Abstract
There has been a growing attention on using machine learning (ML) in pharmacovigilance. This study aimed to investigate the utility of supervised ML algorithms on timely detection of safety signals in the Korea Adverse Event Reporting System (KAERS), using infliximab as a case drug, between 2009 and 2018. Input data set for ML training was constructed based on the drug label information and spontaneous reports in the KAERS. Gold standard dataset containing known AEs was randomly divided into the training and test sets. Two supervised ML algorithms (gradient boosting machine [GBM], random forest [RF]) were fitted with hyperparameters tuned on the training set by using a fivefold validation. Then, we stratified the KAERS data by calendar year to create 10 cumulative yearly datasets, in which ML algorithms were applied to detect five pre-specified AEs of infliximab identified during post-marketing surveillance. Four AEs were detected by both GBM and RF in the first year they appeared in the KAERS and earlier than they were updated in the drug label of infliximab. We further applied our models to data retrieved from the US Food and Drug Administration Adverse Event Reporting System repository and found that they outperformed existing disproportionality methods. Both GBM and RF demonstrated reliable performance in detecting early safety signals and showed promise for applying such approaches to pharmacovigilance.
Similar content being viewed by others
Introduction
Post-marketing surveillance studies are essential in ensuring drug safety through a periodic monitoring of the potential adverse events (AEs) that were not identified during clinical trials1,2. Routine drug safety monitoring has traditionally been based on a spontaneous reporting system (SRS) by applying statistical data mining tools to promptly identify a safety signal, that is an AE related to a specific drug that requires further investigation on a causal relationship3,4. This in turn enables the health authorities to take timely actions to mitigate further safety risks through a regulatory action5,6.
Many data mining methods are available to detect safety signals in SRS, including frequentist-based methods (i.e., proportional reporting ratio [PRR] and reporting odds ratio [ROR]) and Bayesian approaches (i.e., gamma Poison shrinkage model [GPS] and information component [IC] of Bayesian confidence propagation neural network [BCPNN])7,8,9. Performances of these methods have been explored previously, with the latter generally demonstrating higher sensitivity and specificity10. Recently, there has been a growing attention on the application of machine learning (ML) algorithms in pharmacovigilance11. Among studies that explored ML-based AE prediction, one study applied an ensemble ML algorithm trained with the AE profiles extracted from the US Food and Drug administration (FDA) Adverse Event Reporting System (FAERS) to identify 18 of 23 AEs identified during post-marketing surveillance12. Another study utilized information on the known AEs and drug indications extracted from the Side Effect Resource (SIDER) database to train ML algorithms, and successfully predicted AEs responsible for a drug withdrawal13. Besides AE prediction, few studies also have utilized ML algorithms to identify safety signals in the SRSs14,15. Preliminary evidence has shown that ML algorithms were not superior over frequentist-based methods and Bayesian approaches in detecting safety signals; however, this may be attributed to sparseness of certain combination of drug and AE leading to poor representativeness of ML training datasets, which may negatively impact the performance of ML algorithms16.
While the value of the data mining methods has been well established in the SRS databases, relatively little is known on the performance of ML algorithms in post-marketing surveillance. Indeed, ML algorithms had been utilized in predicting potential drug-AE association with the processing of natural languages from social media or health-related forums17,18; however, these approaches were laborious, requiring annotations of the lay languages by qualified medical experts11. Contrastingly, information in the SRS databases is based on standardized international medical terminology or classification systems, thereby requiring less human input for constructing training datasets.
The Korea Adverse Event Reporting System (KAERS) database was established in 1988 and managed by the Korea Institute of Drug Safety and Risk Management (KIDS)19. Similar to FAERS, it contains spontaneous reports of suspected drug and AE (s) reported by the healthcare professionals, consumers, and pharmaceutical companies. All AE reports are processed and stored as an individual case safety report that are periodically submitted to regulatory bodies (i.e., Ministry of Food and Drug Safety, World Health Organization (WHO) - Uppsala Monitoring Centre) or provided for research use. Many efforts have been made to detect safety signals in the KAERS, applying a variety of data mining methods including frequentist-based methods20,21,22, Bayesian approaches and tree-based scan statistics23,24.
Given the growing interest, this study aimed to investigate the utility of ML algorithms on early detection of AEs in the KAERS. Our recent pilot study successfully identified new safety signals for two anti-neoplastic drugs, nivolumab and docetaxel, using ML algorithms15. Here, we expand from our previous work to explore the supervised ML-based early AE prediction, using KAERS data and drug label information as the feature and label data for ML training, respectively. Two ensemble ML algorithms, gradient boosting machine (GBM) and random forest (RF), previously demonstrated to be the most effective in classifying safety signals using real-world data14, were applied to detect early safety signal, defined as an AE detected prior to it being updated in the drug label information. Specifically, we selected 5 AEs identified during post-marketing surveillance of a case drug, infliximab, an TNF-alpha inhibitor commonly prescribed for rheumatoid arthritis.
