Abstract
The fourth year of the COVID-19 pandemic without decreasing trends in the global numbers of new daily cases, high numbers of circulating SARS-CoV-2 variants and re-infections together with pessimistic predictions for the Omicron wave duration force studies about the endemic stage of the disease. The global trends were illustrated with the use the accumulated numbers of laboratory-confirmed COVID-19 cases and deaths, the percentages of fully vaccinated people and boosters (additional vaccinations), and the results of calculation of the effective reproduction number provided by Johns Hopkins University. A new modified SIR model with re-infections was proposed and analyzed. The estimated parameters of equilibrium show that the global numbers of new daily cases will range between 300 thousand and one million, daily deaths—between one and 3.3 thousand.
Similar content being viewed by others
Introduction
The fourth year of the COVID-19 pandemic, the high numbers of circulating SARS-CoV-2 variants1,2,3 and re-infected persons4,5,6, the lack of decreasing trends in the global numbers of new daily cases7,8, and the expected very long duration of the Omicron wave9 make us think about the constant circulation of the pathogen, that is, about the endemic stage of the disease. To illustrate the global trends, we will use the accumulated numbers of laboratory-confirmed COVID-19 cases Vj and deaths Dj, the percentages of fully vaccinated people VCj and boosters BCj, and the results of calculation of the effective reproduction number Rj10. Information corresponding to day tj (starting with January 22, 2020) is available in COVID-19 Data Repository by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU)8. We will use the version the JHU file updated on December 7, 2022 and the smoothed daily and annual characteristics11 (see formulae (8), (9) and Table 1):
The classical SIR-model12,13,14,15,16,17 relates the numbers of susceptible S(t), infectious I(t) and removed persons R(t) over time t and can be successfully used for simulations of the first COVID-19 waves in individual countries and worldwide11,18,19,20. Nevertheless, this model yields only trivial equilibrium point \(I_{**} = R_{**} = 0\) and cannot be used to estimate endemic characteristics. Numerous improvements of this model9,11,21,22,23,24 do not take into account re-infections. In this paper we will modify the classical SIR model by adding the re-infection terms, calculate a non-trivial equilibrium point, and estimate its stability and endemic characteristics of SARS-CoV-2 infection.
Results
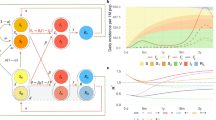
The solid lines in Fig. 1 illustrate the vaccination levels VCj and BCj (green and yellow colors, respectively), Rj values (magenta) and results of calculations of smoothed numbers of new daily cases DVi, deaths DDi, and the case fatality risk CFRi = DDi/DVi listed in Table 2 (blue, black and red, respectively). It can be seen that after October 2020, smoothed global numbers of new daily cases were never less than 300,000. After April 2020, the effective reproduction number10 was close to the critical value 1.025,26,27,28. In 2022 we see the a sufficient drop in CFI figures.
Global characteristics of the COVID-19 pandemic. Green, yellow and magenta lines respectively show VCj, BCj, Rj values listed in8. Solid blue, black and red represent calculations of DVi, DDi and CFRi, respectively (Eqs. (8) and (9)). Corresponding dashed lines represent the average values for different years listed in Table 1. The dotted blue line shows the results of non-linear correlation (Eq. 7).
To estimate the long term trends we calculated average values of DV, DD, CFI for every year of the pandemic with the use of accumulated characteristics on January 22 and December 31, 2020; January 1 and December 31, 2021; and January 1 and December 6, 2022. The results are listed in Table 1 and shown in Fig. 1 by dashed lines.
The DV values approximately doubled every year, the DD figure in 2021 was 1.8 times higher than in 2020. But in 2022 we see a sufficient decrease in mortality and much lower CFR values. This can be explained as a positive effect of vaccinations (the global VC values tend to some saturation in 2022, see green line), lower pathogenicity of the Omicron strain (which began to spread widely at the end of 202129,30), as well as the influence of natural immunity24.
