Abstract
The first gene associated with Crohn disease (CD) has been identified as CARD15 (16q12). Three variants, R702W, G908R and 1007fsinsC are strongly and independently associated with the disease. A second gene, conveying a smaller risk for inflammatory bowel disease (IBD), has been identified as DLG5 (10q23). We assess the frequency of the CARD15 SNPs and of the R30Q mutation in DLG5 and their contribution to the development of CD in a cohort of unrelated IBD patients (151 CD, 325 ulcerative colitis (UC)) and healthy controls (236) from South-east Norway (IBSEN cohort). Genotype-based tests of population differentiation using 23 SNPs across CARD15, together with estimates of FST, indicated that the German and Norwegian background populations could be differentiated at the CARD15 locus. The Norwegian and German CD samples exhibited particularly strong differentiation at the three predisposing loci and those marking their background haplotype. There were significantly lower frequencies of the CARD15 SNPs and no significant association with CD in the Norwegian samples. Only a marginal association was observed for the subphenotypes ileitis and ileocolitis vs colitis (P=0.048). The population attributable risk percentage (PAR%) for CARD15 variants in the Norwegian cohort is the lowest reported for a European population (1.88%), except Iceland. Similarly, the DLG5 variant showed no association with CD or IBD, however, there was a negative correlation with stricture (P=0.035). The present results are consistent with an emerging pattern of a low frequency of the CARD15 variants in Northern countries where the prevalence of IBD is greatest.
Similar content being viewed by others
Introduction
Crohn disease (CD (MIM 266600)) and ulcerative colitis (UC (MIM 191390)) are the two major forms of idiopathic inflammatory bowel diseases (IBD). The current estimated prevalence in North Western countries is 40–250/100 000 individuals, with the highest reported incidence in Scandinavia1, 2, 3, 4 and mid-western Canada. The etiology of IBD is still largely unknown but is believed to reflect the interaction of a multifactorial genetic component, confirmed by consistent evidence of familial clustering,5 an increased concordance of the IBD phenotype in monozygotic twins6, 7 and consistently positive results from genetic linkage studies, and environmental factors.8
The first gene underlying susceptibility to CD has been identified as CARD15 (Protein: NOD2) (16q12, IBD1).9, 10, 11 The NOD2 protein is involved in the interaction between monocytes and bacterial peptidoglycans.12, 13 Three SNPs located the sequence of CARD15 encoding the C-terminal leucine-rich region, two missense mutations C14772T (R702W), G25386C (G908R) and a frame-shift mutation 32629insC (L1007fs) creating a truncated protein, are strongly and independently associated with CD susceptibility in central European white populations. These three SNPs never occur in cis, that is on the same haplotype but share a common background haplotype that is partially marked by the P268S variant.14
The overall frequencies of the three mutant alleles in Caucasian CD populations (European and North American)9, 10, 11, 15, 16, 17, 18, 19 range from 19.1 to 29%; while lower frequencies have been found in African-Americans.20 None of these mutations have been found in Asian IBD populations (Japanese,21 Korean,14 Chinese22). Recently, studies on CD patients from Finland,23 Ireland,24, 25 Scotland25 and Iceland26 have indicated a low frequency of the three mutations in these populations. Considerable variation in frequencies at loci associated with complex disorders among European populations, in particular when these differences are observed in patient samples, may have important implications for the replication of association studies and the clinical impact of the loci in question.
A second IBD susceptibility gene has been recently identified as DLG5 (10q23).27 This gene encodes a scaffolding protein that is believed to be involved in the maintenance of epithelial cell integrity.28 It has been suggested that DLG5, which is a binding partner of vinexin, may play a role in cell–cell contact29 and that mutations in this gene may therefore be involved in defects of the intestinal epithelial barrier function. Such defects are known to be a feature of IBD. In contrast to CARD15, DLG5 conveys a much smaller risk with odds ratios (OR) around 1.5 in two independent studies (in comparison with about 40 for CARD15).27, 28, 29, 30 While the positional information cannot be clearly resolved on the level of coding SNPs, the variant G113A, resulting in the amino-acid substitution R30Q, has been suggested as potentially causative for both CD and IBD (CD plus UC).27
Until now little is known about the genetic background of IBD in the high incidence population of Norway. Given the heterogeneity in allele frequencies reported for CARD15 variants in different European populations, we decided to examine the frequencies of 23 well-studied SNPs in the CARD15 gene in a thoroughly characterized Norwegian, population-based incidence cohort and to contrast these with a long-established German CD cohort.11, 14, 31 We tested the ability of these SNPs to act as markers of population differentiation between the German and Norwegian disease and control (background) populations and assessed the impact and risk conferred by the disease associated SNPs C14772T (R702W), G25386C (G908R) and 32629insC (1007fsinsC) in the Norwegian population. For comparison, the frequency of DLG5 R30Q is also examined and tested for association with IBD, CD and UC. The contribution of CARD15 and DLG5 variants to the localization and behavior of CD in the Norwegian cohort of 476 unrelated IBD patients (the IBSEN cohort) and 236 healthy controls1 is briefly examined.
