Abstract
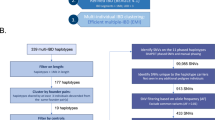
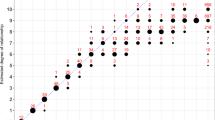
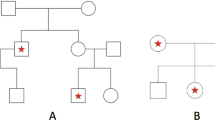
A recent report showed significant associations between several SNPs in a previously unknown EST cluster with schizophrenia.1 The cluster was identified as the human dystrobrevin binding protein 1 gene (DTNBP1) by sequence database comparisons and homology with mouse DTNBP1.2 However, the linkage disequilibrium (LD) among the SNPs in DTNBP1 as well as the pattern of significant SNP–schizophrenia association was complex. This raised several questions such as the number of susceptibility alleles that may be involved and the size of the region where the actual disease mutation(s) could be located. To address these questions, we performed different single-marker tests on the 12 previously studied and 2 new SNPs in DTNBP1 that were re-scored using an improved procedure, and performed a variety of haplotype analyses. The sample consisted of 268 Irish multiplex families selected for high density of schizophrenia. Results suggested a simple structure where the LD in the target region could be explained by 6 haplotypes that together accounted for 96% of haplotype diversity in the whole sample. From these six, a single high-risk haplotype was identified that showed a significant association with schizophrenia and explained the pattern of significant findings in the analyses with individual markers. This haplotype was 30 kb long, had a large effect, could be measured with two tag SNPs only, had a frequency of 6% in our sample, seemed to be of relatively recent origin in evolutionary terms, and was equally distributed over Ireland. Implications of these findings for follow-up and replication studies are discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Gershon ES, DeLisi LE, Hamovit J, Nurnberger Jr, JI, Maxwell ME, Schreiber J et al. A controlled family study of chronic psychoses. Schizophrenia and schizoaffective disorder. Arch Gen Psychiatry. 1988; 45: 328–336.
Kendler KS, McGuire M, Gruenberg AM, O'Hare A, Spellman M, Walsh D . The Roscommon family study: I. Methods, diagnosis of probands, and risk of schizophrenia in relatives. Arch Gen Psychiatry 1993; 50: 527–540.
Gottesman II, Shields J . Schizophrenia in twins: 16 years' consecutive admissions to a psychiatric clinic. Br J Psychiatry 1966; 112: 809–818.
Kendler KS . Overview: a current perspective on twin studies of schizophrenia. Am J Psychiatry 1983; 140: 1413–1425.
Farmer AE, McGuffin P, Gottesman II . Twin concordance for DSM-III schizophrenia. Scrutinizing the validity of the definition. Arch Gen Psychiatry 1987; 44: 634–640.
Kety SS . The significance of genetic factors in the etiology of schizophrenia: results from the national study of adoptees in Denmark. J Psychiatr Res 1987; 21: 423–429.
Tienari P, Wynne LC, Moring J, Lahti I, Naarala M, Sorri A et al. The Finnish adoptive family study of schizophrenia. Implications for family research. Br J Psychiatry 1994; 17(Suppl): 20–26.
McGue M, Gottesman I . The genetic epidemiology of schizophrenia and the design of linkage studies. Eur Arch Psychiatry Clin Neurosci 1991; 240: 174–181.
Coon H, Jensen S, Holik J, Hoff M, Myles-Worsley M, Reimherr F et al. Genomic scan for genes predisposing to schizophrenia. Am J Med Genet 1994; 54: 59–71.
Moises HW, Yang L, Kristbjarnarson H, Wiese C, Byerley W, Macciardi F et al. An international two-stage genome-wide search for schizophrenia susceptibility genes. Nat Genet 1995; 11: 321–324.
Pulver AE, Lasseter VK, Kasch L, Wolyniec P, Nestadt G, Blouin JL et al. Schizophrenia: a genome scan targets chromosomes 3p and 8p as potential sites of susceptibility genes. Am J Med Genet Neuropsychiatr Genet 1995; 60: 252–260.
