Abstract
Gene fusions are known to drive many human cancers. Therefore, the functional characterization of newly discovered fusions is critical to understanding the oncobiology of these tumors and to enable therapeutic development. NPM1–TYK2 is a novel fusion identified in CD30 + lymphoproliferative disorders, and here we present the functional evaluation of this fusion gene as an oncogene. The chimeric protein consists of the amino-terminus of nucleophosmin 1 (NPM1) and the carboxyl-terminus of tyrosine kinase 2 (TYK2), including the kinase domain. Using in vitro lymphoid cell transformation assays and in vivo tumorigenic xenograft models we present direct evidence that the fusion gene is an oncogene. NPM1 fusion partner provides the critical homodimerization needed for the fusion kinase constitutive activation and downstream signaling that are responsible for cell transformation. As a result, our studies identify NPM1–TYK2 as a novel fusion oncogene and suggest that inhibition of fusion homodimerization could be a precision therapeutic approach in cutaneous T-cell lymphoma patients expressing this chimera.
Similar content being viewed by others
Introduction
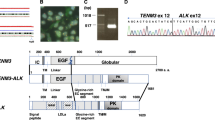
Gene fusions resulting from chromosomal rearrangements create chimeric proteins that play a significant role in the pathogenesis of various lymphomas and leukemias1,2,3. Deciphering the oncogenic biology of new chromosomal translocations is essential to understanding the molecular mechanisms of disease, which is vital to the development of therapeutic interventions4,5,6. NPM1–TYK2 (t; 5:19) is the first identified chimeric tyrosine kinase in 4% of CD30+ lymphoproliferative disorders (LPDs)7. Cutaneous CD30+ LPDs, the second most common type of cutaneous T-cell lymphoma (CTCL), include a clinicopathologic spectrum of benign lymphomatoid papulosis and primary cutaneous anaplastic large-cell lymphoma (ALCL)8. The fusion partners in this arrangement are the nucleophosmin 1 (NPM1) gene (5q35) and the tyrosine kinase 2 (TYK2) gene (19p13). NPM1 is a nucleolar phosphoprotein that performs diverse biological functions, including molecular chaperoning, ribosome biogenesis, DNA repair, and maintaining genomic stability9,10. TYK2 is a non-receptor tyrosine kinase that belongs to the Janus kinases (JAKs) family is associated with cytokine and growth factor receptors which activate STAT signaling11,12. NPM1–TYK2 is an 81 kDa chimera, comprised of the amino-terminal 257 amino acids of the NPM1 protein and carboxyl-terminal 461 amino acids length of the TYK2 protein, including the catalytic domain. The NPM1–TYK2 fusion consists of an almost full-length NPM1 protein except for the last 37 amino acids of the carboxyl-terminus, this contrasts with other NPM1 fusion proteins in which the NPM1 fusion partner is mostly limited to the oligomerization domain13,14,15.
A previous report documented an activated NPM1–TYK2 in vitro, but the characterization of the fusion protein was quite limited7. To understand the pathobiology of the fusion, it is essential to analyze the novel fusion kinase oncogenesis in relevant biological assays. In the present study, we focused on the functional characterization of NPM1–TYK2 utilizing both in vitro transformation assays and in vivo tumorigenicity in mouse xenograft models and detailed downstream signaling responsible for the oncogenicity. Our studies elucidate the oncogenic partnering role of NPM1 in the transformation potential of the NPM1–TYK2 fusion protein. Overall, our in vitro and in vivo models validate NPM1–TYK2 as a novel fusion oncogene and represent essential tools for future drug screening efforts to develop potential targeted therapies for CD30+ LPDs with this fusion event.
Results
In vitro transformation potential of the NPM1–TYK2 fusion gene
The fundamental essay for establishing an oncogenic property of a new gene is the in vitro transformation assay16. The Ba/F3 in vitro transformation assay is a well-established method to determine the oncogenic potential of novel gene mutations in hematologic malignancies17. The Ba/F3 cell line requires IL-3 for normal proliferation and survival18. Oncogene overexpression in Ba/F3 cells allows for cytokine-independent growth through utilization of oncogene mediated survival signaling mechanisms19. The pCDH-EF1-MCS-T2A-Puro lentiviral expression plasmid was used to generate the overexpression constructs. pCDH-empty vector and pCDH-FG–NPM1–TYK2 lentiviral particles were made and transduced into Ba/F3 cells. Stable clones of the pCDH-empty vector (hereafter referred to as Ba/F3-Vector) and pCDH-FG–NPM1–TYK2 (hereafter referred to as Ba/F3–NPM1–TYK2) were selected using puromycin antibiotic selection. Once stable cell lines were established, we used total RNA from Ba/F3-NPM1–TYK2 cells to measure NPM1–TYK2 fusion transcript by real-time quantitative polymerase chain reaction (RT-qPCR). NPM1–TYK2 mRNA transcript levels in Ba/F3–NPM1–TYK2 cells were compared to Myla cells (i.e., the only available NPM1–TYK2 endogenously expressing cell line) while the SU-DHL-1 (NPM1-ALK+ ALCL) T-cell lymphoma cell line served as a negative control (Fig. 1A)20. We further examined the subcellular localization of the chimeric protein by immunofluorescence assay. As shown in Fig. 1B, our immunofluorescence data indicates NPM1–TYK2 fusion protein distribution in both nuclear and cytoplasmic compartments. To explore the oncogenic potential of the NPM1–TYK2 fusion gene, Ba/F3-Vector, and Ba/F3–NPM1–TYK2 cells were grown in the medium without IL-3 for seven days. As shown in Fig. 1C., vector control cells were unable to survive in the absence of IL-3 in the culture medium. In contrast, stable expression of NPM1–TYK2 in Ba/F3 cells promoted IL-3 independent growth utilizing NPM1–TYK2 mediated oncogenic signaling as a survival mechanism (Fig. 1C). Next, we examined the clonogenic potential of transformed Ba/F3–NPM1–TYK2 and Ba/F3-Vector cells by incubating each cell line in Methocult medium without IL-3 supplementation for seven days to allow colony formation. Ba/F3 cells expressing the NPM1–TYK2 fusion gene were able to produce colonies due to the oncogenic gain-of-function, while vector control cells could not survive in the absence of IL-3 growth factor (Fig. 1D). Thus, our in vitro transformation assay results with Ba/F3 cells demonstrate the oncogenic potential of the NPM1–TYK2 gene.