To simulate the re-infections, we add terms \(\pm \delta R\) to the first and last differential equations of the classical SIR-model11,12,13,14,15,16,17, which relates the numbers of susceptible S(t), infectious I(t) and removed persons R(t) over time t:
Now compartment S(t), which includes the persons who are sensitive to the pathogen, is increasing with rate \(\delta R\) due to the persons who have lost their immunity and can be re-infected. The same term with opposite sign appears in Eq. (3).
Parameters \(\alpha\) (primal infection rate), \(\rho_{{}}\) (removal rate) and \(\delta\) (re-infection rate) can be supposed to be constant for every epidemic wave. Corresponding generalized SIR model, initial conditions and parameter identification procedures (at \(\delta\) = 0) were successfully used in9,11,31 to simulate and predict different waves of COVID-19 pandemic in different countries and worldwide.
The set of differential Eqs. (1)–(3) has the non-trivial equilibrium point32,33,34 \((S_{*} ,I_{*} ,R_{*} )\) corresponding to zero values of the derivatives:
At the equilibrium, the daily number of new cases DV* must be equal to the daily number of new infections; to the sum of immunized, isolated and dead persons; and to the number of persons who have lost their immunity and moved from compartment R to S, i.e.:
The Jacobian matrix32,33,34 of set (1)–(3) can be written as follows:
Taking into account (4), the eigenvalues of (6) at the equilibrium point can be calculated as follows:
Eigenvalues \(\lambda_{2,3}\) have negative real parts, but the presence of a zero eigenvalue \(\lambda_{1}\) indicates that the system can approach no equilibrium with increasing time. Its asymptotic stability needs further investigations with the use of non-linear methods.
To estimate the endemic global number of new daily cases we can use the average DV value for 2022 (around 1 million), presented in Table 1. More optimistic prediction \(DV \approx 300,000\) follows from the minimum of DVi values illustrated by the blue solid line and non-linear correlation approach35, which yields the best fitting curve
for the 2022 dataset (shown by the dotted blue line). Thus, the global daily numbers of SARS-CoV-2 cases can vary from 300 thousand to one million and the daily numbers of deaths from 1.0 to 3.3 thousand (if we use the CFR value from the Table 1 corresponding to 2022). According to the recent WHO report “nearly 2.8 million new cases and over 13,000 deaths were reported in the week of 9–15 January 2023. In the last 28 days (19 December 2022–15 January 2023), nearly 13 million cases and almost 53,000 new deaths were reported globally”7. Calculating the average daily characteristics, we obtain approximately 400,000–464,000 new daily cases and 1857–1893 daily deaths. These figures are consistent with the given theoretical estimates of SARS-CoV-2 endemic characteristics.
Discussion
It is necessary to emphasize that the presented estimates refer to the laboratory confirmed cases only. Due to the high numbers of asymptomatic patients and insufficient testing level, the real number of SARS-CoV-2 cases is much higher35,36,37. To estimate the global visibility coefficient let us take the number of cases per million 72,524 registered worldwide as of August 1, 20228. On the same day, the average value was 460,834 for European countries with high enough testing level (more than 3 tests per capita)35. It means that the real numbers of cases can be approximately 6 times higher than registered figures. In the future, reducing the testing level can lead to lower numbers of detected cases and deaths.
Other restrictions on the accuracy of the given estimates may be related to the situation in mainland China, where the Zero-COVID policy38 began to be lifted only after November 30, 202239. Very high numbers of new daily cases in December 2022 reported by the media40 do not look reliable41, nevertheless it is better to use the presented estimations excluding the Chinese datasets.
According to the presented estimations, the annual mortality caused by SARS-CoV-2 will range from 365 thousand to 1.2 million. Unfortunately, these figures are higher than in the case of seasonal influenza (between 294 and 518 thousand annual deaths in the period from 2002 to 201142).
Conclusions
Thus, the proposed modified SIR model with re-infections allowed us to make adequate estimations of the equilibrium (endemic) global daily numbers of SARS-CoV-2 cases and related deaths. In different countries and regions, these characteristics are different and need special investigations. To take into account the influence of other factors (e.g., vaccination and/or testing rates), more complicated mathematical models can be used.