Blouin JL, Dombroski BA, Nath SK, Lasseter VK, Wolyniec PS, Nestadt G et al. Schizophrenia susceptibility loci on chromosomes 13q32 and 8p21. Nat Genet 1998; 20: 70–73.
Faraone SV, Matise T, Svrakic D, Pepple J, Malaspina D, Suarez B et al. Genome scan of European-American schizophrenia pedigrees: results of the NIMH genetics initiative and millennium consortium. Am J Med Genet Neuropsychiatr Genet 1998; 81: 290–295.
Garver DL, Barnes R, Holcombe J, Filbey F, Wilson R, Bowcock A . Genome-wide scan and schizophrenia in African Americans. Am J Med Genet Neuropsychiatr Genet 1998; 81: 454–455.
Kaufmann CA, Suarez B, Malaspina D, Pepple J, Svrakic D, Markel PD et al. NIMH genetics initiative millennium schizophrenia consortium: linkage analysis of African-American pedigrees. Am J Med Genet Neuropsychiatr Genet 1998; 81: 282–289.
Levinson DF, Mahtani MM, Nancarrow DJ, Brown DM, Kruglyak L, Kirby A et al. Genome scan of schizophrenia. Am J Psychiatry 1998; 155: 741–750.
Shaw SH, Kelly M, Smith AB, Shields G, Hopkins PJ, Loftus J et al. A genome-wide search for schizophrenia susceptibility genes. Am J Med Genet Neuropsychiat Genet 1998; 81: 364–376.
Hovatta I, Varilo T, Suvisaari J, Terwilliger JD, Olikainen V, Arajärvi R et al. A genome-wide screen for schizophrenia genes in an isolated Finnish subpopulation suggesting multiple susceptibility loci. Am J Hum Genet 1999; 65: 1114–1124.
Williams NM, Rees MI, Holmans P, Norton N, Cardno AG, Jones LA et al. A two-stage genome scan for schizophrenia susceptibility genes in 196 affected sibling pairs. Hum Mol Genet 1999; 8: 1729–1739.
Rees MI, Fenton I, Williams NM, Holmans P, Norton N, Cardno A et al. Autosome search for schizophrenia susceptibility genes in multiply affected families. Mol Psychiatry 1999; 4: 353–359.
Brzustowicz LM, Hodgkinson KA, Chow EWC, Honer WG,, Bassett AS . Location of a major susceptibility locus for familial schizophrenia on chromosome 1q21–q22. Science 2000; 288: 678–682.
Ekelund J, Lichtermann D, Hovatta I, Ellonen P, Suvisaari J, Terwilliger JD et al. Genome-wide scan for schizophrenia in the Finnish population: evidence for a locus on chromosome 7q22. Hum Mol Genet 2000; 9: 1049–1057.
Schwab SG, Hallmayer J, Albus M, Lerer B, Eckstein GN, Borrmann M et al. A genome-wide autosomal screen for schizophrenia susceptibility loci in 71 families with affected siblings: support for loci on chromosome 10p and 6. Mol Psychiatry 2000; 5: 638–649.
Gurling HM, Kalsi G, Brynjolfson J, Sigmundsson T, Sherrington R, Mankoo BS, et al. Genomewide genetic linkage analysis confirms the presence of susceptibility loci for schizophrenia, on chromosomes 1q32.2, 5q33.2, and 8p21–22 and provides support for linkage to schizophrenia, on chromosomes 11q23.3–24 and 20q12.1–11.23. Am J Hum Genet 2001; 68: 661–673.
Gottesman II . Schizophrenia Genesis. Freeman: New York, 1991.
Risch N . Linkage strategies for genetically complex traits. I. Multilocus models. Am J Hum Genet 1990; 46: 222–228.
Riley BP, Tahir E, Rajagopalan S, Mogudi-Carter M, Faure S, Weissenbach J et al. Psychiatr Genet 1997; 7: 57–74.
