Abstract
Flatfish undergo a remarkable metamorphosis from symmetrical pelagic larvae to fully asymmetrical benthic juveniles. The most distinctive features of this transformation is the migration of one eye. The molecular role of thyroid hormone in the metamorphosis process in flatfishes is well established. However, the regulatory network that facilitates eye movement remains enigmatic. This paper presents a morphological investigation of the metamorphic process in turbot eyes, using advanced imaging techniques and a global view of gene expression. The study covers migrant and non-migrant eyes and aims to identify the genes that are active during ocular migration. Our transcriptomic analysis shows a significant up-regulation of immune-related genes. The analysis of eye-specific genes reveals distinct patterns during the metamorphic process. Myosin is highlighted in the non-migrant eye, while ependymin is highlighted in the migrant eye, possibly involved in optic nerve regeneration. Furthermore, a potential association between the alx3 gene and cranial restructuring has been identified. Additionally, it confirmed simultaneous adaptation to low light in both eyes, as described by changes in opsins expression during the metamorphic process. The study also revealed that ocular migration activates systems asynchronously in both eyes, providing insight into multifaceted reorganization processes during metamorphosis of flatfish.
Similar content being viewed by others
Introduction
The visual system is particularly important in predatory fish for their feeding and hunting behavior. This has forced animals, such as flatfish, to adapt their visual system to environmental conditions. Flatfish, an important group of fish with a high economic value, undergoes one of the most epic metamorphoses in the animal kingdom involving morphological and behavioral changes: a symmetric pelagic larva becomes an asymmetric benthic juvenile, including the migration of one eye to the other side of the body. How these fish have adapted to their new life conditions has been an enigma that has concerned scientists from Darwin to the present day1. It is believed that this unusual morphology and eye migration in flatfish has evolved as a response to the need for ambushing or seeking refuge2,3.
The transition from pelagic to benthic life requires adaptation to environmental photic variations. Flatfish undergo a complex reorganization of the retina’s photoreceptors during metamorphosis to adapt to a demersal lifestyle4,5,6. As both eyes experience attenuation of light conditions, no differences related to development are found in the ocular layers or lens7. However, several asymmetric characteristics related to eye position and visual system have been described in the literature. In order to accommodate the eye on the ocular side, a different orbit composition has been described in migrant and non-migrant eyes, such as a shorter extraocular muscle in the migrant eye8. Furthermore, there is an asymmetric craniofacial remodeling during lateralization, excluding the jaws, which remain mainly symmetrical9,10,11. Regarding differential features between eyes directly related to the visual system, a clear asymmetry in the length of the optic nerve has been found, with the non-migrant eye having a shorter nerve12, as well as a tendency for the optic nerve of the migrant eye to cross over the non-migrant eye at the optic chiasm13. Brinon et al.14 suggested a lower functionality of the migrant eye, such as a smaller left optic tectum size during metamorphosis. However, there appear to be no differences in hunting success during settlement.
Understanding the mechanisms driving eye migration has been a challenge that is still far from being fully unraveled. It is well-studied that thyroid hormone, as in amphibians15, is responsible for triggering metamorphosis and, therefore, eye migration16. Thyroid hormone inhibitors result in incomplete migration and symmetrical juveniles17. Different hypotheses have tried to explain eye migration in flatfish: (i) cranial asymmetry drives eye movement18; (ii) cell proliferation in the suborbital tissue on the blind side pushes the eye to the other side of the body19. Despite these great contributions, there are not enough studies that shed light on what occurs in the eye at a molecular level. We propose a transcriptome-wide study of the left and right eye, including surrounding tissue, at three key developmental stages (pre-metamorphic, climax of metamorphosis, and post-metamorphic) in turbot (Scophthalmus maximus) with the aim of revealing the differentiation of gene expression between an eye undergoing migration (right eye in turbot) and the eye remaining in its position after settlement (left eye in turbot).
Methods
LSFM rapid 3D imaging of skull bone development during turbot metamorphosis
A total of 50 fish reared under standard production conditions at 18 °C, were collected following a temporal sequence (10 individuals per stage): 10, 15, 20, 30, 57 dpf (days post fertilization). Turbot fish were euthanized using MS-222 (500 mg/L, for 20–30 min) (Sigma-Aldrich, USA), followed by fixation in 2% paraformaldehyde. To facilitate bone observation, fish were stained with Alizarin Red protocol specifically modified for turbot larvae and juvenile20. The stained animals were then preserved and observed in glycerol. To observe in detail the bone structure of the pre-selected turbot stages, samples were analyzed using a Zeiss Lightsheet7 imaging system (LSFM). The images were captured using a 5× NA 0.16 plan-neofluid detection objective and a zoom of 0.5 to achieve a larger field of view. The illumination laser was set to 561 nm, combined with an SBS LP 640 detection filter to capture the alizarin red signal. The Pivot Scan system was employed to eliminate any shadowed areas that might appear in the field. The images were acquired at a resolution of 1.88 × 1.88 × 4.73 mm. For each specimen, a suitable number of tiles were selected to cover the entire sample at lateral position (between 1 × 1 and 4 × 4). In the z-axis, all planes with signals were recorded. After the acquisition, tile stitching was performed using the Zen Blue software from Zeiss. Due to the large size of the samples, images were taken from two angles (0 and 180°), which were then fused using the VolumeFusion tool in the Arivis 4D software (Zeiss, Germany). Subsequently, 3D visualizations of each specimen were created using the Arivis 4D software. Structures of interest, such as the skull, bone, pseudomesial bar, and mandible, were selected using the wand tool to enhance the asymmetry of the individuals throughout the development stages. Finally, different 3D views (lateral, dorsal, and frontal) were generated for each individual.
