-
PDF
- Split View
-
Views
-
Cite
Cite
Minna Laatu, Hilpi Rautelin, Marja-Liisa Hänninen, Susceptibility of Campylobacter hyointestinalis subsp. hyointestinalis to antimicrobial agents and characterization of quinolone-resistant strains, Journal of Antimicrobial Chemotherapy, Volume 55, Issue 2, February 2005, Pages 182–187, https://doi.org/10.1093/jac/dkh537
Close - Share Icon Share
Abstract
Objectives:
To study the susceptibility of Campylobacter hyointestinalis subsp. hyointestinalis to several antimicrobial agents and to investigate the mechanisms of nalidixic acid and ciprofloxacin resistance.
Methods:
The disc diffusion method was employed to study the susceptibility of 49 C. hyointestinalis subsp. hyointestinalis strains of reindeer and bovine origin to 12 different antimicrobial agents. In addition, the MICs of nalidixic acid and ciprofloxacin were determined. The nucleotide sequence of a 270 bp fragment of the gyrA gene was determined in ciprofloxacin-susceptible and -resistant strains. The effect of a multidrug efflux pump inhibitor Phe-Arg-β-naphthylamide (PAβN) on the MICs of ciprofloxacin and nalidixic acid was also studied.
Results:
The only decreased susceptibility for antimicrobial agents of this study was observed for sulphonamide compound and streptomycin (24% and 32% of the strains, respectively), and this phenomenon was observed exclusively in the bovine strains. In sequence studies, a Thr-86→Ile change was found in strains with MICs of ciprofloxacin of ≥ 64 mg/L, but this mutation was absent in strains with lower resistance levels. The use of PAβN did not affect the MIC of ciprofloxacin but decreased the MIC of nalidixic acid 2–4-fold.
Conclusions:
The Finnish C. hyointestinalis subsp. hyointestinalis strains are susceptible to a majority of the antimicrobials of veterinary importance. The mechanism of ciprofloxacin resistance at lower levels (≤32 mg/L) is not associated with a specific mutation in the quinolone resistance-determining region of the gyrA gene. Finally, there are distinct differences in the mechanisms of ciprofloxacin resistance compared with nalidixic acid resistance within the studied species.
Introduction
Several Campylobacter species are important agents of gastroenteritis in humans. Whereas the principal Campylobacter spp. causing acute bacterial diarrhoea in humans are Campylobacter jejuni and Campylobacter coli, the importance of other Campylobacter species may have been underestimated due to difficulties in isolation and identification at the species level.1–3
Campylobacter hyointestinalis is one of the 16 described species in the genus Campylobacter and in 1995 this species was divided into two subspecies, namely subsp. hyointestinalis and subsp. lawsonii.4C. hyointestinalis was first isolated from pigs with proliferative enteritis in 1983,5 and later it was identified in various other animals, such as hamsters, cattle and reindeer.6,7 In 1986, C. hyointestinalis was isolated for the first time from a human patient suffering from proctitis,8 and subsequently a number of other cases have been reported involving the isolation of C. hyointestinalis from the stool samples of patients with watery diarrhoea.9 The reported cases concern mainly young children, elderly persons or those suffering from underlying illnesses. In some of the cases, the source of the infection could be traced to unpasteurized milk or an animal carrier,10 suggesting zoonotic food-borne transmission.
Nalidixic acid and ciprofloxacin are members of a large group of quinolones, antimicrobials that primarily target bacterial DNA gyrase, a type II topoisomerase.11 There is also clear evidence of cross-resistance within this class of agents.12 Relevant to this study, C. hyointestinalis possesses a distinctive characteristic of exhibiting high-level inherent resistance to nalidixic acid (128 mg/L)13 while remaining susceptible to ciprofloxacin.