Patients and methods
Study population
A cohort of 476 unrelated, consecutive, newly diagnosed IBD patients (CD: n=151, 31.7%; UC: n=325, 68.3%) from South-east Norway was studied. The patients were registered prospectively from a well-defined geographical area between January 1, 1990 and December 31, 1994 and therefore a follow-up of at least 10 years was available for all patients.1, 32 All patients were seen by IBD specialists and the diagnosis was based on standard clinical, radiological, endoscopic and histological criteria.33 Unrelated healthy, age- and sex-matched controls (n=236) were recruited from the blood donor system of the same geographic region as the disease cohort. A small subgroup of CD patients (n=55) from the Norwegian cohort presented in this study has been included in a previous analysis.31
The following demographic and clinical characteristics were recorded for all the patients (Table 1): gender, age at inclusion and history of familial IBD (first-degree relative affected). For the CD patients the following characteristics were recorded, according to the Vienna Classification34: disease localization at its maximal extent (terminal ileum, ileo colon, colon, upper GI) and disease behavior (inflammatory, stricturing, penetrating). For the UC patients, disease extent was recorded (extensive colitis (disease extending beyond splenic flexure), left side colitis, procto-sigmoiditis and proctitis). All patients gave informed consent to participate in the genetic study.
The German comparison cohort consisted of 462 unrelated IBD patients (CD: n=309, 66%; UC: n=153, 33%), recruited through the German Crohn's and Colitis Foundation and the Competence Network ‘Inflammatory Bowel Disease’ (coordinated at Christian-Albrechts University, Kiel, Germany), plus 540 German unrelated, matched healthy controls, recruited through the Department of Transfusion Medicine at Kiel University. The German samples used in the present paper form a subset of those previously published in association with DLG5 and CARD15.11, 14, 27, 31
Sequencing and genotyping
DNA was extracted using standard techniques (guanidine-detergent lysis) from EDTA blood and dispensed into 96-well plates (20 ng/well). The coding exons (2–12) of the CARD15 gene were screened for novel mutations by direct genomic sequencing in 90 Norwegian CD patients as previously described.14 All CARD15 SNPs (Table 2) and DLG5 R30Q (rs1248696) were genotyped using TaqMan technology on an ABI 7700 Sequence Detector (Applied Biosystems, Foster City, CA, USA) as previously described.14, 27
Statistical analyses
Each marker was tested to ensure Hardy–Weinberg equilibrium in the control populations using a χ2-test. Each of the 23 markers were then tested for their ability to distinguish between the German and Norwegian controls and between the German and Norwegian cases using 10 000 genotype permutations as implemented by FSTAT.35 Values of FST were also calculated using FSTAT. Summary, tests of population differentiation and FST were also carried out, similarly, using all 23 markers as complete multilocus genotypes. This approach does not account for the appreciable level of linkage disequilibrium (LD) between these markers. For comparison, we therefore also explicitly accounted for LD by using HaploRec36 to infer the multilocus haplotypes for all 23 markers in each individual (ie an individuals diplotype) and then repeated the calculation of population differentiation and FST treating the multilocus haplotypes as a single multiallelic locus.
A case–control analysis for the four nonsynonymous mutations (P268S, R702W, G908R and 1007insC) was performed against unrelated healthy controls. Association was tested for genotypes at the appropriate number of degrees of freedom using either a χ2-test or Fisher's exact test as appropriate. OR were calculated by pooling carriers of the rare (predisposing) allele into a single genotype class and confidence intervals (CI) calculated using SISA.37 The population attributable risk percentage (PAR%) was calculated as the attributable risk percentage (AR%) multiplied by the proportion of exposed cases, where AR% was estimated from the OR, assuming that the exposure of the control population to the disease-associated variant reflects the true prevalence of the variant in the general population.38
Results
Mutation detection in CARD15
The coding exons (2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12) and intron–exon boundaries were sequenced in 90 Norwegian CD patients. We confirmed the presence of the following SNPs: SNP5, SNP6, SNP7, SNP8, SNP9, SNP12 and SNP13 (all named according to Hugot et al9); rs2067085 (intron 8), A+3520C and G+4279A (exon 12, 3′UTR)14; C+1833T (A611A) and C+2264T (A755V) (exon 4) and G+2863A (V955I) (exon 9).17 The other rare mutations identified by Lesage et al17 were not detected. SNP numbering is from to the ATG.