Results
Characteristics of the AE reports
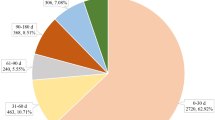
Of 11,376 AE reports, 4482 included infliximab as the suspected drug. Gender was relatively evenly distributed (44.7%, 43.0% for men and women, respectively), and 30.9% of the reports were from age between 30 and 49 years. The report volume increased gradually throughout the study period, with a notable peak of 35.9% in 2016, and 28.0% of the reported AEs were categorized as serious AE. The majority were reported through post-marketing surveillance (60.6%), by manufacturer (83.3%), and from physician (74.8%), implying that these were filed from physician to manufacturer and subsequently got submitted as post-marketing surveillance reports to the KAERS (Table 1).
Primary analysis: early safety signal detection
Characteristics of the 5 pre-specified AEs of infliximab (i.e., agranulocytosis, cervical cancer, cerebrovascular accidents, leukemia, and transient visual loss) are described in Table S1. All AEs were reported by manufacturers and recorded as a serious AE except for transient visual loss.
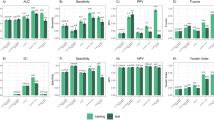
Of the 5 AEs assessed, RF and GBM identified 4 early signals, whereas adjusted ROR and IC did not identify any signal prior to the AEs being updated in the label information of infliximab (Table 2). The 4 early signals identified by RF and GBM were detected in the first year they were reported with infliximab in the KAERS (Fig. 1).
-
Agranulocytosis: WHO-Adverse Reaction Terminology (WHO-ART) Preferred Term “agranulocytosis” first appeared in the cumulative yearly dataset of 2009–2011. Both RF and GBM continuously identified this signal up to 2015, and the AE was updated in the label information of infliximab in 2017.
-
Cervical cancer: WHO-ART Preferred Term “cervical carcinoma” first appeared in the cumulative yearly dataset of 2009–2013. Both RF and GBM continuously identified this signal up until it was updated in the label information of infliximab in 2017.
-
Cerebrovascular accidents: WHO-ART Preferred Terms “cerebellar infarction” and “cerebral infarction” first appeared in the cumulative yearly dataset of 2009–2017. Both RF and GBM continuously identified this signal up until it was updated in the label information of infliximab, and the standardized difference values of GBM remained higher than that of RF throughout the early signal detection period.
-
Leukemia: WHO-ART Preferred Terms “leukemia acute” and “leukemia granulocytic” first appeared in the cumulative yearly dataset of 2009–2015. Both RF and GBM identified this signal in the first year it appeared in the KAERS.
-
Transient visual loss: WHO-ART Preferred Term “vision abnormal” first appeared in the cumulative yearly dataset of 2009–2015. Neither ML algorithms nor conventional data mining methods identified this AE throughout the data period.
Performance of the data mining methods
We measured the performance by comparing the signals detected by each data mining method with the reference standard. GBM demonstrated the best balance between sensitivity (79%) and specificity (79%), followed by RF with sensitivity and specificity of 60% and 91%, respectively. While the ROR and IC had higher specificities (ROR025: 99%; IC025: 95%), their sensitivities (ROR025: 18%; IC025: 21%) were considerably lower than that of ML algorithms. Performance measures expressed by area under receiver operating characteristics curves (AUROC) for the data mining methods are shown in Fig. 2.
Receiver operating characteristic (ROC) curve illustrating the prediction performances of data mining methods used to detect safety signals of infliximab in KAERS (2009–2018) and FAERS (2014–2018). Abbreviation: KAERS, Korea adverse event reporting system; FAERS, FDA adverse event reporting system; AUROC, area under receiver operating characteristic curve; GBM, gradient boosting machine, RF, random forest; ROR, Reporting odds ratio; IC, Information component, Prob, probability. *ROR025 is the lower limit of a 95% confidence interval for estimated reporting odds ratio. †IC025 is the lower limit of a 95% confidence interval for estimated information component.
Secondary analysis: identifying new safety signals of infliximab
Among the unknown 148 AEs of infliximab in the KAERS, 27 safety signals were detected by both GBM and RF, whereas only 2 and 3 were detected by ROR025 and IC025, respectively (Table 3 and Table S2).
Sensitivity analysis: validating the generalizability of ML algorithms
We further applied our models to data retrieved from the FAERS from 3rd quarter of 2014 (Q3 2014) to 4th quarter of 2018 (Q4 2018) and consistently found superior performance of ML algorithms. GBM consistently showed the best performance with sensitivity of 64%, specificity of 83% and overall AUROC of 75% (Fig. 2). RF achieved the second highest overall AUROC (73%) with sensitivity of 64% and specificity of 76%, respectively (Fig. 2). Notwithstanding the relatively lower performance measures in FAERS than in KAERS, ML algorithms consistently performed better than ROR025 and IC025.
Discussion
We applied supervised ML algorithms, GBM and RF, to explore their utility in early signal detection of post-marketing safety signals associated with infliximab in the SRSs. Among the 5 AEs with confirmed causality association to infliximab and added to the drug label information post-approval, GBM and RF identified 4 safety signals in the very first year these AEs appeared in the KAERS, whereas ROR and IC did not generate any early signals. According to the results of this study, ML algorithms performed better than the methods currently used by the regulatory agencies in South Korea (AUROC of 0.82 and 0.79 for GBM and RF, respectively, vs. 0.59 and 0.58 for ROR and IC, respectively). However, applying the same algorithms to FAERS data over a shorter time period of the infliximab lifecycle did not demonstrate the same performance, but they both outperformed ROR and IC respectively. The above findings show promise for applying such approaches to pharmacovigilance.