Since the beginning of the use of fluoroquinolones as antimicrobials in human and veterinary medicine, increasing resistance levels to nalidixic acid and ciprofloxacin have been reported in zoonotic enteric Gram-negative pathogens such as C. jejuni, C. coli and Salmonella.14 Quinolone resistance has been associated with mutations in the quinolone resistance-determining region (QRDR) of the gyrA gene. To further explain the resistance mechanism, the role of an active efflux system that exports toxic compounds has been suggested,15 and more particularly it has been shown that CmeABC, an RND efflux pump, plays a crucial role in preventing the accumulation of many antimicrobial agents, including ciprofloxacin, in C. jejuni.16,17 Recently, Pumbwe et al.18 investigated the overexpression of cmeB and another newly identified RND efflux pump gene, cmeF, in C. jejuni isolates with multiple antibiotic resistance (MAR). cmeB alone or both of these genes were overexpressed in some of these MAR isolates, but no clear evidence was found for their role in ciprofloxacin resistance. The sequenced C. jejuni genome has revealed several other efflux pump genes that may also be involved in the resistance mechanisms of this organism to antimicrobial agents.19
This study was conducted with the purpose of establishing for the first time the susceptibility of C. hyointestinalis subsp. hyointestinalis to a wide range of antimicrobial agents, and to study induction and mechanisms of ciprofloxacin and nalidixic acid resistance. Also the role of the RND transporters in quinolone resistance was further assessed.
Materials and methods
Bacterial strains and growth conditions
C. hyointestinalis subsp. hyointestinalis strains used in this study were isolated from Finnish reindeer (Rangifer tarandus) and bovine faecal samples. The reindeer samples collected from eight different, exclusively reindeer processing slaughterhouses in northern Finland during 1998 have been characterized in detail in our previous publication.7 In addition to a large set of other tests, determination of H2S production is one of the most important tests in the speciation of C. hyointestinalis subsp. hyointestinalis. The bovine strains were obtained in 2003 (with the exception of strain B704, isolated in 1998) from bovine faecal samples by culturing on CAT (cefoperazone 2 mg/L, amphotericin B 2.5 mg/L and teicoplanin 1 mg/L) medium.20 Phenotypic species identification was done similarly to reindeer strains. All strains were stored in skimmed milk supplemented with 15% glycerol at −70°C before use.
Campylobacter strains were grown on Mueller–Hinton (MH) agar (Oxoid Ltd, London, UK) supplemented with 5% horse blood or in MH broth (Oxoid Ltd) and incubated at 37°C for 48 h under microaerobic conditions (5% O2, 10% CO2 and 85% N2) unless otherwise stated.
Antimicrobial susceptibility
The disc diffusion technique21 was used to screen the susceptibility of the studied strains to 12 different antibiotics (discs supplied by Oxoid Ltd). The antimicrobials studied were amoxicillin (10 μg), ciprofloxacin (5 μg), co-amoxiclav (30 μg), ampicillin (25 μg), doxycycline hydrochloride (30 μg), erythromycin (15 μg), gentamicin (10 μg), lincomycin (15 μg), metronidazole (5 μg), streptomycin (10 μg), sulphonamide compound S3 (sulfadiazine, sulfathiazole, sulfamerazine) (300 μg) and tetracycline (10 μg). After 2 days of incubation, the organisms were transferred from blood agar plates into 5 mL of MH broth and adjusted to a turbidity equivalent to that of a 0.5 McFarland standard. After inoculation, the plates were incubated for 48 h before reading the results. The data obtained were analysed using the WHONET 5.1 software (available at http://www.who.int/drugresistance/whonetsoftware/en/).
The agar dilution method was used to determine the MICs of nalidixic acid and ciprofloxacin. Approximately 105 organisms were inoculated on freshly prepared Petri dishes containing MH agar supplemented with 5% horse blood and increasing concentrations (1–512 mg/L) of nalidixic acid (Sigma, St Louis, MO, USA) initially dissolved in 0.2 N NaOH or increasing concentrations (0.064–512 mg/L) of ciprofloxacin (Sigma). The results were read after 48 h of incubation. The applied breakpoints were adapted from the NCCLS22 and strains with MICs of nalidixic acid of ≤8 mg/L were regarded as susceptible, >8 mg/L and <32 mg/L as intermediate and ≥32 mg/L as resistant, and strains with MICs of ciprofloxacin of ≤1 mg/L, >1 mg/L and <4 mg/L, and ≥4 mg/L were termed susceptible, intermediate and resistant, respectively.
C. jejuni 143483, a local control strain, originally isolated in Edinburgh, UK, was used as a control in susceptibility testing.23
Phe-Arg-β-naphthylamide efflux pump inhibitor
In assessing the role of the CmeABC efflux pump, the MICs of ciprofloxacin and nalidixic acid were determined as described above in the presence and absence of efflux pump inhibitor Phe-Arg-β-naphthylamide (PAβN) (Sigma) used at a concentration of 50 mg/L.24
Analysis of gyrA mutations
Three reindeer C. hyointestinalis subsp. hyointestinalis strains (PO0, PO61 and PO271) originally susceptible to ciprofloxacin were grown in gradually increasing concentrations of ciprofloxacin (0.5–64 mg/L). Each mutated strain was isolated at the stage where it had reached resistance to 16, 32 or 64 mg/L of ciprofloxacin.