Frequencies of the CARD15 SNPs in the two populations
The 23 SNPs in the CARD15 gene that were genotyped are detailed in Table 2. For population comparisons these SNPs were genotyped in 79 CD patients and the 236 healthy controls from Norway. Only the four nonsynonymous markers C13470T (P268S), C14772T (R702W), G25386C (G908R), 32629insC (L1007fs) plus G-21889A were then genotyped in the remainder of the Norwegian IBD cohort (n=476). All German samples had been previously genotyped for these 23 markers. Table 3 shows the population comparison between the Norwegian and German samples. When comparing the two control populations, which are taken to be representative of the general population, most markers exhibit low values of FST (FST<0.01) such as would be expected when comparing two similar European populations. Three markers, G-21889A, G162674A and notably 32629insC (1007fsinsC) have FST values greater than 0.01 and correspondingly significant P-values in the genotype-based test of population differentiation. Many other markers show borderline significance in their ability to distinguish the two populations (although only G162674A remains significant after correction for multiple testing (Dunn–Šidák Pcritical=0.002, 23 tests)). With respect to the three markers known to be associated with CD, the C14772T (R702W) T allele occurs with about half the frequency in the Norwegian control population compared with the German population (0.024 vs 0.045, Pallele=0.059), the G25386C (G908R) C allele is slightly more common in the Norwegian sample but not significantly so (0.009 vs 0.006, Pallele=0.504), and the 32629insC (L1007fs) C insertion occurs at about one-third as frequently in the Norwegians (0.015 vs 0.043, Pallele=0.006). When all 23 markers are considered then these CARD15 SNPs are clearly able to distinguish between the Norwegian and German control samples (P‘multi-locus genotype’=0.001, P‘single-locus haplotype’<0.001; Table 3).
When the Norwegian CD sample and the German CD sample are compared (Table 3) a number of markers exhibit a dramatic increase in FST and highly significant tests of population differentiation at the genotype level. All of these markers have FST values well in excess of 0.01 and either correspond to the known CD predisposing mutations or broadly tag the background haplotype as defined in Croucher et al.14 Close inspection of the allele frequencies in Table 3 indicates that these differences result from the dramatic increase in the rare allele frequencies of these SNPs (ie of the disease-associated haplotypes) in the German cases, whereas only minor differences in allele frequencies are observed between the Norwegian cases and controls. Indeed, the C14772T (R702W) and G25386C (G908R) polymorphisms have essentially identical allele frequencies in the Norwegian controls and cases. Consequently, the CD-associated markers all occur at substantially lower frequencies in the Norwegian CD cases compared with the German CD cases (C14772T (R702W) T allele: 0.024 vs 0.095, Pallele <0.001; G25386C (G908R) C allele: 0.010 vs 0.049, Pallele=0.005; 32629insC (L1007fs) C insertion: 0.030 vs 0.162, Pallele≪0.001). When all 23 markers are considered they are clearly able to distinguish between the Norwegian and German CD samples (P‘multi-locus genotype’=0.002, P‘single-locus haplotype’<0.001; Table 3). For the majority of the CARD15 SNPs only 79 individuals were genotyped, if the sample size had been larger then it is likely that an even greater differentiation would be detected between the Norwegian and German CD populations.
CARD15 and disease susceptibility
The four nonsynonymous CARD15 SNPs were examined for association with susceptibility to CD in the Norwegian cohort using a case–control approach. Table 4 shows the P-values for the genotype frequency comparisons and gives the OR and CI for carriership of the rare (predisposing) allele at each of these markers, for both the Norwegian cohort and the German comparison cohort. While all four of these SNPs show significant association with CD in the German sample (as previously reported eg, Hampe et al,11, 31 and Croucher et al14 none were found to be significantly associated with susceptibility to CD in the Norwegian sample (furthermore, there was no apparent association with UC or the combined category IBD – data not shown)). Simple heterozygotes of the three CD implicated variants C14772T (R702W), G25386C (G908R) and 32629insC (L1007fs) were not observed more commonly in the Norwegian CD sample compared with the control group (7.69 vs 8.33%, P=0.887, OR=0.94 (CI 0.43–2.04)). Compound heterozygotes/homozygotes were more frequent in the CD sample, however, the counts were small and did not quite reach significance levels (2.80% (4) vs 0.44% (1), P=0.076, OR=6.50 (CI 0.72–58.80). Overall, carriership of any of the three variants was not significantly different in the CD cases compared to the controls (P=0.582, OR=1.22 (CI 0.60–2.47)) and the PAR% was only 1.88%. Resequencing of the coding exons of the CARD15 gene in 90 Norwegian CD patients did not identify any novel coding mutations in this population.
