-
PDF
- Split View
-
Views
-
Cite
Cite
Rebecca J. Garten, Jinbing Zhang, Shenghan Lai, Wei Liu, Jie Chen, Xiao-Fang Yu, Coinfection with HIV and Hepatitis C Virus among Injection Drug Users in Southern China, Clinical Infectious Diseases, Volume 41, Issue Supplement_1, July 2005, Pages S18–S24, https://doi.org/10.1086/429491
Close - Share Icon Share
Abstract
Background. We sought to examine coinfection with human immunodeficiency virus (HIV) and hepatitis C virus (HCV) among injection drug users (IDUs) in Guangxi, China.
Methods. A longitudinal cohort of IDUs (547 subjects) was established to study risk factors for bloodborne infections. At each visit, participants completed questionnaires defining demographic characteristics, patterns of drug use, and sexual behaviors. Blood samples were collected and analyzed for the presence and genotype of HIV and HCV.
Results. Coinfection with HIV and HCV was found in 17.6% of the IDUs. HCV was present in 95.1% of HIV-positive and 70.4% of HIV-negative heroin users. The prevalence of HIV in HCV-positive and HCV-negative heroin users was 23.4% and 3.6%, respectively. Multivariate logistic regression analysis revealed that sexual activity during the past 6 months and duration of injection drug use were significantly associated with coinfection with HIV and HCV. The main circulating HCV genotypes included 6a (38%), 3b (37%), and 1a (19%), whereas genotypes 6e (4%), 3a (2%), and 1b (1%) were present in only a few IDUs. Multiple HCV genotypes were present at each study site and did not segregate by HIV status or subtype.
Conclusions. HCV is highly prevalent in IDUs throughout southern China. In Guangxi, HIV infections are the result of parenteral and sexual transmission, and, therefore, all IDUs are at high risk of coinfection with HIV and HCV. Molecular tracking of HCV may be a more sensitive predictor of the future spread of the HIV-1 epidemic than is HIV subtyping. This study emphasizes that, without implementation of injection prevention and primary substance abuse programs in China, the extent and effect of coinfection with HIV and HCV will only increase.
Coinfection with HIV and hepatitis C virus (HCV) is a significant global public health problem. This is especially important in populations of drug users, among whom the prevalence of coinfection with HIV and HCV can surpass 90% [1–3]. Coinfection with HIV-1 has been shown to increase HCV loads, hasten the development of cirrhosis, hasten progression to end-stage liver disease, and be a negative prognostic factor for clearance of HCV [4–8]. Although the findings are controversial, in some cases, coinfection with HIV and HCV results in a faster progression to AIDS [9, 10]. Most important, coinfection with HIV and HCV will continue to increase morbidity and mortality in areas such as China, where treatment for either virus is unavailable [11].
The HIV epidemic has been carefully followed since its emergence in southern China, with little emphasis on HCV coinfection. More than 70% of injection drug users (IDUs) in Guangxi Province are infected with HCV [12]. The incidence of HCV infection was found to be extremely high, at 37.6 cases/100 person-years. HCV is usually rapidly acquired after onset of injection drug use through high-risk injection behaviors, whereas HIV is acquired after increasing durations of injection drug use. HIV continues to spread in IDUs at an incidence of ∼7.0 cases/100 person-years, with a prevalence of ∼20%. The major mode of HIV transmission in the province continues to be injection drug use, but sexual transmission is also occurring [13].
Few population-based studies of coinfection with HIV and HCV among drug users in China have been performed, especially with regard to the HCV genotypes present. The aim of the present study was to determine HCV genotypes within the study population and investigate coinfection with HIV and HCV, to identify the risk factors for coinfections among drug users in Guangxi Province, southern China.
Subjects, Materials, and Methods
Study participants. Five hundred forty-seven participants, heroin users ⩾18 years old, were enrolled at study sites in the cities of Pingxiang and Binyang in Guangxi Province. The informed consent procedure has been described elsewhere [14]. In brief, at each visit, participants underwent a physical examination, venipuncture, and an interview about their health and drug and sex risk behaviors. Blood was collected and centrifuged, and the serum either underwent serological assays or was stored at -70°C. Samples were then shipped from Guangxi to our laboratory in Baltimore for further analysis.