Chromosomal DNA of the parent strains and each of their variants with induced resistance was extracted using a previously published modified method of Pitcher et al.25
A 270 bp sequence of the QRDR was amplified using the primers cjgyrA1 and cjgyrA2 as described by Piddock et al.26 PCR products were purified using the QIAquick PCR Purification Kit (Qiagen, Valencia, CA, USA) and subsequently sequenced using the automated cycle sequencer ABI 377XL with Big Dye terminators (PE Applied Biosystems, Warrington, UK).
Protein analysis
The whole cell protein patterns of the induced resistant variants and their parent strains were compared in 10% polyacrylamide gels by SDS–PAGE using Laemmli's method.27
Results
Antimicrobial susceptibility
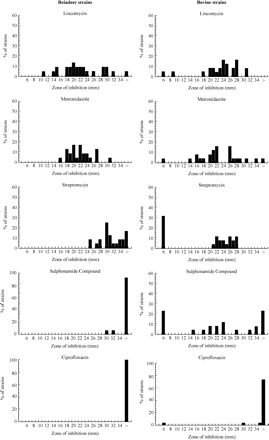
Twenty-five bovine C. hyointestinalis subsp. hyointestinalis strains and 24 reindeer C. hyointestinalis subsp. hyointestinalis strains were studied for resistance to antimicrobial agents using the disc diffusion method. All the studied organisms had distinct growth inhibition zones of at least 24 mm in the presence of gentamicin, erythromycin, ampicillin, co-amoxiclav, doxycycline, amoxicillin and tetracycline indicating susceptibility towards these substances (results not shown). Both sets of strains exhibited decreased susceptibility to metronidazole and lincomycin (Figure 1). The presence of sulphonamide compound or streptomycin generated no growth inhibition zone in, respectively 24% and 32% of the bovine strains, whereas all the reindeer strains had growth inhibition zones of at least 25 mm in the presence of these agents (Figure 1).
Ciprofloxacin susceptibility of the 49 strains from reindeer and cattle was studied (Figure 1) using the disc diffusion method, and apart from one exceptional bovine strain (B704), the observed inhibition zones measured at least 30 mm. These results were confirmed for the reindeer strains by the agar dilution method that resulted in values from 0.064 to 0.25 mg/L. The agar dilution method gave an MIC value of 32 mg/L for strain B704.
As expected, all 49 strains were resistant to nalidixic acid. The resistance of the reindeer strains was analysed by the agar dilution method, and the results indicated MICs of 64–128 mg/L. The MIC of nalidixic acid of the inherently ciprofloxacin-resistant strain (B704) was also determined and an even higher level of resistance was observed (MIC ≥512 mg/L). In strains that gained high-level resistance to ciprofloxacin, the nalidixic acid resistance also increased (MICs ≥512 mg/L).
Phe-Arg-β-naphthylamide efflux pump inhibitor
Involvement of the PAβN efflux pump inhibitor in the ciprofloxacin resistance of strain B704 and induced ciprofloxacin-resistant variants was studied in the presence and absence of PAβN. The presence of PAβN evoked no differences in MIC values of ciprofloxacin in any of the studied strains compared with MIC values without the inhibitor.
Involvement of an efflux pump in the nalidixic acid resistance was studied similarly. The MIC of nalidixic acid in the presence and absence of PAβN was established for eight reindeer C. hyointestinalis strains, the bovine strain B704, the ciprofloxacin-resistant variant strains and their parent strains. There was a 2–4 fold decrease in MICs for wild reindeer strains resulting in resistance levels of 16–32 mg/L. In induced ciprofloxacin-resistant variants showing higher levels of nalidixic acid resistance ( ≥512 mg/L) than the wild strains, the use of PAβN dropped the MIC to 64 mg/L, and the nalidixic acid resistance of the strain B704 decreased from ≥512 to 128 mg/L in the presence of PAβN (Table 1).