A Detection of overexpressed NPM1–TYK2 mRNA transcript levels in transformed Ba/F3 cells by qRT-PCR. Myla and SU-DHL-1 cell lines were used as positive and negative controls, respectively. Columns represent the mean of three independent experiments; bars represent the SEM. B Subcellular localization of NPM1–TYK2. Ba/F3-Vector and transformed Ba/F3 (NPM1–TYK2) cells were cytospun onto glass slides, fixed, permeabilized, and stained for FLAG antibody and DAPI. Images were acquired with a fluorescent microscope using a 60× oil immersion lens. C Transformation of Ba/F3 cells to IL-3 independent growth. Ba/F3-Vector and transformed Ba/F3 (NPM1–TYK2) cells were grown in RPMI medium in the absence of IL-3. Cell viability from each condition was determined daily by trypan blue exclusion assay. Points represent the mean of three independent experiments; bars represent SEM. ∗∗∗P < 0.0005 was considered as statistically extremely significant. D The clonogenic potential of transformed Ba/F3 (NPM1–TYK2) cells. Ba/F3-Vector and transformed Ba/F3 NPM1–TYK2 cells were mixed with MethoCult medium and plated. After 7 days of incubation, the number of colonies was counted. Columns represent the mean of three independent experiments; bars represent the SEM.
Intracellular signaling that governs lymphoid cell transformation
Cancer cells acquire transformation potential through upregulation of survival signaling pathways resulting from oncogenic genetic alterations21. Once we confirmed the oncogenic potential of NPM1–TYK2, we investigated the critical signaling pathway responsible for fusion kinase-driven oncogenicity. In several cancers, TYK2 gene mutations result in constitutive activation of TYK2 kinase leading to upregulation of downstream STAT signaling22,23. Therefore, we examined Ba/F3–NPM1–TYK2 cells for fusion kinase activation and downstream STAT signaling. Our Western blot analysis data clearly indicate the upregulation of phosphorylated NPM1–TYK2 in transformed Ba/F3–NPM1–TYK2 cells. Activation of NPM1–TYK2 fusion kinase resulted in phosphorylation of downstream signaling molecules STAT1, STAT3, and STAT5 (Fig. 2). In contrast, Ba/F3-vector cells showed no activation of fusion kinase or downstream STAT signaling (Fig. 2). Collectively, our NPM1–TYK2 fusion kinase signaling results indicate that Ba/F3–NPM1–TYK2 cells acquired transformation potential through constitutive activation of fusion kinase and downstream STAT signaling.
NPM1–TYK2 fusion kinase activation is an oncogenic driver in lymphoid cell transformation
To further validate the fusion kinase-driven transformation of Ba/F3 cells, we developed a stable Ba/F3–NPM1–TYK2–K462R kinase-dead mutant cell line. Cell lysates were made from Ba/F3-vector, Ba/F3–NPM1–TYK2, Ba/F3–NPM1–TYK2–K462R cell lines, and Western blot analysis was performed for phospho-NPM1–TYK2, phospho-STAT1/3/5, and respective total proteins. Ba/F3-NPM1–TYK2 and Ba/F3-NPM1–TYK2–K462R cells showed ectopic overexpression of FLAG (NPM1–TYK2) protein, whereas no expression was observed in Ba/F3-vector cells (Fig. 3A). Furthermore, wild-type fusion kinase expressing Ba/F3–NPM1–TYK2 cells showed activated fusion kinase and upregulation of downstream STAT1/3/5, whereas no activation of kinase was observed in Ba/F3-vector and Ba/F3–NPM1–TYK2–K462R cells (Fig. 3A). We also examined the IL-3 independent growth and transformation potential of the NPM1–TYK2 kinase-dead mutant compared to wt-NPM1–TYK2 and vector control cells. Ba/F3 vector and NPM1–TYK2 kinase-dead mutant cells were unable to survive without IL-3 in the medium, however, Ba/F3 cells expressing wt-NPM1–TYK2 cells showed IL-3-independent growth (Fig. 3B). The lack of signaling and cell viability in the absence of IL3 of fusion kinase-dead mutant cells clearly demonstrates that the NPM1–TYK2 fusion kinase activity is the oncogenic driver responsible for Ba/F3 cell transformation.
A Fusion kinase-dead mutant blocks kinase activation and downstream signaling. Cell lysates were made from Ba/F3-vector, Ba/F3–NPM1–TYK2, and Ba/F3–NPM1–TYK2–K462R cell lines, and immunoblot analysis was performed for FLAG (NPM1–TYK2), phospho-NPM1–TYK2, phospho-STAT1/3/5, and respective total proteins. β-Actin served as an internal control. B Fusion kinase signaling drives cell transformation. Ba/F3-Vector, Ba/F3–NPM1–TYK2, and Ba/F3–NPM1–TYK2–K462R cells were grown in RPMI medium in the absence of IL-3. Cell viability from each condition was determined daily by trypan blue exclusion assay. Points represent the mean of three independent experiments; bars represent SEM. ∗∗∗P < 0.0005 was considered as statistically extremely significant.
