advantage. These multiple polyploid events have only been detected thanks to widespread large-scale whole-genome sequencing since 2000, and comparative genomic analyses continue to reveal the recurrence and extent of WGD during the evolutionary history of plant lineages (Fig. 1). The ‘-omics’ era has allowed understanding of the evolutionary history of lineages (e.g. Woodhouse et al., 2014; Douglas et al., 2015; Roux and Pannell, 2015; Wendel, 2015; Barker et al., 2016). Several of the most important angiosperm groups include a WGD event that is detected soon after the Cretaceous–Tertiary extinction event (Fig. 1); one can speculate that their genetic structure enabled these polyploids to thrive. One can ask if new polyploids may once again have advantages during the global events including widespread extinction (see Parmesan and Hanley, 2016) now being detected, including climate change. Plants have genetic mechanisms to overcome the challenges of polyploidy, in particular co-regulation of multiple, similar or identical copies of genes, and the adoption of vegetative or apomictic reproduction, or restitution of diploid behaviour during chromosome pairing and recombination at meiosis. Now we can identify the impact of polyploidy, and associated hybridity, on speciation and regulatory mechanisms at the gene expression level, and examine its impact on plant populations, as reported in this Special Issue.
ACKNOWLEDGEMENTS
We particularly thank Eric Jenczewski for helpful discussions, and Alessandra Contento for the images shown in Fig. 2. We also thank Zuzana Münzbergová and two anonymous referees for their helpful comments on our manuscript. We finally thank all our colleagues and collaborators, as well as our students and post-docs for their contribution to our work on polyploidy and interspecific hybridization in plant genome evolution.
LITERATURE CITED