Disease behavior and localization
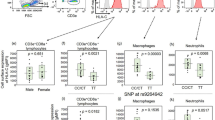
Given the rarity of the R702W, G908R and 1007fs variants and the relatively small sample size of the Norwegian cohort a systematic examination of disease behavior and localization was not feasible. However, a cursory examination of the genotype–phenotype associations that have been previously documented in European Caucasian populations15, 16, 17, 31 is informative as to the role of these rare variants in the Norwegian CD population. Disease localization was examined by comparing patients with ileal involvement (ileitis and ileocolitis) against those with colitis (and no ileal involvement). Patients with ileal disease carried a higher proportion of these CARD15 variants than did patients with colonic disease (14.94% (nine single heterozygotes, four compound heterozygotes/homozygotes) vs 3.70% (two single heterozygotes), P=0.048, OR=4.57 (CI 0.99–21.10)). Disease behavior was classified as stricturing (stenosing), penetrating (fistulizing) or inflammatory (nonstricturing or penetrating). Compared to the simple inflammatory category, patients with penetrating and those with stricturing disease carried a higher proportion of CARD15 variants (13.04 and 20.69 vs 6.59%). However, only stricturing disease achieved nominal significance (stricturing: P=0.028, OR=3.70 (CI 1.09–12.54); penetrating: P=0.383, OR=2.13 (CI 0.49–9.23)).
DLG5 R30Q frequency and disease susceptibility
The IBD-associated DLG5 G113A (R30Q) A-allele occurred with similar frequency in the Norwegian control population to that reported for the German control population by Stoll et al27 (0.109 vs 0.090, Pallele=0.264). The allele frequency in the Norwegian IBD cases did not differ significantly from that observed in the Norwegian controls (0.092 vs 0.109, Pallele=0.338). Consequently, the A-allele was under-represented in the Norwegian IBD cases when compared to the German IBD cases (0.092 vs 0.132, Pallele=0.008).27 Table 5 shows the case–control association statistics for the Norwegian and German samples at the DLG5 G113A (R30Q) locus and includes gender-stratified analyses. The German cohort, which is a subset of that presented by Stoll et al,27 exhibits the expected weak association with IBD (OR=1.48 (1.08–2.04), CD (OR=1.46 (1.03–2.09)) and UC (OR=1.60 (1.03–2.49)). The gender-stratified data suggests that at least the CD component of this association may be male specific (CD, OR=2.28 (1.20–4.35)). The Norwegian data showed no noteworthy evidence of association with IBD, CD or UC at this SNP. However, it must be noted that this cohort is underpowered to demonstrate such a weak to moderate disease association. Further, the 95% CI of, for example, CD (0.60–1.67) overlap with the OR estimate for CD in the German population (OR=1.46), therefore an association with DLG5 R30Q cannot be ruled out in the Norwegian population.
DLG5 R30Q and disease behavior and localization
Despite the low frequency and the apparent lack of association of this variant, the potential influence of DLG5 R30Q on disease behavior and localization in CD patients was examined in the same manner as was carried out for the CARD15 variants. CD patients with a colonic disease localization included a higher frequency of heterozygous carriers of the A-allele compared to those with an ileal presentation, but this difference was not significant (26.00 vs 17.65%, P=0.248, OR=1.64 (CI 0.71–3.81)). Compared to the simple inflammatory category, patients with stricturing disease exhibited a slightly higher carriership of the DLG5 R30Q A-allele but this difference was not significant (30.00 vs 25.29%, P=0.665, OR=1.27 (CI 0.43–3.70)). However, compared to the inflammatory category, stricturing disease showed a negative correlation with carriership of the A-allele (6.45 vs 25.29%, P=0.035, OR=0.20 (CI 0.04–0.92)).
The original publication describing the association of DLG5 with IBD27 suggested a potential interaction between the two genes because a significantly greater transmission of the DLG5 R30Q A-allele was observed in CD trios carrying the CARD15 risk variants compared with those not carrying the CARD15 risk variants. Among the Norwegian CD patients carrying a CARD15 mutation, 20% (three out of 15) also carried the DLG5 R30Q A-allele, compared with 15.79% (three out of 19) of controls carrying a CARD15 mutation. Similar values were observed in individuals not carrying a CARD15 mutation with the carriership of the DLG5 R30Q A-allele being 22.61% in CD patients and 21.50% in controls. Sample sizes are too small to establish whether the differences observed in patients carrying a CARD15 mutation is meaningful (P>0.05).