RNAseq sample collection
Three key stages in turbot development were defined based on Al-Maghazachi and Gilson21 and Suarez-Bregua’s22 morphological criteria. Fish reared under standard production conditions at 18 °C, were collected from three metamorphic stages: pre-metamorphic (15 dpf, symmetrical larvae); the climax of metamorphosis (30 dpf, the migratory eye is visible from the other side of the body); post-metamorphic stage (57 dpf, both eyes lodged in the ocular side). Three independent replicates per stage were collected, and fish were euthanized with a lethal dose of MS-222 (500 mg/L for 20–30 min) (Sigma-Aldrich, USA).
RNA isolation and sequencing
The migrant and non-migrant eyes and their respective surrounding tissue were extracted separately for each stage. In order to obtain enough RNA quantity for sequencing, three eyes per replicate were pooled at the pre-metamorphic stage. Eyes were then fixed in RNAlater (Thermo Fisher Scientific, Waltham, MA, USA) for 24 h at 4 °C and stored at −80 °C until use. Total RNA was extracted from the tissues and purified using the RNeasy Mini Kit (Qiagen) following the manufacturer’s instructions. After verifying the RNA integrity on an Agilent 2100 bioanalyzer (Agilent Technologies, USA), only samples with an RNA integrity number (RIN) equal to or above 8 were used. Libraries were sequenced on a BGISEQ-500 platform and single-end reads of 50 base pairs (bp) length were generated per sample.
Identification of tissue and stage-specific genes
Good-quality raw reads were aligned against the turbot genome assembly (ASM318616v1) using STAR v2.7.0e alignment software23 and subsequently assigned to unique genes with HTseq software v0.10.024. Counts with less than 10 reads in at least 9 of our samples were removed. The read counts matrix was normalized using a median-of-ratios method with the R package DESeq225. We performed pair-wise comparisons of differential gene expression among stages and between tissues belonging to the same stage using DESeq2 and determined as differentially expressed genes (DEG) those with p-value < 0.05 and log2fold change > 1. We combined all DEGs, resulting from stages-by-stage comparison, in a single set and clustered in a heatmap to analyze stage-specific genes for any of the individual tissues. ClusterProfiler R package v3.14.326 was used for GO enrichment analysis, using a homemade functional annotation as background.
Gene expression pattern of tissue-stage interaction
To identify genes related to tissue-time interaction, we used the likelihood ratio test (LRT) statistic with DESeq2 package25. We used a full model with tissue, time, and their interaction. We removed tissue:stage interaction term for the reduced model, and identified as significant those genes with p-value < 0.05. We clustered the expression pattern of genes identified with LRT using DEGreport27 and selected those clusters that correspond to unique expression patterns over time for each tissue. GOs were analyzed using ClusterProfiler for genes that matched expression patterns of interest.
Ethical statements
Ethical approval (ES360570202002/17/FUN.1/BIOLAN.08/JRM) for all studies was obtained from the Institutional Animal Care and Use Committee of the IIM-CSIC Institute in accordance with the National Advisory Committee for Laboratory Animal Research Guidelines licensed by the Spanish Authority (RD53/2013). This work was in conformance with the European animal directive (2010/63/UE) for the protection of experimental animals.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Results
Morphological changes during turbot metamorphosis: skull bone asymmetry
Alizarin red staining was used to show developmental asymmetry in the skull during the turbot metamorphic process, followed by light sheet multiview imaging fluorescence microscopy system (LSFM) analysis and 3D reconstruction (Fig. 1). We observed that most of the skull bone structures (parietal, parasphenoid, supraorbital canal, and maxillary bones structures, dentary and premaxilla) exist symmetrically in the larval period in the early pre-metamorphic stages (10 dpf) (Fig. 1a, f, k) and the ocular position maintains the axis. The first evidence of asymmetry is observed in late pre-metamorphosis (15dpf) (Fig. 1b, g, l), where the separation between the frontal bones (left parietal and right parietal, blue color) decreases significantly, and the right parietal bone starts to become visibly narrower in anticipation of future ocular migration. However, at this stage, we do not yet observe the twisting of the supraorbital canal bone and the presence of nasal bone, which is drastically accentuated in early metamorphosis (20 dpf) (Fig. 1c, h, m) following the same direction that the eye migrates until the end of metamorphosis (Fig. 1e). The pseudomesial bar (red color), a peculiar bone present only in flatfishes, appears about the stage at early metamorphosis (20 dpf) (Fig. 1c, h, m) growing dorsally, as observed at the climax of metamorphosis (30 dpf) (Fig. 1d, i, n), and until completed at the end of metamorphosis (57 dpf) (Fig. 1e, j, o).