Serological assays. At the Guangxi Center for Disease Control and Prevention in Nanning, the presence of HIV antibody was determined by ELISA with the Vironostika HIV-1 Microelisa System (Organon Teknika). All samples that were positive by ELISA were not considered to be positive for HIV until confirmation by a Western blot immunoassay for HIV-1 and -2 (Gene Lab). Antibody to HCV was detected by use of the Ortho HCV ELISA Test System (version 3.0; Ortho Diagnostic Systems).
HCV RNA extraction, PCR amplification, and sequencing. RNA for HCV genotyping was extracted from 140 µL of serum, according to the manufacturer's protocol, by use of the QIAamp Viral RNA kit (QIAGEN). Reverse transcription and nested PCR were done with primers to conserved regions of core and E1, as described elsewhere [15]. After purification with the QIAquick PCR Purification kit (QIAGEN), samples were sequenced by use of the inner forward primer on an automated sequencer (PRISM, version 2.1.1; ABI).
Phylogenetic analyses. Phylogenetic analyses of a 381-bp region of the beginning of HCV E1 were used to determine HCV genotype. Sequences were compiled by use of the BioEdit program (version 5.09) [16] and CLUSTAL_X [17]. Genotypes were assigned after alignment with reference sequence from the HCV database at Los Alamos (available at: http://hcv.lanl.gov/content/hcv-db/GET_ALIGNMENTS/alignments.html). The following controls were used to construct trees: 1a.HC J1 (D00831), 1a.HCV 1 (M62321), 1a.HCV H (M67463), 1b.ARG2 (M74815), 1b.HCV N (AF139594), 1b.HEBEI (L02836), 1c.AY051292 (AY051292), 1c.SR037 (D16191), 2a.AF403230 (AF403230), 2b.DK8 (L16657), 2c.BEBE1 (D50409), 2e.JK020 (D49745), 2f.JK081 (D49754), 2i.HN4 (X76415), 2k.VAT96 (AB031663), 3a.CB (AF046866), 3a.DK12 (L16630), 3a.K3A (D28917), 3b.HCV Tr (D49374), 3b.ST (D11443), 3b.TH527 (D37839), 3c.NE048 (D16612), 3d.NE274 (D16620), 3e.NE145 (D16618), 3f.NE125 (D16614), 3h.SOM1 (AF216786), 3h.SOM2 (AF216787), 3h.SOM3 (AF216788), 3k.JK030 (D49747), 3k.JK055 (D49750), 3k.JK070 (D49752), 4a.ED43 (Y11604), 4c.Z6 (L16678), 4d.DK13 (L16656), 4r.1196E1 4 (D43677), 5a.EUH1480 (Y13184), 5a.FR741 (D50466), 5a.SA13 (AF064490), 6a.EUHK2 (Y12083), 6a.HK2 (L16634), 6a.VN506 (D88469), 6b.NB56 (AY231583), 6b.TH580 (D37841), 6d.VN235 (D84263), 6e.VN787 (D88477), 6e.VN843 (D88478), 6f.TH271 (D37844), 6f.TH552 (D37845), 6f.TH976 (D37846), 6g.JK046 (D49748), 6g.JK065 (D49751), 6h.VN004 (D84265), 6h.VN085 (D88466), 6i.TH555 (D37849), 6i.TH602 (D37850), 6j.TH553 (D37848), 6k.VN405 (D84264), and 6k.VN530 (D88471). Phylogenetic analysis was done with PAUP 4.0 [18], and trees were created by use of the optimality criteria of parsimony and minimum evolution with maximum-likelihood distances. For this large data set, analyses that used likelihood evaluations were not computationally feasible at the time of publication. The GTR + γ + I model of substitution, base frequencies, γ distribution, invariant sites, transition-to-transversion ratios, and rate matrix values were determined by use of Modeltest 3.5 [19]. Bootstrap analysis (100 replicates) was done on each minimum parsimony and evolution tree. Both analyses (minimum parsimony and evolution) gave congruent results, and the minimum evolution bootstrap value is shown.
