Abstract
Post-weaning enteropathies in swine caused by pathogenic E. coli, such as post-weaning diarrhea (PWD) or edema disease (ED), remain a significant problem for the swine industry. Reduction in the use of antibiotics over concerns of antibiotic resistance and public health concerns, necessitate the evaluation of effective antibiotic alternatives to prevent significant loss of livestock and/or reductions in swine growth performance. For this purpose, an appropriate piglet model of pathogenic E. coli enteropathy is required. In this study, we attempted to induce clinical signs of post-weaning disease in a piglet model using a one-time acute or lower daily chronic dose of a pathogenic E. coli strain containing genes for both heat stable and labile toxins, as well as Shiga toxin. The induced disease state was monitored by determining fecal shedding and colonization of the challenge strain, animal growth performance, cytokine levels, fecal calprotectin, histology, fecal metabolomics, and fecal microbiome shifts. The most informative analyses were colonization and shedding of the pathogen, serum cytokines, metabolomics, and targeted metagenomics to determine dysbiosis. Histopathological changes of the gastrointestinal (GI) tract and tight junction leakage as measured by fecal calprotectin concentrations were not observed. Chronic dosing was similar to the acute regimen suggesting that a high dose of pathogen, as used in many studies, may not be necessary. The piglet disease model presented here can be used to evaluate alternative PWD treatment options.
Similar content being viewed by others
Introduction
Post-weaning diarrhea (PWD) and edema disease (ED) following the weaning period in piglets remain significant problems for the swine industry and can result in significant economic losses1,2,3. PWD is characterized by diarrhea which can lead to severe dehydration, emaciation, and death. While ED of swine is characterized by submucosa edemas of the stomach and mesocolon resulting in swelling of eyelids, forehead, and in some cases hemorrhagic gastroenteritis leading to eventual death2. Pathogenic Escherichia coli is the primary cause of these diseases in swine, and the transitionary period of weaning leaves piglets susceptible to infection by pathogenic strains of E. coli3,4. While PWD and ED are generally caused by enterotoxigenic E. coli (ETEC) and Shiga toxin–producing E. coli (STEC), respectively, they affect similarly aged pigs and there can be considerable crossover between serotypes and associated virulence factors. PWD ETEC are primarily associated with E. coli producing heat-stable and/or heat-labile enterotoxin, while ED STEC are associated with Shiga toxin, primarily the 2e subtype (Stx2e), producing strains, which can be expressed with or without other enterotoxins5. The antibiotic colistin has been the classical treatment for pathogenic E. coli in swine, however given concerns over antibiotic resistance, alternative treatment options should be explored1. To evaluate alternative treatment options, a comprehensive model of E. coli pathogenesis in swine is necessary to evaluate efficacy of alternative treatments to antibiotics.
Swine inoculated with ETEC experience sloughing of intestinal villi, increased crypt depths, and scours6,7. It has been shown that ETEC infections in weanling pigs can be caused by a single dose of approximately 109 CFU8,9. However, this high acute single dose most likely does not accurately represent the real-world scenario of PWD or ED in which piglets are more likely initially infected by chronic exposure to lower doses of E. coli as can be found in contaminated feed, water, soil, and elsewhere in the barn environment2. The objective of the present study was to develop and characterize a pathogenic E. coli induced weaned swine model of PWD/ED. The E. coli strain used in this work encodes F18ac + fimbrial adhesin, heat-labile enterotoxin IIA (LT-IIA), heat-stable enterotoxin II (STIIB), as well Shiga toxin (Stx2e), technically classifying it as both an ETEC and STEC. However, there was no evidence of the edemas classically associated with the presence of Stx2e, making significant expression of Shiga toxin under the trial circumstances questionable, thus the trial strain will simply be referred to as a pathogenic E. coli strain. We also sought to evaluate differences in dosing regimens, comparing a one-time high acute dose to a lower daily chronic dose of pathogenic E. coli. To our knowledge, this is the first reported comparison on the effects of a one-time high acute dose vs a lower chronic daily dose, consisting of a tenfold dilution of an overnight culture given for 6 days, in an animal model. Although in the model presented here, it is unclear if the observed differences between dosing regimens are primarily due to E. coli dosing concentration or dosing frequency. The detailed model presented here of E. coli induced gastrointestinal disease provides valuable insights for future novel treatments for post-weaning diarrhea common amongst piglets.