Whole animals were visualized with alizarin red staining. For each specimen, a suitable number of tiles were selected to cover the entire sample at lateral position (between 1 × 1 and 4 × 4). The images were captured using a 5× NA 0.16 plan-neofluar detection objective and a zoom of 0.5 to achieve a larger field of view. Visualizations of each specimen were created using the Arivis 4D software. a–e Lateral view, f–j frontal view and k–o dorsal view of (a, f, k) 10 dpf, (b, g, l) 15 dpf, (c, h, m) 20 dpf, (d, i, n) 30 dpf and (e, j, o) 57 dpf turbot. Dentary (dnt) (orange color), parietal (p) (blue color), pseudomesial bar (pb) (red color), supraorbital canal (sc) (blue color), pre-maxilla (pmx) (orange color), dpf (days post fertilization). Scale bar, 500 µm.
Transcriptome assembly and annotation
Eighteen cDNA libraries were created from three different stages of turbot development and from two tissues, the right and left eye and their surrounding tissue, using three independent replicates. More than 20 million 50 bp single-end reads were generated. The raw reads were checked for a good quality score. All clean, high-quality reads were mapped to the turbot reference genome, with averages of 88.96%, 88.73%, 89.19% in the migrant eye and 89.28%, 88.76%, 88.99% in the non-migrant eye at the pre-metamorphic, metamorphic climax, and post-metamorphic stages, respectively.
We performed principal component analysis (PCA) and samples belonging to the same stage were clustered together, showing a similar distribution pattern in the migrant and non-migrant eye over time (Fig. 2a).
a Principal Component Analysis (PCA) plot showing how samples are grouped according to their similarity. Color code: blue, pre-metamorphic stage; red, climax of metamorphosis; yellow, post-metamorphic stage. Shape code: circle, migrant eye; rhombus: non-migrant eye. b Heatmap and GOs enrichment of differentially expressed genes clustered only as overexpressed in the pre-metamorphic stage (blue bar), climax (red bar), and post-metamorphic stage (yellow bar) in the non-migrant eye (blue) and the migrant eye (red).
Pair-wise comparison using Wald test: stage-specific genes
We investigated the gene expression profile during metamorphosis in turbot eyes, taking three developmental stages as key points. To gain insight into the crucial genes at each stage during the metamorphosis process of turbot, we performed pairwise comparisons among stages in samples belonging to the same tissue. We obtained a total of 2857 differentially expressed genes (DEGs); 1375 genes overexpressed in the pre-metamorphic stage, 308 genes up-regulated during the climax of metamorphosis, and 1174 genes overexpressed in the post-metamorphic stage (Fig. 2b) (Supplementary Data 1–6).
We performed gene ontology (GO) enrichment analysis with genes overexpressed at each stage in both eyes. Genes upregulated in pre-metamorphic and post-metamorphic stages enriched similar GOs in both eyes (Fig. 2b). Thus, GOs enriched during the pre-metamorphic stage in both the migrant and non-migrant eye were related to early development processes such as regulation of neurogenesis (GO:0050767), anterior/posterior pattern specification (GO:0009952) or cell differentiation (GO:0030154) (Supplementary Data 7 and 8). By performing a GO enrichment analysis on genes that show a peak of expression during the climax of metamorphosis according to Wald test analysis in migrant and non-migrant eyes, we identified eye-specific GO terms (Fig. 2b). Thus, in the non-migrant eye we found enriched biological processes of muscle tissue development such as sarcomere organization (GO:0045214), muscle contraction (GO:0006936), skeletal muscle tissue development (GO:0007519), in contrast to the migrant eye in which we found biological processes related to the immune system such as immune response (GO:0006955) or immune system process (GO:0002376) (Supplementary Data 11 and 12). Interestingly, during the post-metamorphic stage, we also identified significant overlaps of GO terms between eyes that were related to the immune system and ocular development: visual perception (GO:0007601), lens development in the camera-type eye (GO: 0002088), immune response (GO: 0006955) or inflammatory response (GO: 0006954) (Supplementary Data 9 and 10). We highlight unique genes overexpressed in the post-metamorphic stage that is specifically expressed in the lens and photoreceptors outer segment (Fig. 3a, b). Focusing on those genes involved in the phototransduction cascade, we found that their expression pattern is almost identical in both eyes over time (Fig. 3c). However, we discovered differences in expression between isoforms expressed primarily in rods than those expressed in cones.