Discussion
Following the identification of CARD15 as the first gene conferring susceptibility to CD in 20019, 10, 11 many studies have confirmed the association of the C14772T (R702W), G25386C (G908R) and 32629insC (1007fsinsC) variants with the development of CD and their contribution to ileal localization of the disease in populations from Europe and North America (Caucasian origin) (eg Hugot et al,9 Ogura et al,10 Vermeire et al,15 Cuthbert et al,16 Lesage et al,17 Abreu et al,18 Ahmad et al19 and Hampe et al31). Differences in the frequencies of these variants between Ashkenazi Jewish20, 39 and non-Jewish populations and their absence in Japanese,21 Korean14 and Chinese22 IBD patients points to a high degree of heterogeneity between ethnically divergent populations. This pattern is consistent with the rarity and therefore probably recent ancestry of these mutations.14, 40
However, significant heterogeneity in the frequencies of these variants has also been observed within Europe. A North-South gradient in allele frequencies is observed in CD patients, with northern European populations exhibiting the lowest frequencies (eg for the 1007fs variant: Finnish 4.8%,23 Icelandic 0%,26 Scottish 4.6% and Irish 3.0%25) and Southern European populations exhibiting the highest frequencies (eg for the 1007fs variant: Italian 9%41 and Spanish 14.2%42). Two aspects are striking about this observation. First, the opposite gradient has been well documented for the prevalence of IBD, with the Scandinavian countries having the highest incidence of IBD.4 Second, although there is some variation in the frequencies of these variants in the respective control populations these differences are largely confined to the CD samples and would therefore appear to represent real differences in the population attributable risk of the CARD15 variants. For a concise summary of the frequencies of these variants in the literature see Arnott et al25 and Economou et al.43
The results presented here add to this pattern. The frequencies of the three CD-associated CARD15 variants were examined in a Norwegian cohort, consisting of 236 healthy controls and 476 sporadic IBD patients (151 patients with CD). The Norwegian cohort was contrasted with a well-studied German cohort consisting of 540 healthy controls and 309 sporadic CD cases. An additional 20 SNPs in the CARD15 gene were also examined. Some differences in allele frequencies were evident when comparing the German and Norwegian control populations and taken together these 23 markers were able to differentiate the German and Norwegian populations in genotype-based tests of population differentiation (P‘multi-locus genotype’=0.0013, P‘single-locus haplotype’<0.001; Table 3).
However, major differences were then observed between the Norwegian and German patients with CD. Compared with the controls, the German cases exhibited significantly higher frequencies of the rare alleles at the three CD-associated CARD15 variants and also in those marking the background haplotype that carries these variants – in agreement with the known CD association in this cohort. With the exception of 32629insC (L1007fs) (which doubled in frequency from 1.5% in the controls to 3.0% in the cases) there was little change in the allele frequencies between the Norwegian controls and cases (Table 3) and no association with CD was evident (Table 4). The PAR% for carriership of the CARD15-CD predisposing alleles was only 1.88%. This value is the lowest so far reported for a European population (with the exception of the Icelandic population which has none of these variants26) and indicates a very minor contribution of CARD15 mutations to CD in the Norwegian population. The lowest PAR% previously reported was 11.0% for the Scottish population.25 The studies of Hugot et al reflected a PAR% of 33.2%.9, 25
The increased frequency of the G25386C (G908R) variant in UC patients compared with healthy controls, observed in the Scottish population described by Arnott et al,25 was not seen in the present cohort.
An examination of an SNP in another gene that has been associated with IBD, DLG5 G113A (R30Q), surprisingly exhibited exactly the same pattern as CARD15 with no association evident in the Norwegian cohort and essentially no associated risk. This SNP tags a haplotype that was found to be significantly overtransmitted in TDT analyses performed on German multiplex families and trios. This association was corroborated in an independent case–control analysis of 525 German CD cases and 515 healthy controls.27 Therefore, the lack of contribution of CARD15 to the genetic risk for CD in Norway is unlikely to make up by this other variant. Interestingly, the Scottish population also shows no association with this variant44 yet exhibits similar allele frequencies to the Norwegian cases and control and the German controls (11.4% in IBD and 13.2% in healthy controls). However, it appears that age and sex distribution could be major confounders obscuring a disease-specific effect.46, 47, 48, 49 In addition, a stronger association with pediatric IBD has been suggested with association signals for DLG5 R30Q being detected in the Scottish population too.49 Therefore, use of tightly age- and sex-matched samples appears of importance in addition to a putative heterogeneity between populations.50 It will be interesting to see if a similar gradient in allele frequencies in Europe is seen for DLG5 as is seen for CARD15.
It may be argued that the Norwegian cohort analyzed here was relatively small and therefore underpowered to detect an association at the CARD15 and DLG5 loci. This is indeed true, the power to detect an association between the CARD15 32629insC (L1007fs) variant and CD in this sample (at a relative risk of 1.8 (carriership) and a risk factor frequency of 22%) is only about 70% and far less for the other variants. Also, it should be noted that the 95% CI of the OR calculated for the Norwegian CARD15 variants are broad and that those for R702W and 1007fs overlap the OR estimates for the German population. CARD15 may play a significant role in Norwegian CD susceptibility, however, in the present study sample both the CARD15 and the DLG5 variants appear to be of relatively minor impact. Furthermore, the Norwegian CD population did not differ substantially from the German CD population in terms of its clinical composition in the most relevant traits; although a strict comparison is difficult. Both cohorts exhibited similar proportions of patients with stricturing or fistulating complications (Norway: 78.8%; Germany: 80.6%) and ilieal involvement (Norway: 59.6%; Germany: 74.4%). It seems unlikely that any such differences would be sufficient to generate that dramatic differences observed here.