In vivo tumorigenic potential of the NPM1–TYK2 fusion gene
Based on the in vitro transformation potential of the NPM1–TYK2 fusion gene, we further investigated the fusion gene’s tumorigenic potential in an in vivo xenograft model. To assess NPM1–TYK2 tumorigenicity, we subcutaneously injected Ba/F3-Vector or transformed Ba/F3–NPM1–TYK2 cells into the flanks of 5- to 7-week-old female Hsd:Athymic Nude-Foxn1nu mice (Envigo, Indianapolis, IN). The control mice (six per group) injected with Ba/F3-Vector cells showed no evidence of tumor formation, while the mice (six per group) injected with transformed Ba/F3–NPM1–TYK2 cells developed palpable tumors in 3 days (Fig. 4A, B). Tumor progression was rapid and reached approximately 1400 mm3 in size within 15 days (Fig. 4D). The size and weight of lymphoma tumors excised from NPM1–TYK2 mice are shown in Fig. 4C. In addition to tumor growth, Ba/F3–NPM1–TYK2 mice also developed splenomegaly and hepatomegaly, while control mice did not (Fig. 4E, F).
A Representative image of mice (n = 6) subcutaneously injected with Ba/F3-Vector showed no tumors. B Representative images of mice (n = 6) subcutaneously injected with transformed Ba/F3–NPM1–TYK2 show tumorigenesis. C Representative images of resected tumors with their respective tumor weight (grams). D Tumor volume was increased significantly in transformed Ba/F3–NPM1–TYK2 injected mice but not in Ba/F3-Vector mice. Bars represent the SEM. E, F In transformed Ba/F3 (NPM1–TYK2) mice, tumor growth was associated with hepatosplenomegaly.
Different analytical assays confirmed the fusion gene expression at mRNA and protein levels in tumor tissues. Our immunocytochemistry results confirm FLAG (NPM1–TYK2) expression in tumor cells (Fig. 5A). We further analyzed NPM1–TYK2 expression using RT-qPCR to measure fusion gene mRNA transcript levels in tumor tissue (Fig. 5B). Since tumor growth was associated with hepatosplenomegaly, we examined spleen and liver tissue samples of Ba/F3-Vector and Ba/F3–NPM1–TYK2 mice for lymphoma cell infiltration by H&E staining. Immunohistochemistry on the spleen and liver tissue from Ba/F3–NPM1–TYK2 injected mice demonstrated the infiltration of lymphoma cells, but not on tissue from Ba/F3-Vector mice (Fig. 5C).
Oncogenic NPM1–TYK2 signaling drives in vivo tumorigenicity
Upon establishing tumorigenicity in xenograft models, we evaluated the molecular pathways responsible for the fusion gene’s tumorigenic potential. Our studies further focused on exploring the activation of the fusion kinase and upregulation of STAT signaling in xenograft tumors, as observed in in vitro transformation assays. Tissue lysates were prepared from each excised frozen tumor and used for Western blot analysis. Since control mice did not develop tumors, we used respective flank tissue as control tissue to compare with tumor tissues. First, we analyzed NPM1–TYK2 expression levels with FLAG (as the overexpressed NPM1–TYK2 protein is tagged with amino-terminus FLAG epitope) and NPM1–TYK2 antibodies. We also examined phosphorylation of fusion kinase NPM1–TYK2, STAT1, STAT3, and STAT5 and respective total proteins. Results showed NPM1–TYK2 fusion protein expression in tumor tissues with both FLAG and NPM1–TYK2 antibodies, whereas no expression was observed in Ba/F3-Vector mice (Fig. 6). In NPM1–TYK2 tumor tissue, high fusion kinase activity resulted in the upregulation of phosphorylated STAT1, STAT3, and STAT5 (Fig. 6). These signaling data analyses clearly indicate that NPM1–TYK2 signaling mediates tumorigenicity in in vivo models and corroborates the in vitro transformation results.
NPM1 fusion partner is essential to NPM1–TYK2 mediated oncogenic signaling
It is essential to study each fusion partner’s role in oncogenic fusions to better understand disease pathogenesis, ultimately aiding in the development of potential targeted therapeutic interventions24. Since NPM1–TYK2 fusion is not fully characterized, here we explored the functional role of the amino-terminus of the NPM1 fusion partner in fusion-driven oncogenicity. To explore the functional role of the NPM1 fusion partner in fusion kinase activity, we deleted the NPM1 partner from full-length NPM1–TYK2 to generate an ΔN-1–257-NPM1–TYK2 deletion construct. We conducted our initial experiments in easy to transfect HEK293T cells. The empty vector, full-length NPM1–TYK2, or ΔN-1–257-NPM1–TYK2 deletion constructs were transfected into HEK293T cells and incubated for 48 h. Total cell lysates were prepared, and Western blot analysis was performed to understand NPM1 deletion effects on NPM1–TYK2 activity. As expected, there was no signal in vector cells (lane 1, Fig. 7A). Full-length NPM–TYK2 (lane 2, Fig. 7A) and NPM1-deletion (lane 3, Fig. 7A) fusion proteins were confirmed by FLAG and NPM1–TYK2 antibodies (Fig. 7A). The full-length fusion protein-induced phosphorylation of NPM1–TYK2 (lane 2, Fig. 7A), but NPM1-deleted fusion protein did not show fusion kinase phosphorylation (lane 3, Fig. 7A). NPM1–TYK2 overexpression induced activation of NPM1–TYK2, resulting in upregulation of STAT1/3/5 signaling as observed in Fig. 2 fusion signaling data. In contrast, deletion of the NPM1 portion from the fusion protein inhibited fusion kinase activity and downstream STAT signaling (lane 3, Fig. 7A). After studying the role of NPM1 fusion partner in HEK293T cells, we made lentiviral particles with ΔN-1–257-NPM1–TYK2 and transduced them into Ba/F3 cells. Stable clones were selected with puromycin and lysates were analyzed for NPM1–TYK2 signaling along with Ba/F3 cells expressing vector and full length-NPM1–TYK2 (Fig. 7B). As observed in HEK293T cells, NPM1 fusion partner deletion from the full-length fusion kinase inhibited NPM1–TYK2 activity and downstream signaling in Ba/F3 cells (Fig. 7B). Together, our results in both HEK293T and Ba/F3 cells exemplify the essential role of the NPM1 fusion partner in NPM1–TYK2 oncogenicity.