Our study successfully applied the ML algorithms in two SRSs, focusing on their ability to generate early signals. Both RF and GBM detected 4 out of 5 pre-specified AEs of infliximab as early as the very first year they were reported in the KAERS. However, our study also has some limitations. First, passive surveillance data are inherently subject to potential selection bias and underestimation of AE reports. Another potential bias may arise from duplication of the reports. Case duplicates, defined as “two or more reports describing the same occurrence of one or more AEs for the same patient that are assigned different case numbers instead of being linked as the same case”25, may potentially alter data mining calculations. However, our limited access to the SRS did not allow for deduplication of the data in hand. Second, potential misclassification of infliximab-AE pairs as either positive or negative control for reference standard construction in our study may had impact on the performance of the ML algorithms. Indeed, quantitative performance of ML algorithm is largely dependent on a quality of reference standard, and misclassification of the negative controls may certainly bias the performance measures in both directions26. However, we used the standard approach presented by the Observational Medical Outcomes Partnership (OMOP) researchers to reduce potential for such misclassification27. Third, the quantity and quality of an input data have major influences on performance of the ML algorithms and can lead to inconsistencies across different data sources14. In the sensitivity analysis for evaluating the generalizability of our findings, ML algorithms could not utilize all features available in the KAERS due to some differences in data characteristics between KAERS and FAERS; for example, information on the report type could not be identified in the FAERS database. Besides, information on an active ingredient of pharmaceutical product was not provided in the FAERS before Q3 2014, which led to the shorter data period for sensitivity analysis, compared with the main analysis. Therefore, the number of label positive and negative derived from FAERS was smaller than that derived from KAERS; Utilizing less samples and features might have resulted in achieving relatively lower performance of ML algorithms in the FAERS. Fourth, safety signals identified from the unknown AEs do not necessarily indicate causal relationship. These signals need to be prioritized by rarity or seriousness and require further investigation by the medical experts.
ML algorithm’s performance is largely determined by the quantity of the training dataset. Sparseness of certain combination of drug and AE due to relatively small report volume size in KAERS, compared with other large-scale SRS such as FAERS or VigiBase of WHO, may have limited the performance of the ML algorithms in our study. For instance, both GBM and RF failed to predict “transient visual loss” as early safety signal partly due to relatively low quantity of reports that included WHO-ART Preferred Term “vision abnormal”. Moreover, apart from low report count, none of the reports that listed “vision abnormal” were recorded as serious AE (Fig. S1). Given that serious AE designation in the KAERS was included in constructing covariate feature, it was likely that the ML algorithms yielded low signaling probability for this particular AE. Nonetheless, both RF and GBM successfully predicted 4 out of 5 AEs, promising as a new approach for early prediction of clinically significant AEs.
Expectedly, ML algorithms outperformed ROR and IC in predicting AEs as well as identifying new signals for infliximab. Such difference may be attributed to the statistical aspect of each method in calculating probability value for a drug-AE pair. Conventional data mining methods such as PRR, ROR, and IC simply calculate signal scores based on 2 × 2 contingency table28. While they are relatively simple and fast to compute, their performance varies across different thresholds and limited by high rate of false positives29,30. On the other hand, ML algorithms utilize a large number of features to calculate signal scores, and their performance is determined by the quality of input dataset. For instance, Schotland et al. used feature data generated by aggregating AEs from the FAERS and drug label information, sequentially, to construct an ensemble ML method, and tested whether unlabeled AEs at the time of drug approval could be predicted. The ML performance improved with increasing volume of input data, from precision, recall and specificity of 0.57, 0.78 and 0.61 with FAERS data to 0.67, 0.81 and 0.71 with addition of drug label information to the feature data, respectively12.
In the recent years, ML-based pharmacovigilance has been extended to studies that predict unlabeled AEs for a new drug at the time of approval by using the pharmacological target adverse event (TAE) profiles based on comparator drugs. An ensemble ML algorithm based on the data from drug label information, literatures and FAERS demonstrated reliable performance in predicting potential AEs of a new drug (AUC of 0.87)31. Besides ML-based AE prediction, they have been applied in routine safety surveillance for generating potential safety signals for evaluation14,15, as well as predicting unknown drug-drug interactions32. Our study findings complement the recently growing evidence on potential application of the ML algorithms, focusing on their ability to facilitate the timeliness of safety signal detection that would in turn reduce patient harm and improve health outcomes.
Of two ML algorithms, RF demonstrated optimal performance with better balance between sensitivity (74%) and specificity (89%), compared with 57% and 95% for GBM. Few studies have also assessed performances of these algorithms. One study based on Australian medication dispensing data showed GBM out-performed RF with sensitivities of 77% and 57%, respectively14. Another study compared performances of several data mining methods, including three ML algorithms, in which IC demonstrated the highest AUC of 0.69 and RF with the lowest AUC of 0.5216. Such discordance across different studies may attribute to the differential data volume, data characteristics, and lack of gold standard. Specifically, in context of differential data characteristics, there are currently no standardized guideline on mapping AEs listed in a drug label with the AE coding system used in each data source (i.e., WHO-ART used to code AE in the KAERS).