Kendler KS, O'Neill FA, Burke J, Murphy B, Duke F, Straub RE et al. Irish study on high-density schizophrenia families: field methods and power to detect linkage. Am J Med Genet 1996; 67: 179–190.
Straub RE, MacLean CJ, O'Neill FA, Burke J, Murphy B, Duke F et al. A potential vulnerability locus for schizophrenia on chromosome 6p24–22: evidence for genetic heterogeneity. Nat Genet 1995; 11: 287–293.
Baron M . Linkage results in schizophrenia. Am J Med Genet 1996; 67: 121–123.
Kendler KS, Straub RE, MacLean CJ, Walsh D . Reflections on the evidence for a vulnerability locus for schizophrenia on chromosome 6p24–22. Am J Med Genet 1996; 67: 124–126.
Schwab SG, Albus M, Hallmayer J, Honig S, Borrmann M, Lichtermann D et al. Evaluation of a susceptibility gene for schizophrenia on chromosome 6p by multipoint affected sib-pair linkage analysis. Nat Genet 1995; 11: 325–327.
Maziade M, Bissonnette L, Rouillard E, Martinez M, Turgeon M, Charron L et al. 6p24–22 region and major psychoses in the Eastern Quebec population. Le Groupe IREP. Am J Med Genet 1997; 74: 311–318.
Lindholm E, Ekholm B, Balciuniene J, Johansson G, Castensson A, Koisti M et al. Linkage analysis of a large Swedish kindred provides further support for a susceptibility locus for schizophrenia on chromosome 6p23. Am J Med Genet 1999; 88: 369–377.
Mowry BJ, Nancarrow DJ, Lennon DP, Sandkuijl LA, Crowe RR, Silverman JM et al. Schizophrenia susceptibility and chromosome 6p24–22. Nat Genet 1995; 11: 233–234.
Gurling H, Kalsi G, Chen AHS, Green M, Butler R, Read T et al. Schizophrenia susceptibility and chromosome 6p24–22. Nat Genet 1995; 11: 234–235.
Riley BP, Rajagopalan S, Mogudi-Carter M, Jenkins T, Williamson R . No evidence for linkage of chromosome 6p markers to schizophrenia in Southern African Bantu-speaking families. Psychiatr Genet 1996; 6: 41–49.
Antonarakis SE, Blouin J-L, Pulver AE, Wolniec P, Lasseter VK, Nestadt G et al. Schizophrenia susceptibility and chromosome 6p24–22. Nat Genet 1995; 11: 235–236.
Daniels JK, Spurlock G, Williams NM, Cardno AG, Jones LA, Murphy KC et al. Linkage study of chromosome 6p in sib-pairs with schizophrenia. Am J Med Genet 1997; 74: 319–323.
Levinson DF, Wildenauer DB, Schwab SG, Albus M, Hallmayer J, Lerer B et al. Additional support for schizophrenia linkage on chromosomes 6 and 8: a multicenter study. Am J Med Genet Neuropsychiatr Genet 1996; 67: 580–594.
Turecki G, Rouleau GA, Joober R, Mari J, Morgan K . Schizophrenia and chromosome 6p. Am J Med Genet Neuropsychiatr Genet 1997; 74: 195–198.
Lewis CM, Levinson DF, Wise LH, DeLisi LE, Straub RE, Hovatta I et al. Meta-analysis of schizophrenia genome scans demonstrates significant linkage findings. 2002; submitted.
Badner JA, Gershon ES . Meta-analysis of whole-genome linkage scans of bipolar disorder and schizophrenia. Mol Psychiatry 2002; 7: 405–411.
Straub RE, Jiang Y, MacLean C, Ma Y, Webb BT, Myakishev MV et al. Genetic variation in the 6p22.3 gene DTNBP1, the human ortholog of mouse dysbindin is associated with schizophrenia. Am J Hum Genet 2002; 71: 337–348.