Nucleotide sequence accession numbers. The HCV partial C/E1 nucleotide sequences determined in this study have been deposited in the GenBank sequence database and have been assigned the accession numbers AY878411–AY878536.
Statistical analysis. SAS software package (version 8.0; SAS Institute) was used for management and analysis of study data. Statistical analysis consisted of univariate analyses and multivariate analyses. Univariate analysis was done with χ2 or Fisher's exact test for each specific risk factor for coinfection and HCV genotype separately. Multivariate analysis was done with the multivariate logistic regression model. For each of the 2 dependent variables, HCV infection and coinfection with HIV and HCV, the factors that were significant at a level of .10 in the univariate analysis were integrated into the logistic regression model. Backward elimination was applied to shape the final model. For a categorical factor with <2 categories, some dichotomous variables were created and integrated into the model. In the final model, if some dichotomous variables corresponding to this variable existed, then all others were also integrated into the final model. Afterward, pairwise interactions among factors were also integrated into the regression model, and a manual elimination procedure was applied to build the final model.
Results
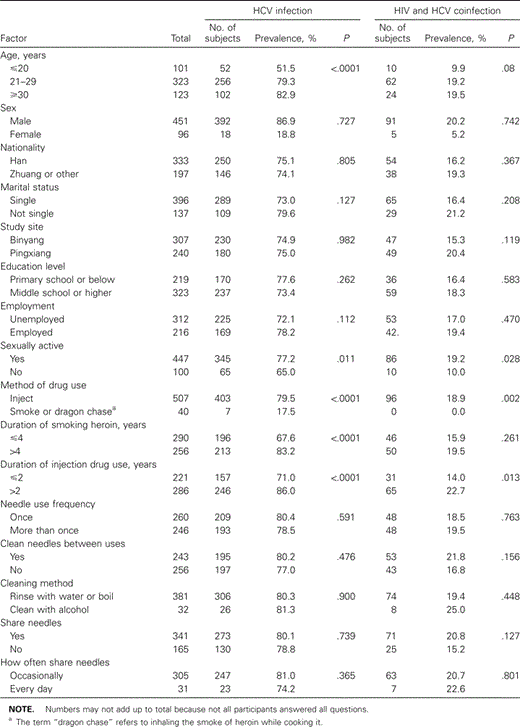
Prevalence of coinfection with HIV and HCV among drug users in Guangxi, China. A total of 547 heroin users from Guangxi Province were investigated in this study. As shown in table 1, prevalences of HCV and HIV were 75.0% and 18.5%, respectively. The prevalence of coinfection with HIV and HCV was 17.6%. The prevalence of HIV infection was 23.4% among HCV-positive heroin users and 3.6% among HCV-negative heroin users. Table 2 lists the results of univariate analysis with χ2 or Fisher's exact test. The results showed that age, method of drug use, duration of smoking, duration of injection drug use, and sexual activity in the past 6 months were associated with HCV infection, whereas method of drug use, duration of injection drug use, and sexual activity in the past 6 months were associated with coinfection with HIV and HCV.
Coinfection with HIV and hepatitis C virus (HCV) in 547 heroin users in the Guangxi cohort.
Risk factors associated with coinfection with HIV and hepatitis C virus (HCV) among 547 heroin users enrolled in the Guangxi cohort.
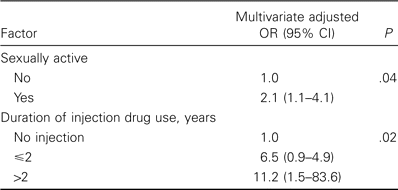
Multivariate analysis was done by logistic regression. For coinfection with HIV and HCV, the method of drug use, duration of injection drug use, and sexual activity in the past 6 months were integrated into the logistic regression model. The results showed that sexual activity and duration of injection drug use were independently associated with coinfection with HIV and HCV (table 3).