In conclusion, our study showed that the ML algorithms performed well in early detecting unknown AEs associated with infliximab. Before discussing the potential routine use of these methods in pharmacovigilance, additional efforts are needed to improve the consistency of the ML algorithms’ performance with other standardized drug-AE reference data. It is also vital to determine the acceptable performance levels in collaboration with the Korean and other regulatory agencies worldwide.
Materials and methods
Data source
Our study data source included AE reports in the KAERS between 2009 and 2018. Each report contains information on the demographic characteristics, administered drug (s), and suspected AE (s). Administered drug (s) are either labeled as “suspected drug”, “concomitant drug”, or “drug-drug interaction”. Each report may include one or more AEs, and classified as “serious AE” if they resulted in one or more of the following conditions: persistent or significant disability, congenital anomaly, life-threatening, death, hospitalization, prolonged hospitalized days, or other unspecified clinical intervention. Moreover, AEs reported by healthcare professionals or consumers are classified as “spontaneous”, pharmaceutical companies as “post-marketing surveillance” and monthly literature search for published case series or reports conducted monthly by KIDS as “literature”. AEs are coded according to the World Health Organization Adverse Reaction Terminology 092 (WHO-ART) and drugs according to the WHO’s Anatomical Therapeutic Chemical (ATC) classifications.
Study scheme
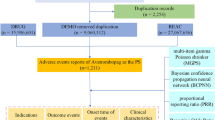
We used the ML algorithms and conventional data mining methods to detect safety signals for infliximab in the KAERS between 2009 and 2018. All AE reports that included two commonly prescribed disease-modifying antirheumatic drugs (i.e., infliximab and methotrexate) were analyzed to identify safety signals for infliximab. We first extracted all AE reports that contained immunosuppressants (ATC: L04) from the KAERS, and then created an initial dataset that listed infliximab or methotrexate as a “suspected drug”. For follow-up report of an initial report, only the latest reports were included. The initial dataset was segmented based on the reporting year, and 10 cumulative yearly datasets were created by merging each subsequent year data to the 2009 data (Fig. 3). Then, retrospective data screening was conducted from 2009 dataset to 2009–2018 dataset to evaluate whether the five pre-specified AEs of infliximab were detected prior to they were updated in the drug label information. After evaluating performance of each data mining method, we applied them to detect safety signals from unknown AEs in the KAERS that are neither listed in the drug label information of infliximab and other drugs belonging to same therapeutic class (i.e., TNF-alpha inhibitor).
Generating input dataset for machine learning
We first constructed an input dataset required for ML algorithm to generate signaling probability of an AE. The input dataset contains information on the known and unknown AEs of a drug (labeled data) and quantifiable properties and characteristics of AEs (feature data).
Labeled data
Labeled data contains information on the known and unknown AEs of the study drugs. Known AEs, referred as reference standard, are retrieved from the drug label information and used to orient for training and testing of the ML algorithms. Unknown AEs are the ones that may be related the reported drug (s), for which causality relationship cannot be ruled out; this data is used by ML algorithms to detect new safety signals of the study drug.
To construct a labeled data, we first created a reference standard by extracting safety data of the study drug (i.e., infliximab) as well as other drugs belonging to the same therapeutic class (i.e., etanercept, adalimumab and golimumab) in the US FDA-approved drug label information; The labels (revised versions between 2009 and 2018) were obtained from the Labeling Archives of the National Library of Medicine DailyMed website33. Using the reference standard, AEs identified in the initial dataset were classified into the following categories: label-positive (Y), label-negative (N), and unknown AE (U). AEs labeled “Y” were the known AEs of the study drugs, “N” the ones not listed in both labeling information of the study drugs and other drugs belonging to same therapeutic class, and “U” the rest of the AEs in the initial dataset. The “Y” and “N” constituted the reference standard, and “U” the unknown AEs of the study drugs in the labeled data, respectively14,34,35. The numbers of AEs labeled “Y”, “N”, and “U” are presented in Table S2. Reference standard was used as an orientation for training and testing the ML algorithms.
Feature data
Supervised ML algorithms generate a signaling probability for each AE based on a feature data. We considered two aspects for selecting information to be included as the features: (1) minimizing potential bias due to missing data, and (2) enhancing the applicability of our methodology to other SRS databases. To take these into account, we used commonly required information for a valid individual case safety report (ICSR) described in “Guideline on good pharmacovigilance practice Module VI”: An identifiable patient, an identifiable reporter, a suspect drug, and an adverse event36. We generated a total of 35 features that included statistical, organ-specific, and covariate features. The statistical feature included the number of reports for a specific AE associated with infliximab from a 2 by 2 contingency table commonly used for disproportionality-based signal detection.
We also included the organ-specific features because typical AE is manifested or closely related to a specific organ, organ system, or mechanism of action. The system organ classes (SOCs) within the WHO-ART represents the category by which a specific AE is distinguished based on etiology, manifestation site, or purpose. A total of 32 SOCs were included as the organ-specific features, and 3 to 4 digits numeric codes (i.e., 100, 200…3200) of the SOCs were assigned a number from 1 to 32 to minimize the potential negative impact that the large numerical value might have on ML algorithms’ performance.