Loss of NPM1 partner abrogates NPM1–TYK2 transformation potential
Next, we tested the loss of the NPM1 partner’s influence on the viability of Ba/F3 cells. Ba/F3 cells expressing vector, full-length NPM1–TYK2, and ΔN-1–257-NPM1–TYK2 were grown in culture media without IL-3. Cell viability was measured using a trypan blue exclusion assay for ten days. As we observed in previous in vitro viability assays (Fig. 1C), full-length NPM1–TYK2 expressing Ba/F3 cells were viable, whereas vector and NPM1 deletion expressing cells were no longer viable (Fig. 8A). Additionally, we analyzed the proliferation potential of the NPM1–TYK2 fusion without the NPM1 fusion partner. We conducted clonogenic assays on vector, full-length NPM1–TYK2, and ΔN-1–257-NPM1–TYK2 in a Methocult medium without IL-3 growth factor. As shown in Fig. 8B, only the cells expressing full-length NPM1–TYK2 fusion kinase were able to proliferate due to constitutive activation of the fusion kinase and downstream survival signaling (Fig. 9). In contrast, deletion of NPM1 from the fusion kinase resulted in inhibition of both kinase activity and downstream signaling activation ultimately leading to cell death. Overall, these results demonstrate that the NPM1 fusion partner is essential to NPM1–TYK2 mediated clonogenicity.
A Deletion of NPM1 portion in NPM1–TYK2 fusion gene abrogates the transformation potential of the NPM1–TYK2 fusion gene in Ba/F3 cells. Ba/F3 cells were transduced with lentiviral particles expressing vector, NPM1–TYK2, and ΔN-1–257-NPM1–TYK2. Cells were grown in RPMI medium without IL-3 growth factor for 10 days. Cell viability from each condition was determined daily by trypan blue exclusion assay. Points represent the mean of three independent experiments; bars represent SEM. ∗∗∗P < 0.0005 was considered as statistically extremely significant. B The deletion of the NPM1 portion in the NPM1–TYK2 fusion gene inhibits the clonogenic potential. Columns represent the mean of three independent experiments; bars represent the SEM.
Discussion
Existing cancer genomics data has shown that gene fusions derived from chromosomal translocations are frequent oncogenic drivers of many hematologic and solid cancers25,26. Targeted therapies are raising hope as effective treatment options for fusion gene-driven maliganancies27,28,29. There has been limited progress in the development of targeted therapies for CTCL due to the lack of understanding of novel genetic alterations and related oncogenic signaling pathways involved in their pathogenesis30,31. An earlier study identified NPM1–TYK2 in CD30+ LPDs7, but the oncogenic potential of NPM1–TYK2 overexpression on lymphoid cell transformation or tumorigenicity in biological assays had not yet been explored. Therefore, our current studies focused on functional characterization of fusion gene oncogenicity utilizing in vitro and in vivo biological assays.
Understanding the protein subcellular localization is critical due to its influence on diverse cellular processes32. In general, wild-type (wt) proteins of NPM1 and TYK2 are localized in the nucleolus and cytoplasm, respectively. Our immunofluorescence data with transformed Ba/F3–NPM1–TYK2 cells showed fusion protein distribution in both nuclear and cytoplasmic compartments. A possible explanation for the nuclear distribution of NPM1–TYK2 is through its heterodimerization with wt-NPM1, as seen in NPM1-ALK expressing ALCL cells33.
Malignant cellular transformation is characterized by continuous uncontrolled proliferation34. The transformed cells are distinguished from normal counterparts by acquired alterations in growth patterns like anchorage-independent growth potential with no intercellular contact inhibition and xenograft formation35. First, we performed in vitro lymphoid cell transformation assays to determine the transformation potential of NPM1–TYK2 using IL-3-dependent lymphoid Ba/F3 cells. The dependency on cytokine signaling of Ba/F3 cell survival is extensively used in hematologic malignancies to understand the oncogenicity of novel kinases, fusion kinases, and downstream signaling of activated tyrosine kinases. Our in vitro transformation assay results distinctly demonstrate that NPM1–TYK2 overexpression leads to the transformation of Ba/F3 cells with survival and proliferation independent of IL-3. Subsequently, our studies focused on identifying the signaling network that is responsible for this transformation potential. In previous fusion kinase oncogenic characterization studies, Ba/F3 cells acquired IL-3-independence through fusion-driven signaling pathways36.