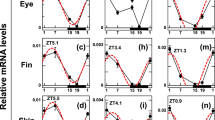
a Genes overexpressed at the post-metamorphosis stage in both eyes lens and retinal genes overexpressed at the post-metamorphosis stage in both eyes. b Representation of the main proteins involved in the phototransduction cascade in vertebrates. PDE phosphodiesterase, CNGC cyclic nucleotide-gated ion channel, NCKX sodium/calcium-potassium exchanger. In the boxes, the isoforms are expressed during the phototransduction cascade. c Gene expression profile throughout pre-metamorphic, climax, and post-metamorphic stages, from migrant and non-migrant eyes samples, of the main genes involved in the turbot phototransduction cascade. The expression profile of isoforms is primarily expressed in rods, and the expression profile of isoforms primarily expressed in cones. Created with BioRender (https://biorender.com).
Pair-wise comparison using Wald test: tissue-specific genes
On the other hand, to know the differences between eyes, we compared the gene expression of the migrant and the non-migrant eye at the pre-metamorphic stage, the climax stage of metamorphosis, and the post-metamorphic stage. We visualized the gene expression dataset using MAplot in order to identify gene expression changes between both eyes in each stage (Fig. 4a). We found large differences during the pre-metamorphic stage and climax, however in post-metamorphic stage, we did not identify marked differences in gene expression. A total of 1890 genes were differentially expressed; 1669 genes were upregulated in migrant eyes as opposed to 221 genes that were upregulated in non-migrant eyes for all the stages. Focusing on each developmental stage, 1026 genes were differentially expressed during pre-metamorphic stage (988 upregulated and 38 down-regulated in the migrant eye) (Supplementary Data 13), 737 genes during the climax of metamorphosis (609 upregulated and 128 genes down-regulated in the migrant eye) (Supplementary Data 14), and 127 genes in the post-metamorphic stage (72 upregulated and 55 genes downregulated in the migrant eye) (Supplementary Data 15). Focusing on the climax stage, we discovered genes expressed in the eye that do not migrate related to fast and slow muscle contraction, such as tnni2a.1 and tnni1, respectively. In addition, we also found different types of myosin, such as myh7I, myl13, or myh6. However, in the migrant eye, we found genes related to neuronal plasticities and synaptic activity, such as epd, bcan, or grpin1 (Fig. 4b).
a From top to bottom, gene expression comparison between migrant eye and non-migrant eye in the pre-metamorphic stage, climax of metamorphosis, and post-metamorphic stage, respectively. Blue dots, upregulated genes in the non-migrant eye; red dots, upregulated genes in the migrant eye. b Top 10 upregulated genes in each tissue and each stage. Blue semicircular bars represent genes upregulated in the non-migrant eye; red semicircular bars genes upregulated in the migrant eye. The figures illustrating turbot stages from individuals sampled by stage were photographed and edited by the authors.
Tissue-stage expression pattern using LRT
We used LRT analysis, a statistical tool of the goodness-of-fit between two models, to exhibit genes whose expression varies not only between developmental stages but also between tissues within developmental stages. Using LRT, we identified 2531 genes with adjusted p-value < 0.05 (Supplementary Data 16). Genes identified as tissue:developmental stage interaction were clustered, and each of the 2531 genes was assigned to one of the 20 clusters obtained. Those clusters in which expression in one of the tissues showed a concave pattern were selected (Fig. 5a). In the non-migrant eye, we found 289 genes that met these conditions (inter-tissue and developmental stage variability and a concave pattern for the non-migrant eye). Genes specifically expressed during climax in the non-migrant eye enrich GOs such as sarcomere organization (GO:0045214), muscle contraction (GO:0006936), skeletal muscle tissue development (GO:0007519) (Fig. 5b). However, for the migrant eye we did not find such a marked concave pattern. We found a few genes that, during the climax stage of metamorphosis exhibit a single peak. Generally, genes with elevated expression at the climax were also up-regulated in the pre-metamorphic stage. In this case, we found 613 genes that enriched GOs related to the nervous system, such as positive regulation of synapse assembly (GO:0051965), synaptic transmission (GO:0035249), glutamatergic or modulation of chemical synaptic transmission (GO:0050804) (Fig. 5b).
a Selected clusters of genes that show a peak of expression during climax, unique to the non-migrant eye (blue), and genes showing a peak of expression during climax, unique to the migrant eye (red). b GOs of genes fitting expression pattern shows in (a). Blue: GOs of genes belonging to the clusters selected as specific to the non-migrant eye; red: GOs of migrant eye-specific genes.
Discussion
As mentioned earlier, flatfish, including the turbot, undergo one of the most remarkable metamorphoses in the animal kingdom, involving morphological and behavioral changes, transitioning from a symmetric larva with a pelagic lifestyle to a completely asymmetric juvenile with a benthic lifestyle. However, one of the most fascinating aspects of this astonishing metamorphic process is the migration of one eye to the other side of the body.