Covariate features included confounding factors, such as gender, age, report type, and report source by occupation and affiliation, which are commonly considered in the post-marketing safety surveillance studies37,38. Age was categorized into seven groups (0–20, 21–29, 30–39, 40–49, 50–59, 60–69, and older), consistent with a previous study on the safety signals of infliximab39. Variables for report type included spontaneous report, post-marketing surveillance study report, and case reports from literature. There were five (physician, pharmacist, nurse, other health professional, and consumer) and three (regional pharmacovigilance center, medical institution, and drug manufacturer) categories for report source by occupation and affiliation, respectively. Relative importance of features in signal detection using the ML algorithms are provided in Fig. S1.
Primary analysis: early safety signal detection
We explored the performance of ML algorithms in detecting early safety signals through retrospective screening of the 10 cumulative yearly datasets. The algorithms were developed using each cumulative dataset to detect safety signals; for example, 2009–2012 data included AE cases reported between 2009 and 2012, with each AE classified as “Y”, “N” or “U” according to the reference standard constructed based on the drug labels updated in 2012. Then, AEs labeled “Y’ and ‘N’ in the 2009–2012 dataset were used to train and evaluate ML algorithms, and data mining methods were used to identify safety signals among the AEs labeled “U”.
Early signal was ascertained by whether it was detected prior to being updated in the drug label information. The five AEs selected a priori were the ones updated post-approval of infliximab through post-marketing surveillance after 2009 and described in details below:
-
Transient visual loss: This term was listed on the infliximab label revised in 2010. WHO-ART Preferred Term “vision abnormal” was considered to be an equivalent term describing this AE.
-
Agranulocytosis: This term was listed on the label revised in 2017. WHO-ART Preferred Term “agranulocytosis” was considered to be an equivalent term describing this AE.
-
Cervical cancer: This term was listed on the label revised in 2017. WHO-ART Preferred Term “cervical carcinoma” was considered to be an equivalent term describing this AE.
-
Cerebrovascular accidents: This term was listed on the label revised in 2017. WHO-ART Preferred Terms “cerebellar infarction” and “cerebral infarction” were considered to be an equivalent term describing this AE.
-
Leukemia: This term was listed on the label revised in 2018. WHO-ART Preferred Terms “leukemia acute” and “leukemia granulocytic” were considered to be an equivalent term describing this AE.
ML algorithms for safety signal detection
Among the available ML algorithms, we chose two commonly used supervised classifiers, RF and GBM, which have been shown to demonstrate high specificity and positive predictive value in detecting AEs from large-scale medication dispensing data of Australia14.
-
RF: The RF classifier is an ensemble model that combines the multiple decision tree classifiers of AEs. Many different subsets of a training dataset with randomly sampled features are used to train RF classifier. Therefore, it averages multiple deep decision trees which are trained on different parts of the same training dataset and eventually aims to reduce the variance and control over-fitting. RF had better predictive accuracy and performance in detecting signals than other supervised ML algorithms such as Support Vector Machine or Logistic Regression14.
-
GBM: The GBM classifier is another ensemble classifier that combines many weak classifiers to generate stronger model. GBM begins by learning a base weak classifier and then generates the next model that reduces predictive errors of the precedent classifier. A stronger machine is eventually made as a result of the repeated process of developing the next model to minimize error.
Each algorithm produces an output value that ranges from 0 to 1, representing a probability of association between infliximab and an AE. When calculating signal detection performance using a reference standard, the signaling threshold is defined as a probability value that has the highest area under receiver operating characteristic curve (AUROC). AUROC is defined as:
Training ML algorithms
From the input data, we constructed gold standard dataset that contained label-positive (i.e., “Y”) and -negative AEs (i.e., “N”). Gold standard dataset was randomly divided into the training (75%) and test sets (25%), adjusting for imbalances in the distribution of the label-positive and -negative AEs with the Synthetic Minority Over-sampling Technique (SMOTE). We fitted RF and GBM with hyperparameters tuned on the training set by using a fivefold stratified cross-validation. Hyperparameter tuning is a strategy to optimize ML’s performance by identifying the optimal values of adjustable parameters in training process, such as the number of nodes in a decision tree to be made that affects the performance of decision tree. Five-fold stratified cross-validation is a resampling technique to evaluate ML algorithm on a finite sample by dividing data into five subsets with the same proportion of labels in the reference standard. The first fold was used as test set and the remaining as training sets, and repeated until all 5 folds were used as the test set.
Conventional data mining methods for safety signal detection
Adjusted ROR: Signal threshold of lower bound of 95% CI for adjusted ROR (ROR025) > 1 was used to detect safety signal40. We applied multivariate logistic model, expressed with the following formula, to present RORs adjusted for the potential confounder41,42,43,44,45:
where Y = reporting year, G = gender, A = age-stratified group, Rt = report type, Rs = report source by occupation, Ra = report source by affiliation, S = serious AE.
BCPNN: The BCPNN is based on the Bayesian statistical principles for quantification of dependencies between drug and AE. Disproportionality that shows the dependencies calculated by BCPNN is called the Information Component (IC) and defined as:
where Px = probability that specific drug is a suspected drug in a case report, Py = probability that specific AE is reported in a case report, Pxy = probability that specific drug-AE combination is listed on a case report. IC > 0 indicates that a particular drug-AE combination is reported more frequently than expected in a dataset; the higher the value of the IC, the more relevant the specific drug-AE combination stands out in the database.