Jiang Y, Straub RE, Sullivan PF, Harris-Kerr C, Webb BT, Wormley B et al. Genomic structure and identification of a human DTNBP1 gene from a putative schizophrenia susceptibility locus on 6p22.3. 2002; submitted.
Benson MA, Newey SE, Martin-Rendon E, Hawkes R, Blake DJ . Dysbindin, a novel coiled-coil-containing protein that interacts with the dystrobrevins in muscle and brain. J Biol Chem 2001; 276: 24232–24241.
Van den Oord EJCG, Jiang Y, Riley BP, Kendler KS, Chen X . FP-TDI SNP scoring by manual and statistical procedures: A study of error rates and types. BioTechniques 34: 610–624.
Daly MJ, Rioux JD, Schaffner SF, Hudson TJ, Lander ES . High-resolution haplotype structure in the human genome. Nat Genet 2001; 29: 229–232.
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B et al. The structure of haplotype blocks in the human genome. Science 2002.
Patil N, Berno AJ, Hinds DA, Barrett WA, Doshi JM, Hacker CR et al. Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science 2001; 294: 1719–1723.
Spitzer RL, Williams JB, Gibbon M, First MB . The structured clinical interview for DSM-III-R (SCID). I. History, rationale, and description. Arch Gen Psychiatry 1992; 49: 624–629.
Kendler KS, Lieberman JA, Walsh D . The structured interview for schizotypy (SIS): a preliminary report. Schizophr Bull 1989; 15: 559–571.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders. American Psychiatric Association: Washington DC, 1987.
Cohen J . A coefficient of agreement for nominal scales. Educ Psychol Meas 1960; 20: 37–46.
Chen X, Levine L, Kwok PY . Fluorescence polarization in homogeneous nucleic acid analysis. Genome Res 1999; 9: 492–498.
Degli-Espost M, Andreas A, Christiansen F, Schalke B, Albert E . An approach to the localization of the susceptibility genes for generalized myasthenia gravis by mapping recombinant ancestral haplotypes. Immunogenetics 1992; 35: 355–364.
Sobel E, Lange K . Descent graphs in pedigree analysis: applications to haplotyping, location scores, and marker-sharing statistics. Am J Hum Genet 1996; 58: 1323–1337.
Hodge SE, Boehnke M, Spence MA . Loss of information due to ambiguous haplotyping of SNPs. Nat Genet 1999; 21: 360–361.
Swafford D . PAUP* Phylogenetic Analysis Using Parsimony (* and other methods). Sinauer Associates: Sunderland, MA, USA, 1998.
Cliff AD, Ord JK . Spatial processes. Models and Applications. London: Pion, 1981.
Geary RC . The contiguity ratio and statistical mapping. The incorporated statistician. 1954; 5: 115–145.
Stokman FN, Sprenger CJA . GRADAP graph definition and analysis package. User's manuel (version 2.0 ed). Groningen.iec ProGamma.
Clayton D . A generalization of the transmission/disequilibrium test for uncertain-haplotype transmission. Am J Hum Genet 1999; 65: 1170–1177.
Clayton D, Jones H . Transmission/disequilibrium tests for extended marker haplotypes. Am J Hum Genet 1999; 65: 1161–1169.
Martin ER, Monks SA, Warren LL, Kaplan NL . A test for linkage and association in general pedigrees: the pedigree disequilibrium test. Am J Hum Genet 2000; 67: 146–154.
Martin ER, Bass MP, Kaplan NL . Correcting for a potential bias in the pedigree disequilibrium test. Am J Hum Genet 2001; 68: 1065–1067.
Rubinstein P, Walker M, Carpenter C, Carrier C, Krassner J, Falk C et al. Genetics of HLA disease associations: the use of the haplotype relative risk (HRR) and the “haplo=delta” (DH) estimates in juvenile diabetes from three racial groups. Hum Immunol 1981; 3: 384.
Spielman RS, McGinnis RE, Ewens WJ . Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am J Hum Genet 1993; 52: 506–516.