Consequently, although there can be little doubt that the CARD15 variants C14772T (R702W), G25386C (G908R) and 32629insC (1007fsinsC), and also variants in the DLG5 gene, represent major risk factors for the development of IBD (in particular CD), this risk appears to be highly population specific. That the risk associated with particular genetic loci varies dramatically between ethnically distinct groups (eg the fact that CARD15 is a risk factor in Caucasians but not Asians) is perhaps not surprising given the complex genetic etiology of a common disease such as IBD. That the risk varies so dramatically between ethnically similar populations (ie within Europeans) is of major importance. First, much of the confidence that is assigned to studies of genetic association in complex disorders is determined by the gold standard of replication in independent populations. Our study, albeit of modest sample size, fails to replicate the association between IBD (CD) and either CARD15 or DLG5. However, a rudimentary analysis of disease localization and behavior does indicate the expected associations with the variants in these genes (ileal disease and stricture43 for CARD15 and general inflammation/lack of stricture for DLG5). Although the PAR% for the CARD15 variants is high (around 30%) in many populations studied, the low PAR% for CD in northern populations, where the prevalence of the disease is highest, brings into question the direct clinical relevance of these mutations. Marsh and McLeod45 have suggested that the frequencies of the CARD15 variants that they observed in healthy populations of Caucasian, African and Asian descent, with the variants being rarer in Africans than Caucasians and absent in Asians, might to some extent contribute to the different incidences of CD in these populations. The European picture suggests that this may not be the case since CARD15-associated risk in Europe is broadly negatively correlated with the incidence of CD. Other loci and environmental factors might have a greater influence on northern European populations. Therefore, until the demographics and biology of CARD15 and other predisposing loci and their interactions in the etiology of IBD are better understood we should approach the clinical interpretation of these variants cautiously.
References
Moum B, Vatn MH, Ekbom A et al: Incidence of Crohn's disease in four counties in Southeastern Norway, 1990–1993. A prospective population-based study. The Inflammatory Bowel South-Eastern Norway (IBSEN) Study Group of Gastroenterologists. Scand J Gastroenterol 1996; 31: 355–361.
Moum B, Vatn MH, Ekbom A et al: Incidence of ulcerative colitis and indeterminate colitis in four counties of Southeastern Norway, 1990–1993. A prospective population-based study. The Inflammatory Bowel South-Eastern Norway (IBSEN) Study Group of Gastroenterologists. Scand J Gastroenterol 1996; 31: 362–366.
Ekbom A, Helmick C, Zack M et al: The epidemiology of inflammatory bowel disease: a large, population-based study in Sweden. Gastroenterology 1991; 100: 350–358.
Shivananda S, Lennard-Jones J, Logan R et al: Incidence of inflammatory bowel disease across Europe: is there a difference between North and South? Results of the European Collaborative Study on Inflammatory Bowel Disease (EC-IBD). Gut 1996; 39: 690–697.
Orholm M, Munkholm P, Langholz E et al: Familial occurrence of inflammatory bowel disease. N Engl J Med 1991; 324: 84–88.
Thompson NP, Driscoll R, Pounder RE et al: Genetics versus environment in inflammatory bowel disease: results of a British twin study. BMJ 1996; 312: 95–96.
Tysk C, Lindberg E, Jarnerot G et al: Ulcerative colitis and Crohn's disease in an unselected population of monozygotic and dizygotic twins. A study of heritability and the influence of smoking. Gut 1988; 29: 990–996.
Satsangi J, Morecroft J, Shah NB et al: Genetics of inflammatory bowel disease: scientific and clinical implications. Best Pract Res Clin Gastroenterol 2003; 17: 3–18.
Hugot JP, Chamaillard M, Zouali H et al: Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. Nature 2001; 411: 599–603.
Ogura Y, Bonen DK, Inohara N et al: A frameshift mutation in NOD2 associated with susceptibility to Crohn's disease. Nature 2001; 411: 603–606.
Hampe J, Cuthbert A, Croucher PJ et al: Association between insertion mutation in NOD2 gene and Crohn's disease in German and British populations. Lancet 2001; 357: 1925–1928.
Ogura Y, Inohara N, Benito A et al: Nod2, a Nod1/Apaf-1 family member that is restricted to monocytes and activates NF-kappaB. J Biol Chem 2001; 276: 4812–4818.