Multivariate logistic regression analysis for factors associated with coinfection with HIV and hepatitis C virus among 547 heroin users in the Guangxi cohort.
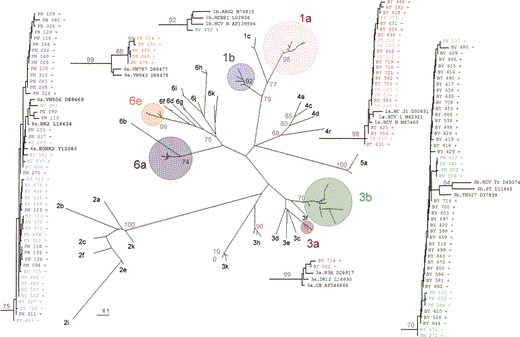
HCV genotyping. HCV genotype was determined for 126 injection drug users (IDUs) from 2 sites within Guangxi Province, southern China. The 2 study sites are geographically separated—Pingxiang is in the south, bordering Vietnam, whereas Binyang is near the capital city of Nanning in the center of the province. Samples obtained from all available HIV-coinfected persons (n = 70), as well as a random sample of non—HIV-infected persons (n = 56), were included in this study. Samples from Guangxi IDUs were classified into genotypes 1a, 1b, 3a, 3b, 6a, and 6e on the basis of phylogenetic analysis of a partial env region (figure 1). Surprisingly, there was no significant clustering within each study site. The majority of IDUs were infected with genotypes 6a (38%), 3b (37%), and 1a (19%), whereas genotypes 6e (4%), 3a (2%), and 1b (1%) were present in only a few IDUs.
Hepatitis C virus phylogenetic tree for 126 injection drug users from Guangxi Province, southern China. The tree was constructed by the maximum evolution method by use of maximum likelihood distances with nucleotide sequences corresponding to the E1 region. Bootstrap values <70% are indicated at the nodes of the corresponding branches. PN, Pingxiang; BY, Binyang; +, HIV positive; -, HIV negative.
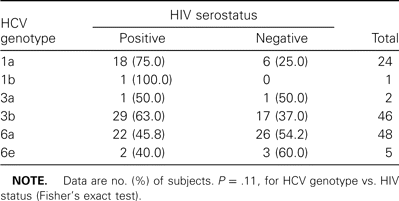
HCV genotypes present in HIV-coinfected persons did not significantly differ from those found in HIV-negative IDUs (table 4). HCV genotype 1a was represented in more HIV-positive than HIV-negative IDUs, whereas the remaining HCV genotypes were relatively equally represented.
Hepatitis C virus (HCV) genotypes, by HIV serostatus, among 126 heroin users in the Guangxi cohort.
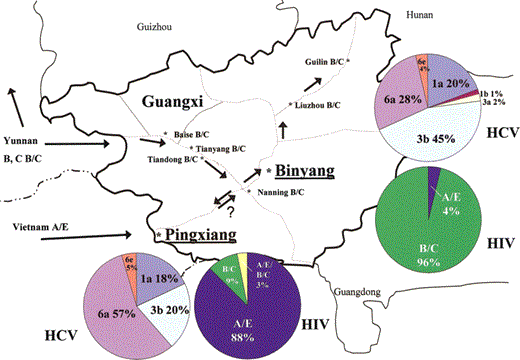
The study sites are not only geographically separated but also are on 2 separate heroin trafficking routes (figure 2). Pingxiang is situated on the Vietnam border and has been experiencing an epidemic of infection due to HIV A/E subtype originating from Vietnam. Binyang is centrally located within the province and, like the majority of cites in Guangxi, has an ongoing epidemic of infection due to HIV B/C recombinant subtype. Unlike the segregated HIV epidemics between these cities, HCV genotypes did not differ by HIV subtype epidemic. HCV genotypes 1a, 3b, 6a, and 6e were found in both Pingxiang and Binyang, whereas genotypes 3a and 1b were present in only a few IDUs from Binyang (figure 2). In Pingxiang, the majority of samples were genotype 6a (57%), whereas, in Binyang, most samples were genotype 3b (45%).