Analysis of gyrA mutations
The partial gyrA sequence obtained was submitted to BLASTn for sequence comparison against GenBank databases, which indicated that this sequence in the wild-type C. hyointestinalis subsp. hyointestinalis was most closely similar to the equivalent sequence in Campylobacter fetus subsp. fetus, and the derived amino acid sequence differed from it in only one position (Asn-99→Ser).
The inherently ciprofloxacin-resistant strain B704 differed from the susceptible strains by the presence of isoleucine instead of threonine at position 86 (C. jejuni numeration) of the GyrA (Table 1). The induced resistant variants of the C. hyointestinalis strains initially susceptible to ciprofloxacin with MICs of ≥64 mg/L also possessed the Thr-86→Ile mutation (Table 1). This amino acid change was due to a single point mutation of cytosine to thymine in the nucleotide sequence. This mutation was not present in strains with MICs of ciprofloxacin of ≤32 mg/L. There appeared to be no other mutations in the sequenced region.
Protein analysis
Visual comparison of SDS–PAGE patterns of the three wild strains and their induced ciprofloxacin-resistant variants (MICs of 16 and 64 mg/L) showed no obvious differences (results not shown).
Discussion
The antimicrobial spectrum used in the susceptibility testing was based on their veterinary and/or human medical use. Because some studies have reported increased resistance of C. jejuni to metronidazole,28 this compound was also included in our study. In dairy cattle, the most important disease requiring antimicrobial treatment is mastitis, for which β-lactams, aminoglycosides, sulphonamides, tetracyclines and macrolides are used. Enrofloxacin and danafloxacin are fluoroquinolones that are used for treating mastitis, and respiratory and intestinal infections in cattle. Reindeer, in contrast, live and feed freely as semi-domesticated animals and are only rarely treated with antimicrobials. In 2002, a total amount of 13 000–15 000 kg of antimicrobials, including about 70 kg of enrofloxacin and danafloxacin as injectables, was used in veterinary medicine in Finland and approximately 60% of this was used for treating cattle.29 Compared with the number of cattle (1.2 million), the use of antimicrobials in Finland is moderate. The results of our study indicate that C. hyointestinalis. subsp. hyointestinalis remains very susceptible to most of the studied substances, supporting the trend of moderate use of antimicrobials in cattle in Finland.
However, cattle and reindeer strains differed distinctly in their susceptibility to streptomycin and sulphonamide compound, as no growth inhibition was observed in 32% and 24% of the bovine strains, respectively, whereas all the organisms of reindeer origin produced distinct inhibition zones. This phenomenon is likely to reflect the veterinary use of these substances for the treatment of mastitis in cattle.
There was one particular bovine strain exhibiting high-level ciprofloxacin resistance (MIC 32 mg/L) and in the course of this study, we induced high-level ciprofloxacin resistance in three inherently susceptible strains. This allowed us to establish that in comparison to susceptible strains, highly resistant C. hyointestinalis subsp. hyointestinalis strains possess a Thr-86→Ile change in their GyrA. This same mutation has previously been reported in nalidixic acid and ciprofloxacin-resistant C. jejuni and C. coli.30 However, in our study, this mutation was present only in highly ciprofloxacin-resistant (i.e. MIC ≥ 64 mg/L) strains, and since the MIC for C. hyointestinalis is naturally at the level of about 0.1 mg/L and there were no other mutations in the QRDR of gyrA, there must be some other mechanisms that would explain the occurrence of the lower level of resistance.
The amino acid sequence of the QRDR of GyrA in wild-type C. hyointestinalis subsp. hyointestinalis shared 73% identity with that of C. jejuni, in which the mutations at Ala-70, Thr-86 or Asp-90 in the QRDR have been associated with nalidixic acid resistance.31 However, at these locations C. hyointestinalis subsp. hyointestinalis possessed the same amino acids as wild-type C. jejuni, thereby suggesting that the inherent resistance of C. hyointestinalis to nalidixic acid is not explained by the same mechanism as the resistance in C. jejuni strains.
In addition to C. hyointestinalis, two other Campylobacter species are inherently nalidixic acid-resistant, i.e. C. fetus and Campylobacter lari.9 The amino acid sequence of the partial GyrA of C. hyointestinalis shared high similarity (98%) with the respective sequence of C. fetus.32 The reported mutation at position Asp-91 of C. fetus associated with lower-level ciprofloxacin resistance (MIC 8–16 mg/L) was not found in our study of C. hyointestinalis.