Methods
Since numbers of new cases and deaths are random and characterized by some weekly periodicity, the use of the smoothed characteristics is recommended11:
Some different smoothing procedure is used in8 based on the values corresponding to the fixed and previous 6 days. To estimate the smoothed numbers of new daily cases DVi, deaths DDi, and the case fatality risk CFRi =DDi/DVi, the numerical derivatives of the smoothed values (8) are calculated as follows11:
The results of calculations are listed in Table 2.
Human/animals involved
No humans or human data was used during this study.
Data availability
All data generated or analysed during this study are included in this published article.
References
https://www.who.int/activities/tracking-SARS-CoV-2-variants.
https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-classifications.html.
https://coronavirus.health.ny.gov/covid-19-reinfection-data.
Guedes, A. R. et al. Reinfection rate in a cohort of healthcare workers over 2 years of the COVID-19 pandemic. Sci. Rep. 13, 712. https://doi.org/10.1038/s41598-022-25908-6 (2023).
Flacco, M. E. et al. Risk of SARS-CoV-2 reinfection 18 months after primary infection: Population-level observational study front public health, 02 May 2022. Sec. Infect. Dis. Epidemiol. Prevent. https://doi.org/10.3389/fpubh.2022.884121 (2022).
World Health Organization. Coronavirus disease (COVID-2019) situation reports. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/. Retrieved Mar 14 2020.
COVID-19 Data Repository by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU). https://github.com/owid/covid-19-data/tree/master/public/data.
Nesteruk, I. Epidemic waves caused by SARS-CoV-2 omicron (B.1.1.529) and pessimistic forecasts of the COVID-19 pandemic duration. MedComm 3, 1. https://doi.org/10.1002/mco2.122 (2022).
Arroyo-Marioli, F., Bullano, F., Kucinskas, S. & Rondón-Moreno, C. Tracking R of COVID-19: A new real-time estimation using the Kalman filter. PLoS One 16(1), e0244474. https://doi.org/10.1371/journal.pone.0244474 (2021).
Nesteruk, I. COVID-19 pandemic dynamics. Springer Nat. https://doi.org/10.1007/978-981-33-6416-5 (2021).
Kermack, W. O. & McKendrick, A. G. A Contribution to the mathematical theory of epidemics. J. R. Stat. Soc. Ser. A 115, 700–721 (1927).
Weiss, H. The SIR model and the foundations of public health. MatMat 3, 1–17 (2013).
Daley, D. J. & Gani, J. Epidemic Modeling: An Introduction (Cambridge University Press, 2005).
Hethcote, H. W. The mathematics of infectious diseases. SIAM Rev. 42(4), 599–653 (2000).
Brauer, F. & Castillo-Chávez, C. Mathematical Models in Population Biology and Epidemiology (Springer, 2001).
Huppert, A. & Katriel, G. Mathematical modelling and prediction in infectious disease epidemiology. Clin. Microbiol. Infect. 19(11), 999–1005. https://doi.org/10.1111/1469-0691.12308 (2013).
Alireza, M., Ievgen, M., Kseniia, B., Sergey, Y. & Dmytro, C. Comparative study of linear regression and SIR models of COVID-19 propagation in Ukraine before vaccination. Radioelectron. Comput. Syst. 3, 5–18. https://doi.org/10.32620/reks.2021.3.01 (2021).
José, E. A., Jérémie, D. & José, N. O. Global analysis of the COVID-19 pandemic using simple epidemiological models. Appl. Math. Model. 90, 995–1008. https://doi.org/10.1016/j.apm.2020.10.019 (2021).
de Andres, P. L., de Andres-Bragado, L. & Hoessly, L. Monitoring and forecasting COVID-19: Heuristic regression, susceptible-infected-removed model and spatial stochastic. Front. Appl. Math. Stat. https://doi.org/10.3389/fams.2021.650716 (2021).
Nakamura, G. M., Cardoso, G. C. & Martinez, A. S. Improved susceptible–infectious–susceptible epidemic equations based on uncertainties and autocorrelation functions. R. Soc. Open Sci. 7(2), 191504 (2020).
Britton, N. F. Essential Mathematical Biology 352 (Springer, 2004).