Results and discussion
Growth performance
Pigs used as an experimental model for PWD were challenged with a spontaneous nalidixic acid-resistant mutant of Escherichia coli strain NCDC 62–57 (ATCC 23,545) referred to hereafter simply as E. coli 62-57nal in either a single acute high-titer dose (1.5 × 1010 CFU), or in a series of daily lower-dose challenges (average daily dose 5.4 × 108 CFU). This strain was chosen for its ability to cause illness during preliminary trials amongst F18 + and K88 (F4) ETEC strains. All pigs were held for two days prior to the start of the trial and were asymptomatic for gastroenteritis. Additionally, pigs were not colonized by organisms capable of forming colonies on MacConkey amended with 50 µg/ml nalidixic acid (MacConkey + nal), and no endogenous phage infecting E. coli 62-57nal were identified. Thirty-six presumptive coliform colonies from pooled fecal samples plated on MacConkey agar (0 µg/ml Nal; three colonies per pen) were also tested by PCR for the presence of Shiga toxin type 1 (Stx1), Shiga toxin type 2 (Stx2), heat-stable enterotoxin I (ST1), heat-stable enterotoxin II (ST2) and heat-labile toxin (LTI). All colonies were negative for Stx1, Stx2, ST2 and LTI, but three colonies were positive for ST1. Presence of ST1 gene alone is not a strong predictor of ability to cause disease8,10,11 and pigs were asymptotic, so all animals were retained in the study.
In general, pigs administered E. coli 62-57nal via both the acute and chronic dosing regimens presented similar clinical signs with the majority of pens developing scours by day 2 and continuing through day 6. Control pens had visibly soft feces on day 5 and 6 with a single incident of scours on day 6, however the animals in control pens remained visibly healthy throughout the trial period. Additionally, the control pen with the incidence of scours was culture negative for the inoculated E. coli 62-57nal throughout the trial, so scours may have been induced by stress or other native microbiota. There was no evidence of difference for overall average daily gain (ADG), average daily feed intake (ADFI), and gain:feed (G:F) of the different treatment groups (P > 0.184, Table 1). However, there were numerical differences between pigs fed the treatments, suggesting that the modest number of replicates and the inherently high post weaning variability in performance were responsible for the failure to detect significant differences in growth performance. This lack of evidence for significant growth differences is similar to previously reported results12; however, significant changes to growth performance in response to ETEC challenge have also been reported13. Pigs administered the acute and chronic dose of E. coli 62-57nal had a 54.7% and 14.9% reduction in ADG compared to the control pigs, respectively (Table 1). The control group had the lowest ADFI among the three treatments with acute and chronic dosing regimens increasing feed intake by 17.3% and 29.95%. These findings are in agreement with previous work that showed a 24% decrease in control pigs ADFI compared to the pigs inoculated with ETEC O149 on d 3 to d 6 9. Madec et al.14 had similar results with a decrease in weight of weaned piglets inoculated with pathogenic E. coli expressing K88 fimbriae from day 0 to day 2 which then recovered by day 9 of the trial. In this study, the acute challenge group had the poorest mean G:F conversion with the control group having the highest mean feed efficiency. Piglets experiencing PWD have been reported to exhibit reduced weight gains3,15, however statistically significant reductions in weight performance were not observed, perhaps due to the relatively brief duration of the trial or small sample sizes.
Bacterial colonization and fecal shedding
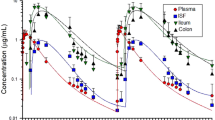
The ability of E. coli 62-57nal to colonize the gastrointestinal tract of inoculated piglets was determined by measuring colony-forming units recovered from intestinal mucosa, intestinal luminal contents, and in feces. Inoculated strain counts adherent to the mucosal lining were found to be variable, with ~ 50% of samples, ranging from 0.21 to 1.71 g of intestinal scraping, yielding counts above the detection limit (5000 CFU/ml of tissue homogenate). Of the samples yielding enumerable colonies, bacterial counts ranged from ~ 104 to ~ 107 CFU/g in the duodenum, jejunum, ileum, cecum and colon (Fig. 1A). Bacterial counts in the cecal and colonic luminal contents, ranging from 0.14 to 11.07 g of digesta, were more reliably above the detection limit and ranged from ~ 103 to ~109 CFU/g, suggesting bacterial proliferation in the unattached population.
Acute and chronic regimens are sufficient to cause colonization of E. coli 62-57nal. (A) At time of necropsy, intestinal scrapings from the duodenum, jejunum, ileum, cecum, and colon were collected and plated for bacterial enumeration. Additionally, cecum and colon contents were collected for direct plating. (B) Fecal samples were collected from each pen on day 1,3, and 6. These samples were serial diluted and plated for direct enumeration of bacteria. LOD ~ 5 × 102 CFU/G. Both (A) and (B) Circles indicate sampling data points, diamonds indicate mean. Sample data was transformed using LOG10. Treatment means grouped by day/section with different letters were significantly different (P < 0.05), those lacking letters indicate no level of significance.