In order to reveal the role of the previously described CmeABC efflux pump in nalidixic acid and ciprofloxacin resistance of C. hyointestinalis subsp. hyointestinalis, we conducted an investigation using PAβN, an efflux pump inhibitor that has been shown to be efficient in restoring the efficacy of antimicrobial agents, such as ciprofloxacin, against Escherichia coli, Pseudomonas aeruginosa and C. jejuni.17,33,34 Nevertheless, it had no effect on ciprofloxacin-resistant C. hyointestinalis in this study. It did, however, decrease the MIC of nalidixic acid 2–4 fold.
In conclusion, our studies showed that Finnish C. hyointestinalis strains remain mainly susceptible to most antimicrobials of veterinary importance, but with some decreased susceptibility to streptomycin and sulphonamide compound in bovine strains, implicating the use of these substances in cattle. The high-level ciprofloxacin resistance was associated with a specific mutation in the gyrA gene, but the QRDR sequence did not explain the nalidixic acid resistance, which instead was associated with the CmeABC-like pump. The mechanisms of nalidixic acid resistance and acquired ciprofloxacin resistance at MIC levels ≤32 mg/L need further study.
Susceptibility of 25 bovine and 24 reindeer C. hyointestinalis subsp. hyointestinalis strains to lincomycin, metronidazole, streptomycin, sulphonamide compound and ciprofloxacin as tested by the disc diffusion method; 6 mm is the diameter of the disc and indicates the complete absence of inhibition zone.
Characteristics of the Campylobacter hyointestinalis subsp. hyointestinalis strains selected in vitro
| Bacterial strain . | Susceptibility to ciprofloxacin . | CIP induction level (mg/L) . | NAL MIC (mg/L)a . | Amino acid 86 in QRDR . |
|---|---|---|---|---|
| B704 | inherently resistant | 32 | ≥512 (128) | isoleucine |
| PO0 | inherently susceptible | – | 64 (32) | threonine |
| PO0Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO0Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO0Cip64 | induced resistance | 64 | ND | isoleucine |
| PO61 | inherently susceptible | – | 128 (32) | threonine |
| PO61Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO61Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO61Cip64 | induced resistance | 64 | ND | isoleucine |
| PO271 | inherently susceptible | – | 128 (32) | threonine |
| PO271Cip16 | induced resistance | 16 | ND | threonine |
| PO271Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO271Cip64 | induced resistance | 64 | ND | isoleucine |
| Eight randomly selected strains | inherently susceptible | – | 64–128 (16–32) | ND |
| Bacterial strain . | Susceptibility to ciprofloxacin . | CIP induction level (mg/L) . | NAL MIC (mg/L)a . | Amino acid 86 in QRDR . |
|---|---|---|---|---|
| B704 | inherently resistant | 32 | ≥512 (128) | isoleucine |
| PO0 | inherently susceptible | – | 64 (32) | threonine |
| PO0Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO0Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO0Cip64 | induced resistance | 64 | ND | isoleucine |
| PO61 | inherently susceptible | – | 128 (32) | threonine |
| PO61Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO61Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO61Cip64 | induced resistance | 64 | ND | isoleucine |
| PO271 | inherently susceptible | – | 128 (32) | threonine |
| PO271Cip16 | induced resistance | 16 | ND | threonine |
| PO271Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO271Cip64 | induced resistance | 64 | ND | isoleucine |
| Eight randomly selected strains | inherently susceptible | – | 64–128 (16–32) | ND |
CIP, ciprofloxacin; NAL, nalidixic acid; ND, not done.
Values in brackets correspond to the MIC in the presence of the efflux pump inhibitor Phe-Arg-β-naphthylamide at 50 mg/L.