Our morphological study of this process using advanced imaging techniques such as Light sheet Multiview Imaging fluorescence microscopy (LSFM) system analysis and high-resolution three-dimensional image reconstruction (Fig. 1) reveals that changes in skull shape, particularly in the frontal bones (right and left parietal), take place before ocular migration is readily discernible. These findings support the idea that ocular migration and cranial asymmetry develop as slightly asynchronous processes. In addition, we observed that the pseudomesial bar, a bone unique to flatfishes, forms during the process of eye migration. Therefore, asymmetric presence and asymmetry of the pseudomesial bar on the blind side is possibly responsible for the migration of the right eye during metamorphosis in turbot, similar to what has already been demonstrated in Japanese flounder28.
Although the molecular basis seems clear, with thyroid hormone being essential for eye migration in flatfish29,30,31, the network of regulatory mechanisms that allows an eye to move across the head to accommodate its partner remains an undiscovered puzzle where different biochemical processes and physiological functions may be altered.
Therefore, we propose this transcriptomic study in turbot eyes, aiming to connect the genome to gene function by uncovering transcriptionally active genes during the specific right eye migration in turbot that takes place during the metamorphosis process with further adaptations imposed by the ecological environment.
Transcriptomics analyses comparing the two eyes have unexpectedly revealed to us that immune-related genes are significatively up-regulated, particularly in the migrant eye during metamorphosis, compared to other developmental stages analyzed. Because of ocular immune privilege, the mechanism by which the eye limits the local inflammatory response to preserve vision32, we would not expect genes such as dhx58, irf7, or ifih1 to be overexpressed in eyes. However, our samples include the surrounding ocular tissue, so we cannot confirm that there is no immune privilege in the ocular tissue of metamorphic flatfish. Nevertheless, immune system signaling pathways are activated during tissue reorganization, and inflammation in the eyes promotes axon regeneration after ocular injury33. Our data also show an increased expression during the climax and post-metamorphic stage of the CIITA gene, which controls MHCII expression34. CIITA expression, probably induced by IFNγ34,35, could play a role in the maturation and reorganization of the immune system36 from larva to juvenile, as well as in recognizing and eliminating larval tissue that drives morphological remodeling around eyes37. However, we found no expression differences in the lc3b gene, described in autophagy processes around the eye during metamorphosis of Paralichtlys olivaceus38, between the migrant eye and the non-migrant eye, nor between different stages.
Analyzing eye-specific genes during the climax of metamorphosis, we found specifically in non-migrant eye, overexpressed myosin genes such as myhb, myl13 and mybpc1, which play an important role in skeletal muscle development or klhl41a involved in sarcomere organization. In contrast, in the migrant eye, we highlight the expression pattern of the epd gene, which, interestingly, seems to be involved in the sharpening of synaptic connections in optic nerve regeneration39. Although other authors have found no differences in neural projections between the migrant and the non-migrant eye in flatfish40, our results show a particular peak of epd gene expression during the climax phase of metamorphosis in the migratory eye, and we believe that the peculiar expression pattern of the epd gene shown in our results could open the debate on the asymmetry of visual projections between the eyes in flatfish. On the other hand, GOs related to the nervous system are specifically enriched in the migrant eye. It is clear that adaptative changes in the central nervous system, as well as changes in neuronal connectivity, are necessary for successful settlement41. Thus, our data show that these changes are most evident in the migrant eye and/or in its surrounding tissue, identifying asymmetry in nerve tissue29.
Accompanying the ocular migration, there is a cranial remodeling that is practically focused on the flexion of the frontal bones and the formation of pseudomesial bar28, as shown in our images (Fig. 1). In this study, we identified a specific pattern, with significant overexpression in the migrant-eye side and no significant differences along the timeline in the non-migrating eye, of the alx3 gene. This gene is specifically expressed in frontonasal neural crest cells and associated with craniofacial disorders42,43,44, and appears to play a key role in craniofacial development in several organisms45, exerting control over the specific timing of differentiation and cell morphologies in frontonasal neural crest cells46. The expression profile obtained for this gene suggests that localized and differential expression of alx3 during metamorphosis may contribute to skull restructuring, coordinating the modulation of bone growth that will allow the unique morphological adaptation observed in flatfishes. Although some genes showed asymmetry in expression between eyes, there was no significant differential expression of genes related to the thyroid hormones cascade, such as trb, trab, and traa, between the migrant and non-migrant eye. The expression profile of thyroid hormone action genes in our data is consistent with that described in other studies47,48 and is equal in both eyes. This lack of substantial variation is unexpected, given the different developmental pathways of the two eyes. However, the expression of matrix metalloproteinase genes, such as mmp9, mmp14, or sparc, required for tissue resorption and remodeling49, is much more pronounced during metamorphosis in the migrating eye compared to the non-migrating eye. In the postmetamorphic stage, the expression of these genes remains elevated in the non-migrating eye.