Map of HIV and hepatitis C virus epidemics in Guangxi Province. Arrows indicate heroin trafficking routes. Dotted lines indicate major highways within the province. A/E, HIV CRF01_AE; B/C, HIV CRF08_BC.
Discussion
Injection drug use continues to fuel the HIV and HCV epidemics spreading throughout southern China. In Guangxi Province, injection drug use continues to be the primary risk factor for the acquisition of HCV and HIV, with HCV usually acquired first, very early after onset of injection drug use. Sexual transmission of HIV within the cohort is likely occurring, because a small proportion of HIV-positive persons are negative for HCV, and sexual activity continues to be associated with both HCV seroprevalence and coinfection with HIV and HCV. Although sexual transmission of HCV is still controversial, it is possible that sexual activity also represents a marker for other high-risk factors, such as sharing injection paraphernalia (e.g., “cookers” [small containers used to dissolve drug], filters, or rinse water) or sharing drug by front loading (i.e., using one syringe to squirt drug into the front of another syringe after removing the needle) [20, 21].
IDUs in China do not routinely have access to treatment for HCV or HIV infection. Similar rates of infection with HCV and coinfection with HIV and HCV among IDUs have been found in Sichuan Province (11.3%), whereas neighboring Yunnan Province reports even higher prevalence of coinfection among IDUs (71.9%) [2, 3]. HIV coinfections may dramatically increase the progression of hepatic fibrosis, risk of end-stage liver disease, risk of hepatocellular carcinoma, and death in Chinese IDUs infected with HCV. Without serious implementation of prevention strategies and treatment options, the morbidity and mortality associated with coinfection with HIV and HCV among Chinese IDUs will only be increasing in the years to come.
HCV genotypes 1a and 3a are highly represented in IDU communities in Europe, whereas genotypes 1a and 1b are the majority in the United States [22–26]. The majority of persons in Europe and the United States who are coinfected with HIV and HCV are infected with HIV subtype B [27]. In our cohort, 3 main HCV genotypes, 1a, 3b, and 6a, are circulating in relatively equal proportions. Genotypes 3 and 6 are common throughout Southeast Asia, especially in Vietnam and Thailand [28, 29]. The study sites of Binyang and Pingxiang were originally chosen because of their separate HIV epidemics of 2 recombinant strains, CRF01_AE (A/E) and CFR08_BC (B/C). Our recent study found that the HIV B/C strain is now present in the city of Pingxiang, where only the A/E strain was previously circulating [13]. IDUs in Pingxiang were previously thought to have little interaction with IDUs from other cities within Guangxi. HCV genotyping did not show significantly different strains between Binyang and Pingxiang, although Binyang had additional genotypes. These results suggest that there are currently more exchanges within Guangxi cities than were seen before. Because HCV is rapidly acquired after onset of injection drug use, HCV genotyping may be a more sensitive risk predictor of the spread of HIV epidemics than is HIV subtyping. Although Southeast Asia has very high rates of HCV infections, very few studies in these HCV genotypes and HIV subtypes have been done.
In this study, we found HIV coinfection resulting from both injection drug use and sexual transmission in IDUs infected with HCV. Social and economic reforms in China have coincided with increased injection drug use and relaxed attitudes toward sex before marriage [30–32]. Persons coinfected with HIV and HCV did not cluster separately from HIV-negative persons. HIV is therefore not restricted to the subpopulation of the cohort, and all HCV-infected persons are at risk for HIV coinfection. HCV genotyping revealed higher levels of interactions throughout the province, which may predict major changes in current HIV epidemics. Without adequate treatment options or effective prevention strategies in drug users, coinfection with HIV and HCV in China will continue to rise, resulting in higher disease burden.
Acknowledgments
Financial support. National Institutes of Health (grant DA-16540 to X.-F.Y.).
Potential conflicts of interest. All authors: no conflicts.






Comments