TYK2, the first identified JAK family member, is associated with cytokine and growth factor receptors which activate STAT signaling37. TYK2 is activated by various cytokines, including numerous interleukins and interferons38. TYK2 point mutations found in T-ALL (T-cell acute lymphoblastic leukemia) cell lines exhibited transformation potential via the STAT1/BCL2 pathway39. Several activating TYK2 mutations are known to trigger TYK2 signaling, which is essential to the development of T-ALL39. Screening of cancer datasets revealed more than fifty TYK2 chromosomal translocations, mostly in hematologic malignancies40. As shown in other TYK2 mutated cancers, we investigated NPM1–TYK2 fusion kinase hyperactivation and downstream STAT signaling by Western blot analysis. Our data displayed constitutive phosphorylation of the TYK2 fusion kinase and upregulated downstream effectors STAT1, 3, and 5 in transformed Ba/F3 cells. Furthermore, our NPM1–TYK2–K462R kinase-dead mutant cells showed abrogated signaling and cell viability, confirming that wt-NPM1–TYK2 mediated constitutive activation of fusion kinase is the oncogenic driver responsible for cell transformation.
In vivo xenograft models play an essential role in understanding molecular mechanisms and pathobiology of the disease41. Once our in vitro transformation studies characterized NPM1–TYK2 chimera’s oncogenic potential and survival signaling that governs transformation, we conducted in vivo studies using NPM1–TYK2 transformed xenograft models. In line with in vitro transformation, we demonstrated the in vivo tumorigenic potential of the fusion gene in NPM1–TYK2 transformed xenograft models. These mice developed robust tumor progression with associated hepatosplenomegaly. Further immunohistochemistry results revealed infiltration of lymphoma cells in tumors, spleen, and liver. From the excised tumors, signaling data revealed hyperactivation of fusion kinase and upregulation of STAT signaling.
Several fusion proteins involving receptor tyrosine kinases have demonstrated transformation potential in many solid cancers and hematologic malignancies25. The majority of these oncogenic fusion kinases contain the carboxyl-terminus kinase and amino-terminus non-kinase partners42. The oligomerization sequences of the amino terminus fusion partner are usually responsible for the constitutive activation of carboxyl-terminus kinase partner24. Studying the role of each fusion partner helps improve the understanding of disease pathogenesis and directly promotes the development of potential targeted therapeutic interventions. Here, we explored the amino-terminus NPM1 fusion partner’s functional role in fusion kinase oncogenicity. Native NPM1 protein exists as dimeric and oligomeric forms through the amino-terminal oligomerization domain13,43. In other NPM1-fusion proteins, the amino-terminus NPM1 fusion partner provides a self-dimerization interface for fusion kinase to form homodimers14,44. In our fusion mapping studies, NPM1 deletion from the NPM1–TYK2 fusion gene completely inhibited fusion gene activation and downstream STAT signaling. Further, in vitro transformation results showed that removal of the amino-terminus NPM1 fusion partner abrogated the transformation and clonogenic potential of the fusion kinase. Our results suggest that the NPM1 fusion partner facilitates a homodimerization interface for NPM1–TYK2 which is necessary for its constitutive activation and transformation potential.
In conclusion, our in vitro and in vivo preclinical findings provide functional evidence to identify NPM1–TYK2 as a novel fusion oncogene. Our lymphoid cell transformation and tumorigenicity in xenograft models represent tools necessary to understand the uncovered cellular mechanisms underlying the disease and to assist in future drug screening to develop precision therapies. Importantly, our fusion mapping studies suggest inhibition of NPM1–TYK2 homodimerization by targeting the NPM1 fusion partner could be a potential therapeutic strategy to treat CD30+ LPDs expressing the chimera. Additionally, future preclinical studies focused on inhibition of the TYK2 fusion partner must also be considered in the development of treatment for these fusion-driven lymphomas.
Methods
Ethics statement
All animal experiments were approved by the University of Kansas Medical Center institutional animal care and use committee and performed in accordance with relevant regulations and guidelines.
Cell culture
Ba/F3, SU-DHL-1 (obtained from DSMZ (Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Braunschweig, Germany), HEK293T, and WEHI-3B cell lines were obtained from American Type Culture Collection (ATCC, Manassas, VA, USA). Myla’s cell line was kindly provided by Ryan Wilcox, University of Michigan, USA. HEK293T and WEHI-3B cells were cultured in DMEM medium supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. Myla, SU-DHL-1, and Ba/F3 cells were maintained in RPMI-1640 medium supplemented with 10% FBS and 1% penicillin/streptomycin. Ba/F3 cells were additionally supplemented with a 10% WEHI-3B conditioned medium (as a source of interleukin 3/IL-3), and all stable clones were maintained with 2 µg/ml of puromycin.
Chemicals and reagents
All reagents and antibodies were purchased from the following: Thermo Scientific, Waltham, MA, USA (RPMI-1640-SH30027; DMEM-SH30081); Corning Life Sciences, USA (Fetal Bovine Serum-MT35010CV, Penicillin/Streptomycin-30-002-CI); InvivoGen, San Diego, CA, USA (Puromycin-58-58-2); STEMCELL Technologies, Vancouver, BC, Canada (MethoCult-H4100 −04100); Cell Signaling Technology, Beverly, MA, USA (TYK2/NPM1–TYK2-14193, phospho-TYK2/phospho-NPM1–TYK2-9321, pSTAT3-9145, STAT5-9363, STAT1-14994); BD Biosciences San Jose, CA, USA (STAT1-610185, phospho-STAT1-612132, STAT3-610189, Phospho-STAT5-611964, β-Actin-612656); Sigma-Aldrich, St. Louis, MO, USA (FLAG-F3165, Trypan Blue-T8154).