Sham P . Statistics in Human Genetics. Arnold: London, 1998.
Weinberg CR . Allowing for missing parents in genetic studies of case–parent triads. Am J Hum Genet 1999; 64: 1186–1193.
Horvath S, Laird NM . A discordant-sibship test for disequilibrium and linkage: no need for parental data. Am J Hum Genet 1998; 63: 1886–1897.
Lewontin RC . On measures of gametic disequilibrium. Genetics 1988; 120: 849–852.
Johnson GC, Esposito L, Barratt BJ, Smith AN, Heward J, Di Genova G et al. Haplotype tagging for the identification of common disease genes. Nat Genet 2001; 29: 233–237.
Ardlie KG, Kruglyak L, Seielstad M . Patterns of linkage disequilibrium in the human genome. Nat Rev Genet 2002; 3: 299–309.
Kitching I, Forey P, Humphries C, Williams D . Cladistics: The Theory and Practice of Parsimony Analysis. Oxford University Press: Oxford, 1998.
Ardlie K, Liu-Cordero SN, Eberle MA, Daly M, Barrett J, Winchester E et al. Lower-than-expected linkage disequilibrium between tightly linked markers in humans suggests a role for gene conversion. Am J Hum Genet 2001; 69: 582–589.
Frisse L, Hudson RR, Bartoszewicz A, Wall JD, Donfack J, Di Rienzo A . Gene conversion and different population histories may explain the contrast between polymorphism and linkage disequilibrium levels. Am J Hum Genet 2001; 69: 831–843.
Cheung KH, Osier MV, Kidd JR, Pakstis AJ, Miller PL, Kidd KK . ALFRED: an allele frequency database for diverse populations and DNA polymorphisms. Nucleic Acids Res 2000; 28: 361–363.
Lõhmussaar E . SNP analysis and linkage disequilibrium map of chromosome 22 using APEX arrays. European Hum Genet Conference, P1058 (Abstract), 2002.
Schaid DJ, Sommer SS . Genotype relative risks: methods for design and analysis of candidate–gene association studies. Am J Hum Genet 1993; 53: 1114–1126.
Van den Oord EJCG, Vermunt JK . Testing for linkage disequilibrium, maternal effects, and imprinting with (in)complete case–parent triads, by use of the computer program LEM. Am J Hum Genet 2000; 66: 335–338.
Acknowledgements
This project was largely supported by Grants MH41953, MH52537, and MH45390 from the United States National Institute of Mental Health (NIMH). Data collection was conducted under the supervision of S Humphries, M Healy, and A Finnerty. Additional interviews were conducted by J Burke, B Murphy, F Duke, R Shinkwin, M Ni Nuallain, F McMahon, J Downing, T Hebron, B Hanratty, E Crowe, M Doherty, J Bray, and L Lowry. We also wish to acknowledge Dr Richard Straub who supervised the SNP genotyping and initial DTNBP1-schizophrenia data analyses.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
van den Oord, E., Sullivan, P., Jiang, Y. et al. Identification of a high-risk haplotype for the dystrobrevin binding protein 1 (DTNBP1) gene in the Irish study of high-density schizophrenia families.. Mol Psychiatry 8, 499–510 (2003). https://doi.org/10.1038/sj.mp.4001263
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.mp.4001263
Keywords
This article is cited by
-
The DTNBP1 (dysbindin-1) gene variant rs2619522 is associated with variation of hippocampal and prefrontal grey matter volumes in humans
European Archives of Psychiatry and Clinical Neuroscience (2013)
-
The dystrobrevin-binding protein 1 gene: features and networks
Molecular Psychiatry (2009)
-
Korrelation zwischen Risikogenvarianten für Schizophrenie und Hirnstrukturanomalien
Der Nervenarzt (2009)
-
Dysbindin gene (DTNBP1) and schizophrenia in Korean population
European Archives of Psychiatry and Clinical Neuroscience (2009)
-
Genomic structural variation and schizophrenia
Current Psychiatry Reports (2008)