Comparing performance of the data mining methods
Given that signal detection thresholds differ across the data mining methods, we calculated standardized differences to compare performance between the methods. The standardized difference refers to a difference between predictive value and signaling threshold in the standard deviation unit, and it was calculated as:
where Ps = Predictive value of an outcome of the signal detection algorithms, To = Optimal thresholds of the signal detection algorithms with the highest AUC, SD = Standard deviation which represents amount of variation or dispersion of the predictive values across all unknown AEs.
Infliximab-AE pairs with standardized difference > 0 were considered as the safety signals.
Also, we used the principles for evaluation of clinical diagnostic tests to quantify performances of the ML algorithms and conventional data mining methods in correctly differentiating AEs of infliximab. Reference standard was used to determine whether the safety signals were either true positives, false positives, true negatives, or false negatives to calculate sensitivity and specificity of each data mining method. Sensitivity was defined as the proportion of “label-positive” AEs that were correctly identified as signals (i.e., true positive/ [true positive + false negative]), and specificity as the proportion of “label-negative” that were correctly identified as non-signals (i.e., true negative/ [true negative + false positive]).
Secondary analysis: identifying new safety signals of infliximab
Among the AEs designated “U” in the labeled data (unknown AE dataset), we used the ML algorithms to identify safety signals for infliximab. For each detected signal, we also calculated signal detection scores of ROR025 and IC05 to determine whether it was also detected by these data mining methods.
Sensitivity analysis: validating the generalizability of ML algorithms
To validate our study findings, we further applied the ML algorithms to data retrieved from FAERS. First, we converted WHO-ART terms for defining AEs in the standard reference to Medical Dictionary for Regulatory Activities (MedDRA) terms. Then, we constructed an input dataset using AE reports in the FAERS from Q3 2014 to Q4 2018. Then, we applied ML algorithms to the input dataset to investigate drug-AE pairs in the standard reference and compared their predictive performance using the AUC of ROC plot.
All statistical analyses were performed using Python software version 3.7.5 (Python Software Foundation, Wilmington, DE, United States), SAS® software, version 9.4 (© 2002–2012 by SAS Institute Inc., Cary, NC, United States), and Microsoft Office 365 ProPlus (Microsoft Corp., Redmond, WA, United States). All methods used in this study were performed in accordance with the relevant guidelines and regulations.
Ethical approval
The Institutional Review Board of Sungkyunkwan University (IRB no. 2019-04-020-001) approved this study and waived the need for an informed consent as only deidentified data were used.
Data availability
The proposed framework implemented using SAS and Python, along with results generated in the study are available in the “Early-detection” repository, https://github.com/SKKUPEPV/Early-detection.git. Our study used the Korea Adverse Event Reporting System (KAERS) database, established by the Korea Institute of Drug Safety & Risk Management (KIDS) in South Korea (Data number: 1905A0020). KIDS forbids the transfer, rent, or sale of the database to any third party other than the researcher, who obtained the approval for the provided database (Official website of KIDS: http://open.drugsafe.or.kr/; Contact information of data access committee: + 82-2-2172-6700). We accessed the data used in our study in the above mentioned manner, which we expect future researchers to do so in the same manner, and did not receive special privileges from KIDS.
References
Vlahović-Palčevski, V. & Mentzer, D. Postmarketing surveillance. Handb. Exp. Pharmacol. 205, 339–351. https://doi.org/10.1007/978-3-642-20195-0_17 (2011).
Meyboom, R. H. et al. Principles of signal detection in pharmacovigilance. Drug Saf. 16, 355–365. https://doi.org/10.2165/00002018-199716060-00002 (1997).
CIOMS. (CIOMS, Geneva, 2010).
Waller, P. C., Coulson, R. A. & Wood, S. M. Regulatory pharmacovigilance in the United Kingdom: Current principles and practice. Pharmacoepidemiol. Drug Saf. 5, 363–375. https://doi.org/10.1002/(sici)1099-1557(199611)5:6%3c363::Aid-pds249%3e3.0.Co;2-7 (1996).
Paludetto, M. N., Olivier-Abbal, P. & Montastruc, J. L. Is spontaneous reporting always the most important information supporting drug withdrawals for pharmacovigilance reasons in France?. Pharmacoepidemiol. Drug Saf. 21, 1289–1294. https://doi.org/10.1002/pds.3333 (2012).
Clarke, A., Deeks, J. J. & Shakir, S. A. An assessment of the publicly disseminated evidence of safety used in decisions to withdraw medicinal products from the UK and US markets. Drug Saf. 29, 175–181. https://doi.org/10.2165/00002018-200629020-00008 (2006).
Egberts, A. C., Meyboom, R. H. & van Puijenbroek, E. P. Use of measures of disproportionality in pharmacovigilance: Three dutch examples. Drug Saf. 25, 453–458. https://doi.org/10.2165/00002018-200225060-00010 (2002).
Bate, A. et al. A Bayesian neural network method for adverse drug reaction signal generation. Eur. J. Clin. Pharmacol. 54, 315–321 (1998).
Rothman, K. J., Lanes, S. & Sacks, S. T. The reporting odds ratio and its advantages over the proportional reporting ratio. Pharmacoepidemiol. Drug Saf. 13, 519–523. https://doi.org/10.1002/pds.1001 (2004).