Based on our findings and previous results from other studies, although one eye undergoes a dramatic migration involving changes described above in muscle, nervous system, and bone, there appear to be no differences in development between the migrant and non-migrant eye. However, we do find significant differences in the development of the eye itself throughout the different stages related to the imposed habitat change. As previously mentioned, the turbot larvae undergo a habitat transition from a pelagic to a benthic state during the metamorphosis process. Life in the benthos is associated with reduced light, which triggers a reorganization of photoreceptors in many fish. This reorganization is critical in establishing the asymmetric pigmentation pattern in the skin of flatfish50 and in developing scotopic vision in the retina, as observed in sole51. Thus, this new lifestyle requires changes in the expression of various opsins to adapt to the new conditions of light restriction49,52. Our results show that rh1, an opsin involved in scotopic vision and expressed in rod photoreceptors, is highly expressed during the post-metamorphic stage when the fish has completed settlement.
In contrast, the gene expression of long-wavelength sensitive opsins and medium-wavelength opsins (rh2a and rh2b) decrease throughout ontogeny. These changes in the expression pattern of opsins are consistent with the adaptation to the different environments to which flatfish are exposed during development49. Additionally, the primary isoforms of the phototransduction cascade present in the postmetamorphic stage are primarily expressed in the rods responsible for scotopic vision. These include gucy2f, pde6a, pde6g, and cnga1a, which exhibit comparable expression patterns in both eyes. It is important to note that our study also found a decrease in opn1sw2 gene expression during metamorphosis, which is consistent with previous research. Opn1sw2 is believed to promote retinoic acid synthesis, which could contribute to inhibiting the metamorphosis process50. In addition, we found overexpression of lens genes (cryaa, crygs4, crygm3, cryba1b, cryba2b, cryba4, and crybb1), which enrich lens development in camera-type eye GO at the post-metamorphic stage in both eyes. So, our data indicate that the visual function of the migrant eye remains unaltered even after a significant metamorphosis process.
In conclusion, this study uncovers the intricate biological orchestra behind turbot metamorphosis, revealing a fascinating mix of morphological, genetic, and environmental complexities. The study highlights the ocular migration and cranial restructuring, which involves the formation of the pseudomesial bar and frontal bone flexion. These processes are supported by specific genetic expressions, such as the alx3 gene in craniofacial development and differential activation of immune-related genes. This study intricately links asynchronous eye development in the same individual during turbot metamorphosis with differential expression of musculoskeletal and regenerative genes. The non-migrant eye exhibits specific overexpression of musculoskeletal genes, which are crucial in skeletal muscle development, and this aligns with the relatively static structural demands of this eye. In contrast, the migrant eye exhibits a higher prominence of regeneration genes, specifically the epd gene, which is involved in synaptic connections during optic nerve regeneration. This reflects the dynamic and complex developmental processes required for ocular migration. This deeper understanding of the genetic underpinnings of eye-specific development provides novel insights into the intricate coordination of growth and adaptation in response to both internal genetic programming and external environmental challenges.
Data availability
RNA-seq data from all 18 samples representing the three developmental stages have been deposited in the NCBI Gene Expression Omnibus (NCBI GEO). Both raw files in fastq format and processed files can be readily accessed via GEO: Series GSE245741 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE245741)50.
References
Zimmer, C. The evolution of extraordinary eyes: the cases of flatfishes and stalk-eyed flies. Evol. Educ. Outreach 1, 487–492 (2008).
Friedman, M. The evolutionary origin of flatfish asymmetry. Nature 454, 209–212 (2008).
Schreiber, A. M. Asymmetric craniofacial remodeling and liberalized behavior in larval flatfish. J. Exp. Biol. 209, 610–621 (2006).
Bolstad, K., & Novales Flamarique, I. Photoreceptor distributions, visual pigments and the opsin repertoire of Atlantic halibut (Hippoglossus hippoglossus). Sci. Rep. 12, 1–24 (2022).
Evans, B. I. & Fernald, R. D. Retinal transformation at metamorphosis in the winter flounder (Pseudopleuronectes americanus). Vis. Neurosci. 10, 1055–1064 (1993).
Hoke, K. L., Evans, B. I. & Fernald, R. D. Remodeling of the cone photoreceptor mosaic during metamorphosis of flounder (Pseudopleuronectes americanus). Brain. Behav. Evol. 68, 241–254 (2006).
Villegas, E. D. M., Dans, M. J. D., Castillo, C. P. A. & Alvarez, R. A. Development of the eye in the turbot Psetta maxima (Teleosti) from hatching through metamorphosis. J. Morphol. 233, 31–42 (1997).