Mustafa, T. An extended epidemic model with vaccination: Weak-immune SIRVI. Phys. A 598, 127429. https://doi.org/10.1016/j.physa.2022.127429 (2022).
Nesteruk, I. Influence of possible natural and artificial collective immunity on new COVID-19 pandemic waves in Ukraine and Israel. Explor. Res Hypothesis Med. https://doi.org/10.14218/ERHM.2021.00044 (2021).
https://www.r-bloggers.com/2020/04/effective-reproduction-number-estimation/
van der Heiden, M. & Hamouda, O. Schätzung Der Aktuel-Len Entwicklung Der Sars-Cov-2-Epidemie in Deutsch-land—nowcasting. Epid. Bull. 17, 10–15. https://doi.org/10.25646/669 (2020).
Cori, A., Ferguson, N. M., Fraser, C. & Cauchemez, S. A new framework and software to estimate time-varying reproduction numbers during epidemics. Am. J. Epidemiol. 178(9), 1505–1512. https://doi.org/10.1093/aje/kwt133 (2013).
Lorenzo-Redondo, R., Ozer, E. A. & Hultquist, J. F. Covid-19: Is omicron less lethal than delta?. BMJ 378, 1806. https://doi.org/10.1136/bmj.o1806 (2022).
Sigal, A., Milo, R. & Jassat, W. Estimating disease severity of Omicron and Delta SARS-CoV-2 infections. Nat. Rev. Immunol. 22, 267–269. https://doi.org/10.1038/s41577-022-00720-5 (2022).
Nesteruk, I. Improvement of the software for modeling the dynamics of epidemics and developing a user-friendly interface. Infect. Dis. Model. 8(3), 806–821. https://doi.org/10.1016/j.idm.2023.06.003 (2023).
Anthony, N. M., Ling, H. & Derong, L. Stability of Dynamical Systems (Birkhäuser, 2008). https://doi.org/10.1007/978-0-8176-4649-3.
Bohmer, C. G., Harko, T. & Sabau, S. V. Jacobi stability analysis of dynamical systems—applications in gravitation and cosmology. Adv. Theor. Math. Phys. 16, 1145–1196 (2012).
Nesteruk, I. & Rodionov, O. The COVID-19 pandemic in rich and poor countries. Res. Square https://doi.org/10.21203/rs.3.rs-2348206/v1 (2022).
Ryan, M. B. et al. Estimating global, regional, and national daily and cumulative infections with SARS-CoV-2 through Nov 14, 2021: A statistical analysis. Lancet 399(10344), 2351–2380. https://doi.org/10.1016/S0140-6736(22)00484-6 (2022).
Lazarus, J. V. et al. A multinational Delphi consensus to end the COVID-19 public health threat. Nature https://doi.org/10.1038/s41586-022-05398-2 (2022).
https://news.cctv.com/2022/12/03/ARTIbCeNratl6uA4kXhzEdBG221203.shtml.
Bloomberg. Internet information. https://www.bloomberg.com/news/articles/2022-12-23/china-estimates-covid-surge-is-infecting-37-million-people-a-day. Retrieved 24 Jan 2023.
Nesteruk, I. What is wrong with Chinese COVID-19 statistics?. Epidemiol. Biostat. Public Health 18(1), 9–11. https://doi.org/10.54103/2282-0930/20637 (2023).
Paget, J. et al. Global mortality associated with seasonal influenza epidemics: New burden estimates and predictors from the GLaMOR Project. J. Glob. Health 9(2), 020421. https://doi.org/10.7189/jogh.09.020421 (2019).
Acknowledgements
The author is grateful to Tetsuro Kobayashi and Oleksii Rodionov for their help in providing very useful information.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nesteruk, I. Endemic characteristics of SARS-CoV-2 infection. Sci Rep 13, 14841 (2023). https://doi.org/10.1038/s41598-023-41841-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-41841-8
This article is cited by
-
Intranasal delivery of PEA-producing Lactobacillus paracasei F19 alleviates SARS-CoV-2 spike protein-induced lung injury in mice
Translational Medicine Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.