Lindquist, M., Ståhl, M., Bate, A., Edwards, I. R. & Meyboom, R. H. A retrospective evaluation of a data mining approach to aid finding new adverse drug reaction signals in the WHO international database. Drug Saf. 23, 533–542. https://doi.org/10.2165/00002018-200023060-00004 (2000).
Lee, C. Y. & Chen, Y. P. Machine learning on adverse drug reactions for pharmacovigilance. Drug Discov. Today 24, 1332–1343. https://doi.org/10.1016/j.drudis.2019.03.003 (2019).
Schotland, P. et al. Target-adverse event profiles to augment pharmacovigilance: A pilot study with six new molecular entities. CPT Pharmacomet. Syst. Pharmacol. 7, 809–817. https://doi.org/10.1002/psp4.12356 (2018).
Liu, M. et al. Large-scale prediction of adverse drug reactions using chemical, biological, and phenotypic properties of drugs. J. Am. Med. Inform. Assoc. 19, e28–e35. https://doi.org/10.1136/amiajnl-2011-000699 (2012).
Hoang, T., Liu, J., Roughead, E., Pratt, N. & Li, J. Supervised signal detection for adverse drug reactions in medication dispensing data. Comput. Methods Programs Biomed. 161, 25–38. https://doi.org/10.1016/j.cmpb.2018.03.021 (2018).
Bae, J.-H. et al. Machine learning for detection of safety signals from spontaneous reporting system data: Example of nivolumab and docetaxel. Front Pharmacol. 11, 602365–602365. https://doi.org/10.3389/fphar.2020.602365 (2021).
Pham, M., Cheng, F. & Ramachandran, K. A comparison study of algorithms to detect drug-adverse event associations: Frequentist, bayesian, and machine-learning approaches. Drug Saf. 42, 743–750. https://doi.org/10.1007/s40264-018-00792-0 (2019).
Nikfarjam, A. & Gonzalez, G. H. Pattern mining for extraction of mentions of Adverse Drug Reactions from user comments. In AMIA ... Annual Symposium proceedings/AMIA Symposium. AMIA Symposium 2011, 1019–1026 (2011).
Patki, A. et al. Mining adverse drug reaction signals from social media: Going beyond extraction. Proc. BioLinkSig 1, 8 (2014).
Shin, J. Y. et al. New initiatives for pharmacovigilance in South Korea: Introducing the Korea institute of drug safety and risk management (KIDS). Pharmacoepidemiol Drug Saf. 23, 1115–1122. https://doi.org/10.1002/pds.3715 (2014).
Lee, H., Kim, H. J., Choe, Y. J. & Shin, J. Y. Signals and trends of Guillain-Barré syndrome after the introduction of live-attenuated vaccines for influenza in the US and South Korean adverse event reporting systems. Vaccine 38, 5464–5473. https://doi.org/10.1016/j.vaccine.2020.06.038 (2020).
Jeong, D., Kim, W. & Shin, J. Y. Disproportionality analysis of spontaneously reported hypoglycemia events due to insulin use: A comparison between insulin detemir and insulin degludec using the Korea adverse event reporting system. Int. J. Clin. Pharmacol. Ther. https://doi.org/10.5414/cp203768 (2020).
Kim, S. et al. Data-mining for detecting signals of adverse drug reactions of fluoxetine using the Korea adverse event reporting system (KAERS) database. Psychiatry Res. 256, 237–242. https://doi.org/10.1016/j.psychres.2017.06.038 (2017).
Lee, H., Kim, J. H., Choe, Y. J. & Shin, J.-Y. Safety surveillance of pneumococcal vaccine using three algorithms: Disproportionality methods, empirical bayes geometric mean, and tree-based scan statistic. Vaccines (Basel) 8, 242. https://doi.org/10.3390/vaccines8020242 (2020).
Kim, J. H., Lee, H. & Shin, J. Y. Bacillus calmette-guérin (BCG) vaccine safety surveillance in the Korea adverse event reporting system using the tree-based scan statistic and conventional disproportionality-based algorithms. Vaccine 38, 3702–3710. https://doi.org/10.1016/j.vaccine.2020.04.007 (2020).
Kreimeyer, K. et al. Using probabilistic record linkage of structured and unstructured data to identify duplicate cases in spontaneous adverse event reporting systems. Drug Saf. 40, 571–582. https://doi.org/10.1007/s40264-017-0523-4 (2017).
Slattery, J. Measuring signal detection performance: Can we trust negative controls and do we need them?. Drug Saf. 39, 371–373. https://doi.org/10.1007/s40264-016-0407-z (2016).
Hauben, M., Aronson, J. K. & Ferner, R. E. Evidence of misclassification of drug-event associations classified as gold standard “negative controls” by the observational medical outcomes partnership (OMOP). Drug Saf. 39, 421–432. https://doi.org/10.1007/s40264-016-0392-2 (2016).
Wang, C. et al. Exploration of the association rules mining technique for the signal detection of adverse drug events in spontaneous reporting systems. PLoS ONE 7, e40561. https://doi.org/10.1371/journal.pone.0040561 (2012).
Bate, A. & Evans, S. J. Quantitative signal detection using spontaneous ADR reporting. Pharmacoepidemiol Drug Saf. 18, 427–436. https://doi.org/10.1002/pds.1742 (2009).