Cloning and generation of stable Ba/F3–FLAG–NPM1–TYK2 and Ba/F3–FLAG-∆1–257–NPM1–TYK2 cell lines
Human full-length NPM1–TYK2 cDNA7 was amplified by PCR. The PCR product was then gel purified, digested with XbaI and NheI restriction enzymes, and subcloned into a lentiviral plasmid pCDH-EF1-MCS-T2A-Puro vector (purchased from System Biosciences, Palo Alto, CA, USA). The colonies were screened, and positive clones were confirmed by Sanger DNA sequencing. The lentiviral plasmids, either empty vector (pCDH-EF1-MCS-T2A-Puro) or recombinant plasmid (pCDH-EF1-FG-NPM1–TYK2) were transfected along with packaging plasmid (psPAX2) and envelop plasmids (pMD2.G) into HEK293T cells using Fugene HD transfection reagent (Roche Applied Science). The lentiviral particles were collected after 48 h. Ba/F3 cells were transduced with either empty vector or pCDH-EF1-FG-NPM1–TYK2 lentivirus and the selection was carried out after 48 h with 2 µg/ml puromycin. Ba/F3-pCDH-vector cell growth was IL-3-dependent, while the transformed Ba/F3–FG–NPM1–TYK2 cell growth was IL-3 independent. The fusion gene NPM1–TYK2 mRNA expression levels were confirmed by qPCR and protein expression levels were confirmed by Western blotting using FLAG and NPM1–TYK2 antibodies. The deletion constructs Ba/F3–FG–∆N-1–257–NPM1–TYK2 were generated from NPM1–TYK2 cDNA using primers to amplify the amplicon spanning only the TYK2 portion. The clones were screened, and positive clones were confirmed by Sanger DNA sequencing. Lentiviral particles were made, as explained above. The deletion constructs protein expression levels were confirmed by Western blotting for FLAG and NPM1–TYK2. Stable cells expressing the deletion construct were IL-3 dependent due to the loss of the NPM1 fusion partner.
Generation of NPM1–TYK2 kinase-dead mutant stable cell line
FG-NPM1–TYK2-K462R kinase-dead mutant was created by performing Site-Directed Mutagenesis using site-specific mutagenesis primers (FP 5′-ACTGGCGAGATGGTGGCGGTGCGGGCCCTCAAGGCAGACTGCGGC-3′ and RP 5′-GCCGCAGTCTGCCTTGAGGGCCCGCAC CGCCACCATCTCGCCAGT-3′) and wt-NPM1–TYK2 cDNA as a template (Q5 site-directed mutagenesis kit, New England BioLabs Inc, Ipswich, Massachusetts, USA)7. The mutant sequence was confirmed by Sanger sequencing. Ba/F3 cells were transduced with FG–NPM1–TYK2–K462R lentiviral particles and stable clones were established using puromycin antibiotic selection. Ectopic kinase-dead fusion protein expression levels were confirmed by Western blot using FLAG antibody.
Western blot analysis
The cells were harvested, washed with phosphate-buffered saline (PBS), added in lysis buffer (25 mM Tris. HCl, 150 mM NaCl, 25 mM NaF, 0.5 mM Na-orthovanadate, 1% Triton-X, 1 mM Benzamidine) with protease and phosphatase inhibitors and incubated on ice for 20 min. Cell lysates were centrifuged at 10,000 RPM for 15 min, and protein concentrations were determined using the BCA protein assay kit (Pierce Biotechnology, Rockford, IL USA). The protein samples were resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis and immunoblotting was performed. The blots were scanned using the Odyssey IR scanner (Li-COR Biosciences, Lincoln, NE, USA). All blots derive from the same experiment and were processed in parallel.
Immunofluorescence
To analyze the cellular distribution of NPM1–TYK2, Ba/F3–FG–NPM1–TYK2 cell slides were prepared by cytospin technique. Similarly, Ba/F3-pCDH-vector cells were also processed and used as a control. Both control and FG–NPM1–TYK2 overexpressed cells were fixed with 4% paraformaldehyde and permeabilized with methanol/glacial acetic acid (3:1). After fixation, the cells were incubated with FLAG primary antibody overnight, washed with PBS, and then incubated with secondary antibody conjugated with Dylight-594 for an hour (D1-2594, Vector Laboratories, Burlingame, CA). Coverslips were mounted with Vectashield antifade mounting medium containing DAPI (4′, 6′-diamidino-2-phenylindole, Vector laboratories, Burlingame, CA). The images were acquired using an Eclipse E1000 microscope (Nikon).
Immunohistochemistry
Paraffin-embedded tissues were sectioned at 4 µM size and mounted on the glass slide. Sections were dewaxed in xylene overnight and rehydrated through in gradient ethanol series, followed by washing with PBS. Hematoxylin staining was performed using Leica Surgipath SelecTech Hematoxylin. This is followed by brief immersion in Richard-Allan Scientific Eosin-y with Phloxine, then dehydration with reagent alcohol, xylene, and mounted with a coverslip using permanent mounting media. The slides were examined with Nikon Eclipse E1000 microscope under a 40× objective.
RT-qPCR
Reverse transcription
Total RNA was isolated using an RNA isolation kit (Roche Applied Bioscience) and reverse transcribed with High-Capacity cDNA reverse transcription kit (Applied Biosystems) according to the manufacturer’s protocol. The resulting cDNA was diluted and used in SYBR green qPCR assay.
SYBR green qPCR assay
qPCR was performed using Power SYBR Green Mastermix (Applied Biosystems, Carlsbad, CA, USA) on an Applied Biosystems StepOne Plus Real-Time PCR System. All oligonucleotide primers were obtained from Integrated DNA Technologies (Coralville, IA). All assays were performed in triplicates and repeated twice, and results were plotted as average fold change relative to GAPDH. Primers used for validation of NPM1–TYK2 fusion transcripts in cell lines by SYBR green assay using the following pair of forward and reverse primers; 5′-ACTCAAAACCATCATCAACACCA-3′; 5′-GTTCCGGCCACACACATTAC-3′.