Inohara N, Ogura Y, Fontalba A et al: Host recognition of bacterial muramyl dipeptide mediated through NOD2 implications for Crohn's disease. J Biol Chem 2003; 278: 5509–5512 [Epub 2003 Jan 5504].
Croucher PJ, Mascheretti S, Hampe J et al: Haplotype structure and association to Crohn's disease of CARD15 mutations in two ethnically divergent populations. Eur J Hum Genet 2003; 11: 6–16.
Vermeire S, Wild G, Kocher K et al: CARD15 genetic variation in a Quebec population: prevalence, genotype–phenotype relationship, and haplotype structure. Am J Hum Genet 2002; 71: 74–83 [Epub 2002 May 2017].
Cuthbert AP, Fisher SA, Mirza MM et al: The contribution of NOD2 gene mutations to the risk and site of disease in inflammatory bowel disease. Gastroenterology 2002; 122: 867–874.
Lesage S, Zouali H, Cezard JP et al: CARD15/NOD2 mutational analysis and genotype–phenotype correlation in 612 patients with inflammatory bowel disease. Am J Hum Genet 2002; 70: 845–857.
Abreu MT, Taylor KD, Lin YC et al: Mutations in NOD2 are associated with fibrostenosing disease in patients with Crohn's disease. Gastroenterology 2002; 123: 679–688.
Ahmad T, Armuzzi A, Bunce M et al: The molecular classification of the clinical manifestations of Crohn's disease. Gastroenterology 2002; 122: 854–866.
Bonen DK, Ogura Y, Nicolae DL et al: Crohn's disease-associated NOD2 variants share a signaling defect in response to lipopolysaccharide and peptidoglycan. Gastroenterology 2003; 124: 140–146.
Yamazaki K, Takazoe M, Tanaka T et al: Absence of mutation in the NOD2/CARD15 gene among 483 Japanese patients with Crohn's disease. J Hum Genet 2002; 47: 469–472.
Leong RW, Armuzzi A, Ahmad T et al: NOD2/CARD15 gene polymorphisms and Crohn's disease in the Chinese population. Aliment Pharmacol Ther 2003; 17: 1465–1470.
Helio T, Halme L, Lappalainen M et al: CARD15/NOD2 gene variants are associated with familially occurring and complicated forms of Crohn's disease. Gut 2003; 52: 558–562.
Bairead E, Harmon DL, Curtis AM et al: Association of NOD2 with Crohn's disease in a homogenous Irish population. Eur J Hum Genet 2003; 11: 237–244.
Arnott ID, Nimmo ER, Drummond HE et al: NOD2/CARD15, TLR4 and CD14 mutations in Scottish and Irish Crohn's disease patients: evidence for genetic heterogeneity within Europe? Genes Immun 2004; 5: 417–425.
Thjodleifsson B, Sigthorsson G, Cariglia N et al: Subclinical intestinal inflammation: an inherited abnormality in Crohn's disease relatives? Gastroenterology 2003; 124: 1728–1737.
Stoll M, Corneliussen B, Costello CM et al: Genetic variation in DLG5 is associated with inflammatory bowel disease. Nat Genet 2004; 36: 476–480 [Epub 2004 Apr 2011.
Nakamura H, Sudo T, Tsuiki H et al: Identification of a novel human homolog of the Drosophilla dlg, P-dlg, specifically expressed in the gland tissues and interacting with p55. FEBS Lett 1998; 433: 63–67.
Wakabayashi M, Ito T, Mitsushima M et al: Interaction of lp-dlg/KIAA0583, a membrane-associated guanylate kinase family protein, with vinexin and beta-catenin at sites of cell-cell contact. J Biol Chem 2003; 278: 21709–21714 [Epub 22003 Mar 21725].
Daly MJ, Pearce AV, Farwell L et al: Association of DLG5 R30Q variant with inflammatory bowel disease. Eur J Hum Genet 2005; 20: 20.
Hampe J, Grebe J, Nikolaus S et al: Association of NOD2 (CARD 15) genotype with clinical course of Crohn's disease: a cohort study. Lancet 2002; 359: 1661–1665.
Moum B, Ekbom A, Vatn MH et al: Inflammatory bowel disease: re-evaluation of the diagnosis in a prospective population based study in South Eastern Norway. Gut 1997; 40: 328–332.
Podolsky DK : Inflammatory bowel disease. N Engl J Med 1991; 325: 928–937.
Gasche C, Scholmerich J, Brynskov J et al: A simple classification of Crohn's disease: report of the Working Party for the World Congresses of Gastroenterology, Vienna 1998. Inflamm Bowel Dis 2000; 6: 8–15.
Goudet J : FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available at http://www.unil.ch/izea/softwares/fstat.html. Updated from Goudet (1995). 2001.