Characteristics of the Campylobacter hyointestinalis subsp. hyointestinalis strains selected in vitro
| Bacterial strain . | Susceptibility to ciprofloxacin . | CIP induction level (mg/L) . | NAL MIC (mg/L)a . | Amino acid 86 in QRDR . |
|---|---|---|---|---|
| B704 | inherently resistant | 32 | ≥512 (128) | isoleucine |
| PO0 | inherently susceptible | – | 64 (32) | threonine |
| PO0Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO0Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO0Cip64 | induced resistance | 64 | ND | isoleucine |
| PO61 | inherently susceptible | – | 128 (32) | threonine |
| PO61Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO61Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO61Cip64 | induced resistance | 64 | ND | isoleucine |
| PO271 | inherently susceptible | – | 128 (32) | threonine |
| PO271Cip16 | induced resistance | 16 | ND | threonine |
| PO271Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO271Cip64 | induced resistance | 64 | ND | isoleucine |
| Eight randomly selected strains | inherently susceptible | – | 64–128 (16–32) | ND |
| Bacterial strain . | Susceptibility to ciprofloxacin . | CIP induction level (mg/L) . | NAL MIC (mg/L)a . | Amino acid 86 in QRDR . |
|---|---|---|---|---|
| B704 | inherently resistant | 32 | ≥512 (128) | isoleucine |
| PO0 | inherently susceptible | – | 64 (32) | threonine |
| PO0Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO0Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO0Cip64 | induced resistance | 64 | ND | isoleucine |
| PO61 | inherently susceptible | – | 128 (32) | threonine |
| PO61Cip16 | induced resistance | 16 | ≥512 (64) | threonine |
| PO61Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO61Cip64 | induced resistance | 64 | ND | isoleucine |
| PO271 | inherently susceptible | – | 128 (32) | threonine |
| PO271Cip16 | induced resistance | 16 | ND | threonine |
| PO271Cip32 | induced resistance | 32 | ≥512 (64) | threonine |
| PO271Cip64 | induced resistance | 64 | ND | isoleucine |
| Eight randomly selected strains | inherently susceptible | – | 64–128 (16–32) | ND |
CIP, ciprofloxacin; NAL, nalidixic acid; ND, not done.
Values in brackets correspond to the MIC in the presence of the efflux pump inhibitor Phe-Arg-β-naphthylamide at 50 mg/L.
This study was supported by grants from the Academy of Finland (to M. L.) and the Walther Ehrström Foundation.
References
Lastovica, A. J. & Skirrow, M. B. (
Skirrow, M. B. & Blaser, M. J. (
Engberg, J., On, S. L., Harrington, C. S. et al. (
On, S. L., Bloch, B., Holmes, B. et al. (
Gebhart, C. J., Ward, G. E., Chang, K. et al. (
Walder, M., Sandstedt, K. & Ursing, J. (
Hanninen, M. L., Sarelli, L., Sukura, A. et al. (
Fennell, C. L., Rompalo, A. M., Totten, P. A. et al. (
Edmonds, P., Patton, C. M., Griffin, P. M. et al. (
Gorkiewicz, G., Feierl, G., Zechner, R. et al. (
Drlica, K. & Zhao, X. (
Barry, A. L. & Fuchs, P. C. (
Gebhart, C. J., Edmonds, P., Ward, G. E. et al. (
McEwen, S. A. & Fedorka-Cray, P. J. (
Charvalos, E., Tselentis, Y., Hamzehpour, M. M. et al. (
Lin, J., Michel, L. O. & Zhang, Q. (
Luo, N., Sahin, O., Lin, J. et al. (
Pumbwe, L., Randall, L. P., Woodward, M. J. et al. (
Parkhill, J., Wren, B. W., Mungall, K. et al. (
Aspinall, S. T., Wareing, D. R., Hayward, P. G. et al. (
Bauer, A. W., Kirby, W. M., Sherris, J. C. et al. (
National Committee for Clinical Laboratory Standards. (
Hakanen, A., Huovinen, P., Kotilainen, P. et al. (
Saenz, Y., Ruiz, J., Zarazaga, M. et al. (
Pitcher, D. G., Saunders, N. A. & Owen, R. J. (
Piddock, L. J., Ricci, V., Pumbwe, L. et al. (
Laemmli, D. K. (
Jones, K. & Stanley, K. N. (
National Veterinary and Food Research Institute. (2004). [Online.] http://www.nam.fi/uploads/laakeinfo/ellaakkeet2002.pdf (8 july 2004, date last accessed).
Bachoual, R., Ouabdesselam, S., Mory, F. et al. (
Wang, Y., Huang, W. M. & Taylor, D. E. (
Taylor, D. E. & Chau, A. S. (
Lomovskaya, O., Warren, M. S., Lee, A. et al. (
Author notes
1Department of Food and Environmental Hygiene, Faculty of Veterinary Medicine, PO Box 57, FIN-00014 University of Helsinki; 2Department of Bacteriology and Immunology, Haartman Institute, University of Helsinki and HUSLAB, Helsinki University Central Hospital, Helsinki, Finland