Clonogenic assay
The clonogenic assays were performed with Ba/F3 cells to determine the oncogenic potential of ΔN-1–257-NPM1–TYK2 in comparison to full-length NPM1–TYK2 along with vector control. Stable Ba/F3 cells were grown in Methocult media without IL-3 in triplicates. The cells were allowed to grow and form colonies for seven days. The colonies were counted and presented as percentages.
Cell viability
To compare the oncogenic mechanisms of the full-length NPM1–TYK2 and ∆N-1–257-NPM1–TYK2, resulting in the survival of Ba/F3 cells, we used trypan blue exclusion assays, and cell viability was represented by a percentage. Further, we conducted similar viability experiments with NPM1–TYK2–K462R kinase-dead mutant cells to validate the fusion kinase-driven transformation potential.
Animal experiments
Upon confirming in vitro transformation potential of the NPM1–TYK2 fusion gene, we next aimed to characterize the in vivo tumorigenic potential of a novel fusion gene utilizing an NPM1–TYK2-transformed Ba/F3-xenograft model. We used five to seven-week-old female Hsd:Athymic Nude-Foxn1nu mice (Envigo, Indianapolis, IN) to validate the tumorigenic potential of fusion kinase NPM1–TYK2. We injected 5 million Ba/F3-vector control and Ba/F3–FG–NPM1–TYK2 cells in the flank via a subcutaneous route (six mice in each group). Tumor size was measured by the Vernier caliper and tumor volumes were calculated using the modified ellipsoid formula: V = 1/2(length × width2). The mice were euthanized when the tumor size reached 2000 mm3 or the tumors became necrotic. Tumor tissues, spleen, and liver were collected from euthanized mice to perform Western blot analysis for signaling and hematoxylin and eosin (H&E) staining to evaluate lymphoma cell infiltration associated with hepatosplenomegaly.
Statistical analysis
Statistical analysis was performed using Microsoft Excel. Significant differences were determined using the Student’s t-test.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that all relevant data for this study are included within the paper. For any additional information regarding the supporting data, please contact the corresponding author with a reasonable request.
References
Davare, M. A. & Tognon, C. E. Detecting and targetting oncogenic fusion proteins in the genomic era. Biol. Cell 107, 111–129 (2015).
Mertens, F., Johansson, B., Fioretos, T. & Mitelman, F. The emerging complexity of gene fusions in cancer. Nat. Rev. Cancer 15, 371–381 (2015).
Lu, H. et al. Engineering and functional characterization of fusion genes identifies novel oncogenic drivers of cancer. Cancer Res. 77, 3502–3512 (2017).
Nussenzweig, A. & Nussenzweig, M. C. Origin of chromosomal translocations in lymphoid cancer. Cell 141, 27–38 (2010).
Dupain, C., Harttrampf, A. C., Urbinati, G., Geoerger, B. & Massaad-Massade, L. Relevance of fusion genes in pediatric cancers: toward precision medicine. Mol. Ther. Nucleic Acids 6, 315–326 (2017).
Malone, E. R., Oliva, M., Sabatini, P. J. B., Stockley, T. L. & Siu, L. L. Molecular profiling for precision cancer therapies. Genome Med. 12, 8 (2020).
Velusamy, T. et al. A novel recurrent NPM1-TYK2 gene fusion in cutaneous CD30-positive lymphoproliferative disorders. Blood 124, 3768–3771 (2014).
Prieto-Torres, L. et al. CD30-positive primary cutaneous lymphoproliferative disorders: molecular alterations and targeted therapies. Haematologica 104, 226–235 (2019).
Gu, X. et al. Leukemogenic nucleophosmin mutation disrupts the transcription factor hub that regulates granulomonocytic fates. J. Clin. Invest. 128, 4260–4279 (2018).
Brunetti, L., Gundry, M. C. & Goodell, M. A. New insights into the biology of acute myeloid leukemia with mutated NPM1. Int. J. Hematol. 110, 150–160 (2019).
Hubbard, S. R. Mechanistic insights into regulation of JAK2 tyrosine kinase. Front. Endocrinol. 8, 361 (2017).
Hammaren, H. M., Virtanen, A. T., Raivola, J. & Silvennoinen, O. The regulation of JAKs in cytokine signaling and its breakdown in disease. Cytokine 118, 48–63 (2019).
Kunchala, P., Kuravi, S., Jensen, R., McGuirk, J. & Balusu, R. When the good go bad: mutant NPM1 in acute myeloid leukemia. Blood Rev. 32, 167–183 (2018).
Falini, B. et al. Translocations and mutations involving the nucleophosmin (NPM1) gene in lymphomas and leukemias. Haematologica 92, 519–532 (2007).
Darracq, A. et al. NPM and NPM-MLF1 interact with chromatin remodeling complexes and influence their recruitment to specific genes. PLoS Genet. 15, e1008463 (2019).
Alvarez, A., Barisone, G. A. & Diaz, E. Focus formation: a cell-based assay to determine the oncogenic potential of a gene. J. Vis. Exp. 51742 (2014).
Cussac, D. et al. Nucleophosmin-anaplastic lymphoma kinase of anaplastic large-cell lymphoma recruits, activates, and uses pp60c-src to mediate its mitogenicity. Blood 103, 1464–1471 (2004).
Liu, C. B., Itoh, T., Arai, K. & Watanabe, S. Constitutive activation of JAK2 confers murine interleukin-3-independent survival and proliferation of BA/F3 cells. J. Biol. Chem. 274, 6342–6349 (1999).
Palmer, R. H., Vernersson, E., Grabbe, C. & Hallberg, B. Anaplastic lymphoma kinase: signalling in development and disease. Biochem. J. 420, 345–361 (2009).
Thaler, S. et al. Establishment of a mouse xenograft model for mycosis fungoides. Exp. Dermatol. 13, 406–412 (2004).