Eronen L, Geerts F, Toivonen H : A Markov chain approach to reconstruction of long haplotypes. Proceedings of the Ninth Symposium Pacific on Biocomputing (PSB). Singapore: World Scientific, 2004; 104–115.
Uitenbroek DG : SISA Binomial. World Wide Web http://home.clara.net/sisa/bionomial.htm, 1997.
Hennekens CH, Buring SL : Epidemiology in Medicine. Lippioncott Williams and Wilkins: Philadelphia, PA, 1987, pp 87–93.
Sugimura K, Taylor KD, Lin YC et al: A novel NOD2/CARD15 haplotype conferring risk for Crohn disease in Ashkenazi Jews. Am J Hum Genet 2003; 72: 509–518 [Epub 2003 Feb 2007].
Cho JH : The Nod2 gene in Crohn's disease: implications for future research into the genetics and immunology of Crohn's disease. Inflamm Bowel Dis 2001; 7: 271–275.
Annese V, Palmieri O, Latiano A et al: Frequency of NOD2/CARD15 variants in both sporadic and familial cases of Crohn's disease across Italy An Italian Group for Inflammatory Bowel Disease Study. Dig Liver Dis 2004; 36: 121–124.
Mendoza JL, Murillo LS, Fernandez L et al: Prevalence of mutations of the NOD2/CARD15 gene and relation to phenotype in Spanish patients with Crohn disease. Scand J Gastroenterol 2003; 38: 1235–1240.
Economou M, Trikalinos TA, Loizou KT, Tsianos EV, Ioannidis JPA : Differential effects of NOD2 variants on Crohn's disease risk and phenotype in diverse populations: a meta analysis. Am J Gastroenterol 2004; 99: 2393–2404.
Noble CL, Nimmo ER, Drummond H, Smith L, Arnott ID, Satsangi J : DLG5 variants do not influence susceptibility to inflammatory bowel disease in the Scottish population. Gut 2005; 54: 1416–1420.
Marsh S, McLeod HL : Crohn's disease: ethnic variation in CARD15 genotypes. Gut 2003; 52: 770.
Daly MJ, Pearce AV, Farwell L et al.: Association of DLG5 R30Q variant with inflammatory bowel disease. Eur J Hum Genet 2005; 13: 835–839.
Russell RK, Drummond HE, Nimmo ER et al.: The DLG 5-113A mutation is associated with susceptibility to early onset inflammatory bowel disease and demonstrates a complex genotype phenotype relationship: PG2-05. J Pediatr Gastroenterol Nutr 2005; 40: 641–642.
Tenesa A, Noble C, Satsangi J et al: Association of DLG5 and inflammatory bowel disease across populations. Eur J Hum Genet 2006; 14: 259–260.
Friedrichs F, Brescianini S, Annese V et al.: Evidence of transmission ratio distortion of DLG5 R30Q variant in general and implication of an association with Crohn disease in men. Hum Genet 2006; 31: 1–7, [Epub ahead of print].
Trinh TT, Rioux JD : Understanding association and causality in the genetic studies of inflammatory bowel disease. Gastroenterology 2005; 129: 2106–2110.
Acknowledgements
We particularly wish to thank our technicians, the cooperating clinical centers, physicians, families and patients with IBD that have contributed to the German and Norwegian IBSEN collections. We also thank the German Crohn's and Colitis Foundation (DCCV e.v.) for their continued support and participation. Grant support: Deutsche Forschungsgemeinschaft (For 423), a competence network ‘Chronisch-entzündliche Darmerkrankungen’ and the German National Genome Research Network.
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
About this article
Cite this article
Medici, V., Mascheretti, S., Croucher, P. et al. Extreme heterogeneity in CARD15 and DLG5 Crohn disease-associated polymorphisms between German and Norwegian populations. Eur J Hum Genet 14, 459–468 (2006). https://doi.org/10.1038/sj.ejhg.5201576
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201576
Keywords
This article is cited by
-
Meta-analysis of associations between DLG5 R30Q and P1371Q polymorphisms and susceptibility to inflammatory bowel disease
Scientific Reports (2016)
-
Assessment of heterogeneity between European Populations: a Baltic and Danish replication case-control study of SNPs from a recent European ulcerative colitis genome wide association study
BMC Medical Genetics (2011)
-
The multidrug resistance 1 (MDR1) gene polymorphism G-rs3789243-A is not associated with disease susceptibility in Norwegian patients with colorectal adenoma and colorectal cancer; a case control study
BMC Medical Genetics (2009)
-
Detailed assessment of NOD2/CARD15 exonic variation in inflammatory bowel disease in Scotland: implications for disease pathogenesis
Genes & Immunity (2008)
-
Analysis of the NOD2/CARD15 Gene in Patients Affected with the Aseptic Abscesses Syndrome with or without Inflammatory Bowel Disease
Digestive Diseases and Sciences (2008)