Sakaeda, T., Kadoyama, K., Minami, K. & Okuno, Y. Commonality of drug-associated adverse events detected by 4 commonly used data mining algorithms. Int. J. Med. Sci. 11, 461–465. https://doi.org/10.7150/ijms.7967 (2014).
Schotland, P. et al. Target adverse event profiles for predictive safety in the postmarket setting. Clin. Pharmacol. Ther. 109, 1232–1243. https://doi.org/10.1002/cpt.2074 (2021).
Cheng, F. & Zhao, Z. Machine learning-based prediction of drug-drug interactions by integrating drug phenotypic, therapeutic, chemical, and genomic properties. J. Am. Med. Inform. Assoc. JAMIA 21, e278-286. https://doi.org/10.1136/amiajnl-2013-002512 (2014).
National Library of Medicine. DailyMed Label Archives (National Library of Medicine, Bethesda, 2020).
Wahab, I. A., Pratt, N. L., Wiese, M. D., Kalisch, L. M. & Roughead, E. E. The validity of sequence symmetry analysis (SSA) for adverse drug reaction signal detection. Pharmacoepidemiol Drug Saf. 22, 496–502. https://doi.org/10.1002/pds.3417 (2013).
Harpaz, R. et al. A time-indexed reference standard of adverse drug reactions. Sci. Data 1, 140043. https://doi.org/10.1038/sdata.2014.43 (2014).
European Medicines Agency. Guideline on good pharmacovigilance practices (GVP) Module VI–Collection, management and submission of reports of suspected adverse reactions to medicinal products https://www.ema.europa.eu/en/documents/regulatory-procedural-guideline/guideline-good-pharmacovigilance-practices-gvp-module-vi-collection-management-submission-reports_en.pdf.
Ran, J. et al. Signal detection of human papillomavirus vaccines using the Korea adverse events reporting system database, between 2005 and 2016. Int. J. Clin. Pharm. 41, 1365–1372. https://doi.org/10.1007/s11096-019-00881-9 (2019).
Soukavong, M. et al. Signal detection of adverse drug reaction of amoxicillin using the Korea adverse event reporting system database. J. Korean Med. Sci. 31, 1355–1361. https://doi.org/10.3346/jkms.2016.31.9.1355 (2016).
Matsui, T. et al. Age-related trends in injection site reaction incidence induced by the tumor necrosis factor-α (TNF-α) inhibitors etanercept and adalimumab: The food and drug administration adverse event reporting system, 2004–2015. Int. J. Med. Sci. 14, 102–109. https://doi.org/10.7150/ijms.17025 (2017).
van Puijenbroek, E. P. et al. A comparison of measures of disproportionality for signal detection in spontaneous reporting systems for adverse drug reactions. Pharmacoepidemiol Drug Saf. 11, 3–10. https://doi.org/10.1002/pds.668 (2002).
Van Puijenbroek, E. P., Egberts, A. C., Meyboom, R. H. & Leufkens, H. G. Signalling possible drug-drug interactions in a spontaneous reporting system: Delay of withdrawal bleeding during concomitant use of oral contraceptives and itraconazole. Br. J. Clin. Pharmacol. 47, 689–693. https://doi.org/10.1046/j.1365-2125.1999.00957.x (1999).
van Puijenbroek, E. P., Egberts, A. C., Heerdink, E. R. & Leufkens, H. G. Detecting drug-drug interactions using a database for spontaneous adverse drug reactions: An example with diuretics and non-steroidal anti-inflammatory drugs. Eur. J. Clin. Pharmacol. 56, 733–738. https://doi.org/10.1007/s002280000215 (2000).
Qian, Y. et al. A computerized system for detecting signals due to drug-drug interactions in spontaneous reporting systems. Br. J. Clin. Pharmacol. 69, 67–73. https://doi.org/10.1111/j.1365-2125.2009.03557.x (2010).
Suzuki, Y. et al. Analysis of the interaction between clopidogrel, aspirin, and proton pump inhibitors using the FDA adverse event reporting system database. Biol. Pharm. Bull. 38, 680–686. https://doi.org/10.1248/bpb.b14-00191 (2015).
Abe, J. et al. Evaluation of dabigatran- and warfarin-associated hemorrhagic events using the FDA-Adverse event reporting system database stratified by age. Int. J. Med. Sci. 12, 312–321. https://doi.org/10.7150/ijms.10703 (2015).
Acknowledgements
This research was supported by Government-wide R&D Fund project for infectious disease research (GFID), Republic of Korea (grant No HG18C0068). This research was supported by a grant (21153MFDS607) from Ministry of Food and Drug Safety of South Korea in 2021–2025. This work was supported by the Bio Industry Technology Development Program (No. 20015086) By the Ministry of Trade, Industry & Energy (MOTIE, Korea).
Author information
Authors and Affiliations
Contributions
J.L. conceived the study design, performed data analysis, and drafted the manuscript. J.H.K. conceived the study design, and drafted the manuscript. J.B. conceived the study design, and performed data analysis. I.S. conceived the study design. J.S. conceived the study design, and oversighted all aspect of the study. All authors read and approve the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, JE., Kim, J.H., Bae, JH. et al. Detecting early safety signals of infliximab using machine learning algorithms in the Korea adverse event reporting system. Sci Rep 12, 14869 (2022). https://doi.org/10.1038/s41598-022-18522-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18522-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.