Ballard, K. A., Pickett, R. L. & Sivak, J. G. Comparison of the musculoskeletal structure of the orbits of the migrating and non-migrating eyes in the winter flounder (Pseudopleuronectes americanus). Exp. Biol. 47, 23–26 (1987).
Wagemans, F., Focant, B. & Vandewalle, P. Early development of the cephalic skeleton in the turbot. J. Fish. Biol. 52, 166–204 (1998).
Saele, O., Smáradóttir, H. & Pittman, K. Twisted story of eye migration in flatfish. J. Morphol. 267, 730–738 (2006).
Wagemans, F. & Vandewalle, P. Development of the cartilaginous skull in solea solea: trends in pleuronectiforms. Ann. Sci. Nat. 20, 39–52 (1999).
Murray, M. Axonal transport in the asymmetric optic axons of flatfish. Exp. Neurol. 42, 636–646 (1974).
Parker, G. The optic chiasma in teleost and its bearing. Bull. Mus. Comp. Zool. 40, 219 (1903).
Briñón, J. G. et al. Volumetric analysis of the telencephalon and tectum during metamorphosis in a flatfish, the Turbot Scophthalmus maximus. Brain Behav. Evol. 41, 1–5 (1993).
Tata, J. R. Amphibian metamorphosis as a model for studying the developmental actions of thyroid hormone. Cell Res. 8, 259–272 (1998).
de Jesus, E. G., Hirano, T. & Inui, Y. Flounder metamorphosis: its regulation by various hormones. Fish. Physiol. Biochem. 11, 323–328 (1993).
Gomes, A. S., Alves, R. N., Rønnestad, I. & Power, D. M. Orchestrating change: the thyroid hormones and GI-tract development in flatfish metamorphosis. Gen. Comp. Endocrinol. 220, 2–12 (2015).
Sæle, Ø., Silva, N. & Pittman, K. Post-embryonic remodelling of neurocranial elements: a comparative study of normal versus abnormal eye migration in a flatfish, the Atlantic halibut. J. Anat. 209, 31–41 (2006).
Bao, B. et al. Proliferating cells in suborbital tissue drive eye migration in flatfish. Dev. Biol. 351, 200–207 (2011).
Walker, M. B. & Kimmel, C. B. A two-color acid-free cartilage and bone stain for zebrafish larvae. Biotech. Histochem. 82, 23–28 (2007).
Al-Maghazachi, S. J. & Gibson, R. The developmental stages of larval turbot, Scophthalmus maximus (L.). J. Exp. Mar. Biol. Ecol. 82, 35–51 (1984).
Suarez-Bregua, P., Pérez-Figueroa, A., Hernández-Urcera, J., Morán, P. & Rotllant, J. Temperature-independent genome-wide DNA methylation profile in turbot post-embryonic development. J. Therm. Biol. 88, 1–7 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-A Python framework to work with high-throughput sequencing data. Bioinformatics https://doi.org/10.1093/bioinformatics/btu638 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. ClusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16, 284–287 (2012).
Pantano, L. DEGreport: Report of DEG Analysis. (2023).
Okada, N., Takagi, Y., Seikai, T., Tanaka, M. & Tagawa, M. Asymmetrical development of bones and soft tissues during eye migration of metamorphosing Japanese flounder, Paralichthys olivaceus. Cell Tissue Res. 304, 59–66 (2001).
Campinho, M. A. et al. A thyroid hormone regulated asymmetric responsive centre is correlated with eye migration during flatfish metamorphosis. Sci. Rep. 8, 1–13 (2018).
Manchado, M., Infante, C., Asensio, E., Planas, J. V. & Cañavate, J. P. Thyroid hormones down-regulate thyrotropin β subunit and thyroglobulin during metamorphosis in the flatfish Senegalese sole (Solea senegalensis Kaup). Gen. Comp. Endocrinol. 155, 447–455 (2008).
Miwa, S. & Inui, Y. Effects of various doses of thyroxine and triiodothyronine on the metamorphosis of flounder (Paralichthys olivaceus). Gen. Comp. Endocrinol. 67, 356–363 (1987).
Schepens, T. Immune privilege in the eye: an evolutionary adaptation. Dev. Comp. Immunol. 18, 421–431 (1994).
Wong, K. A. & Benowitz, L. I. Retinal ganglion cell survival and axon regeneration after optic nerve injury: role of inflammation and other factors. Int. J. Mol. Sci. 23, 10179 (2022).
Boss, J. M. Regulation of transcription of MHC class II genes. Curr. Opin. Immunol. 9, 107–113 (1997).
Chang, B. C., Fontes, J. D., Peterlin, M. & Flavell, R. A. Class II transactivator (CIITA) is sufficient for the inducible expression of major histocompatibility complex class II genes. J. Exp. Med. 180, 1367–1374 (1994).
McGuire, C. C. & Robert, J. R. Developmental exposure to thyroid disrupting chemical mixtures alters metamorphosis and post-metamorphic thymocyte differentiation. Curr. Res. Toxicol. 3, 100094 (2022).