Sever, R. & Brugge, J. S. Signal transduction in cancer. Cold Spring Harb. Perspect. Med. 5, a006098 (2015).
Borcherding, D. C., He, K., Amin, N. V. & Hirbe, A. C. TYK2 in Cancer Metastases: Genomic and Proteomic Discovery. Cancers (Basel). 13, 4171 (2021).
Leitner, N. R., Witalisz-Siepracka, A., Strobl, B. & Muller, M. Tyrosine kinase 2—surveillant of tumours and bona fide oncogene. Cytokine 89, 209–218 (2017).
Charest, A. et al. Oncogenic targeting of an activated tyrosine kinase to the Golgi apparatus in a glioblastoma. Proc. Natl Acad. Sci. USA 100, 916–921 (2003).
Tuna, M., Amos, C. I. & Mills, G. B. Molecular mechanisms and pathobiology of oncogenic fusion transcripts in epithelial tumors. Oncotarget 10, 2095–2111 (2019).
Latysheva, N. S. & Babu, M. M. Discovering and understanding oncogenic gene fusions through data intensive computational approaches. Nucleic Acids Res. 44, 4487–4503 (2016).
Ma, H., Mallampati, S., An, G. & Wang, J. Targeted therapy in hematological malignancies: from basic research to clinical practice. Biomed. Res. Int. 2015, 157570 (2015).
Pikman, Y. & Stegmaier, K. Targeted therapy for fusion-driven high-risk acute leukemia. Blood 132, 1241–1247 (2018).
Seebacher, N. A., Stacy, A. E., Porter, G. M. & Merlot, A. M. Clinical development of targeted and immune based anti-cancer therapies. J. Exp. Clin. Cancer Res. 38, 156 (2019).
Prasad, A. et al. Identification of gene mutations and fusion genes in patients with Sezary syndrome. J. Invest. Dermatol. 136, 1490–1499 (2016).
Brunner, P. M., Jonak, C. & Knobler, R. Recent advances in understanding and managing cutaneous T-cell lymphomas. F1000Res 9, 331 (2020).
Itzhak, D. N., Tyanova, S., Cox, J. & Borner, G. H. Global, quantitative and dynamic mapping of protein subcellular localization. Elife 5, e16950 (2016).
Chen, Y. & Hu, J. Nucleophosmin1 (NPM1) abnormality in hematologic malignancies, and therapeutic targeting of mutant NPM1 in acute myeloid leukemia. Ther. Adv. Hematol. 11, 2040620719899818 (2020).
Feitelson, M. A. et al. Sustained proliferation in cancer: mechanisms and novel therapeutic targets. Semin. Cancer Biol. 35 Suppl, S25–S54 (2015).
Mori, S. et al. Anchorage-independent cell growth signature identifies tumors with metastatic potential. Oncogene 28, 2796–2805 (2009).
Sanchez-Garcia, I. & Grutz, G. Tumorigenic activity of the BCR-ABL oncogenes is mediated by BCL2. Proc. Natl Acad. Sci. USA 92, 5287–5291 (1995).
Ghoreschi, K., Laurence, A. & O’Shea, J. J. Janus kinases in immune cell signaling. Immunol. Rev. 228, 273–287 (2009).
Tran, N. V., Nguyen, L. T. A., Lim, K. W. & Phan, A. T. Potent and selective knockdown of tyrosine kinase 2 by antisense oligonucleotides. Immunohorizons 5, 70–80 (2021).
Sanda, T. et al. TYK2-STAT1-BCL2 pathway dependence in T-cell acute lymphoblastic leukemia. Cancer Discov. 3, 564–577 (2013).
Woss, K., Simonovic, N., Strobl, B., Macho-Maschler, S. & Muller, M. TYK2: an upstream kinase of STATs in cancer. Cancers (Basel) 11, 1728 (2019).
Jung, J. Human tumor xenograft models for preclinical assessment of anticancer drug development. Toxicol. Res. 30, 1–5 (2014).
Neel, D. S. et al. Differential subcellular localization regulates oncogenic signaling by ROS1 kinase fusion proteins. Cancer Res. 79, 546–556 (2019).
Lopez, D. J., Rodriguez, J. A. & Banuelos, S. Nucleophosmin, a multifunctional nucleolar organizer with a role in DNA repair. Biochim. Biophys. Acta Proteins Proteom. 1868, 140532 (2020).
Falini, B. et al. Altered nucleophosmin transport in acute myeloid leukaemia with mutated NPM1: molecular basis and clinical implications. Leukemia 23, 1731–1743 (2009).
Acknowledgements
R. Balusu acknowledges the Sosland Family Foundation Research Award, Hale Family Foundation, Frontiers Clinical and Translational Pilot Award UL1T, and Lied Preclinical Pilot Award. R. Jensen is a recipient of P30-CA168524 from NCI.
Author information
Authors and Affiliations
Contributions
S.K. and R.B. designed the experiments. S.K., C.J.V., D.R.W., and R.B. carried out experiments and generated data. S.K., R.W.B., M.U.M., T.L.L., I.S., C.J.V., A.V., S.A., S.G., W.C., D.R.W., R.A.J., Y.S., J.P.M., and R.B. analyzed and interpreted data. K.S.J.E.-J. provided reagents. R.A.J., J.P.M., and R.B. provided resources and acquired funding. S.K. and R.B. wrote the original paper. R.B. supervised the work. All authors reviewed and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kuravi, S., Baker, R.W., Mushtaq, M.U. et al. Functional characterization of NPM1–TYK2 fusion oncogene. npj Precis. Onc. 6, 3 (2022). https://doi.org/10.1038/s41698-021-00246-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-021-00246-4