Izutsu, Y. The immune system is involved in Xenopus metamorphosis. Front. Biosci. 14, 141–149 (2009).
Gao, L., Huang, Y., Sun, M. & Bao, B. The role of autophagy on eye migration during the metamorphosis of Paralichthys olivaceus. Cells Dev. 169, 203751 (2022).
Schmidt, J. T. & Shashoua, V. E. Antibodies to ependymin block the sharpening of the regenerating retinotectal projection in goldfish. Brain Res. 446, 269–284 (1988).
Medina, M., Rep, J., Ward, R., Rio, P. & Lemire, M. The primary visual system of flatfish: an evolutionary perspective. Anat. Embryol. 187, 167–191 (1993).
Graf, W. & Baker, R. Neuronal adaptation accompanying metamorphosis in the flatfish. J. Neurobiol. 21, 1136–1152 (1990).
Beverdam, A., Brouwer, A., Reijnen, M., Korving, J. & Meijlink, F. Severe nasal clefting and abnormal embryonic apoptosis in Alx3/Alx4 double mutant mice. Development 3986, 3975–3986 (2001).
Twigg, S. R. F. et al. Frontorhiny, a distinctive presentation of frontonasal dysplasia caused by recessive mutations in the ALX3 Homeobox Gene. Am. J. Hum. Genet. 84, 698–705 (2009).
Lakhwani, S., García-Sanz, P. & Vallejo, M. Alx3-deficient mice exhibit folic acid-resistant craniofacial midline and neural tube closure defects. Dev. Biol. 344, 869–880 (2010).
McGonnell, I. M. et al. Evolution of the Alx homeobox gene family: parallel retention and independent loss of the vertebrate Alx3 gene. Evol. Dev. 13, 343–351 (2011).
Mitchell, J. M. et al. The alx3 gene shapes the zebrafish neurocranium by regulating frontonasal neural crest cell differentiation timing. Development 148, 1–12 (2021).
Alves, R. N. et al. The transcriptome of metamorphosing flatfish. BMC Genomics 17, 1–24 (2016).
Louro, B., Marques, J. P., Manchado, M., Power, D. M. & Campinho, M. A. Sole head transcriptomics reveals a coordinated developmental program during metamorphosis. Genomics 112, 592–602 (2020).
Wang, Y. et al. Evolutionary ecology of the visual opsin gene sequence and its expression in turbot (Scophthalmus maximus). BMC. Ecol. Evol. 21, 114 (2021).
Guerrero-Peña, L. et al. Unravaling the transcriptomic landscape of eye migration and visual adaptations during flatfish metamorphosis data sets. Gene Expression Omnibus https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE245741 (2024).
Frau, S., Novales Flamarique, I., Keeley, P. W., Reese, B. E. & Muñoz-Cueto, J. A. Straying from the flatfish retinal plan: cone photoreceptor patterning in the common sole (Solea solea) and the Senegalese sole (Solea senegalensis). Physiol. Behav. 176, 139–148 (2017).
Mader, M. M. & Cameron, D. A. Photoreceptor Differentiation during Retinal Development Growth and Regeneration in a Metamorphic Vertebrate. J. Neurosci. 24, 11463–11472 (2004).
Acknowledgements
This study was financed by funds from MCIN/AEI/10.13039/501100011033 (AGL2017-89648P and PID2021-126511OB-100 to J.R.) and by “ERDF A way of making Europe”, by the “European Union”. L.G. was the recipient of an FPI predoctoral fellow (PRE2018-085475) from the Spanish State Agency of Research (AEI). The authors thank to Katherina García and José María Urbano from the Advanced Microscopy and Imaging Service at CABD-CSIC for their invaluable assistance and expertise in microscopy, which significantly contributed to the success of this research, and Ms. Carolina Costas for maintenance of the experimental animals and assistance during sampling.
Author information
Authors and Affiliations
Contributions
Conceptualization: P.S.-B., J.J.T. and J.R.; methodology and investigation: L.G.-P., P.S.-B., L.M.-M., J.J.T., L.S.-R.; validation: L.G.-P., P.S-B.; formal analysis: L.G.-P., P.S.-B., J.J.T.; resources: R.T., P.G.-F., and J.R; data curation: L.G-P., P.S-B., J.J.T.; writing—original draft preparation: L.G.-P., P.S.-B., and J.R.; writing—review and editing: J.J.T., J.R.; supervision, P.S.-B. and J.R.; project administration: J.R.; and funding acquisition: J.R. All authors have read and agreed to the published version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Alexander Schreiber, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Luke R. Grinham.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guerrero-Peña, L., Suarez-Bregua, P., Sánchez-Ruiloba, L. et al. Unraveling the transcriptomic landscape of eye migration and visual adaptations during flatfish metamorphosis. Commun Biol 7, 253 (2024). https://doi.org/10.1038/s42003-024-05951-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-05951-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.