Acute and chronic treatments had higher levels of E. coli 62–57-nal in feces (~ 105 to 107 CFU/g) than control pigs on all sampling days (Fig. 1B). Despite the preventative efforts described in the Materials and Methods, control pens sporadically shed E. coli 62–57-nal in the feces at levels near the lower detection limit (500 CFU/g), likely reflecting low levels of pen cross-contamination. Pigs administered the acute pathogenic E. coli dose exhibited significantly higher fecal shedding on day 1 (~ 107 CFU/g, P = 0.001) compared to the chronic dose, however there was no statistically significant difference in fecal E. coli 62-57nal counts between the acute and chronic treatments after d 1. This result is consistent with other piglet studies which observed peak shedding between 24 and 48 h post-inoculation16,17. Pathogen shedding in the acute group remained high through d 6, indicating that the E. coli 62-57nal was able to persistently colonize the gastrointestinal tract of swine. Although, the possibility of low levels of reinfection from the contaminated pens cannot be completely ruled out. High inter-animal variability in intestinal bacterial loads and fecal shedding has been observed in other studies examining F18 E. coli challenge in immature pigs, commonly spanning 3–4 orders of magnitude in both genetically susceptible and resistant populations18,19.
Markers of inflammation and intestinal leakage
Infection-induced inflammation is mediated by increased levels of pro-inflammatory cytokines20. To evaluate the level of E. coli induced inflammation, serum interleukins 6 and 8 (IL-6 and IL-8) were measured. IL-6 and IL-8 are useful biomarkers since they have been linked to intestinal inflammation21,22,23,24 and have been employed as markers of porcine immune activation in response to E. coli previously25,26,27,28. On d 6 of the study, pigs challenged with E. coli 62-57nal had increased (P < 0.05) concentrations of serum IL-6 compared to control pigs (Fig. 2). However, there was no difference in IL-6 concentrations between acute and chronic treatments. Similar elevations of IL-6 were also observed in the treatment groups of a bacterially induced murine model of chronic intestinal inflammation29. For concentrations of IL-8, there was a marginally significant overall treatment effect on d 6 (P = 0.089): chronic pigs had increased (P < 0.05) serum IL-8 concentrations compared to control pigs, and acute dose pigs were intermediate (P = 0.5423). Lee et al.27 observed peak serum IL-8 levels in ETEC-challenged piglets between day 0 and 2 which then declined through d 7. This could explain the lower levels of serum IL-8 in the acute challenge group as serum cytokines were measured six days after acute challenge, while the daily chronic challenge may maintain elevated IL-8 concentrations. Increased levels of IL-6 and IL-8 in response to challenge with E. coli 62-57nal is in agreement with prior work demonstrating these cytokines as markers of inflammation and infection27,29.
Increased concentrations of fecal calprotectin have been positively correlated with the histological activity of inflammatory bowel disease in humans30, and serve as a marker of neutrophilic intestinal inflammation31. Fecal calprotectin has also been suggested as a noninvasive marker of intestinal inflammation in swine32, however additional study is necessary. Past studies have investigated calprotectin in swine plasma, intestinal lumen, and jejunal mucosa, all of which were found to be correlated with bacterial infection33,34. In the present study, there were no significant treatment effects on fecal calprotectin concentration. This is the first study to our knowledge to test calprotectin levels in the feces of pigs inoculated with E. coli, and this indicates fecal calprotectin may not be an informative biomarker in this model.
Villi length and histopathology
Previous studies involving swine challenged with ETEC strains have reported villous atrophy and reductions in crypt depth35; similar symptoms have also been reported in chronic intestinal enteropathies in humans36. At time of necropsy, sections were collected to evaluate villi length in the piglet model, but no morphologic changes were observed between treatment groups. General bacterial rod attachment was evaluated by an anatomic pathologist and observed sporadically in all samples with no apparent correlation between rod attachment and direct bacterial plating as only 38% (11/29) of samples with rod attachment tested positive for E. coli 62-57nal by direct plating. Villus length in E. coli challenged animals did not differ from the controls in the duodenum (P = 0.7125), jejunum (P = 0.3719), and ileum (P = 0.778). Lack of villus blunting may be due to the limited duration of this study. A prior longer-term study (21 d), with a murine model of chronic intestinal inflammation obtained villus blunting through a combination of bacterial challenge and malnutrition29. Similarly, post-weaning anorexia in piglets has been shown to be associated with reduced villus heights37. Therefore, given a longer trial period and/or malnourishment, blunting may have been eventually observed in our present model. Histology is also only able to evaluate a tiny fraction of the intestinal tract, so lesions must be broadly distributed throughout the tissue to be detectable by this method. Based on this data, histologic analysis does not appear to be a useful method for evaluating this model.
Effects on the microbiome by 16S qPCR analysis
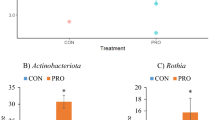
To observe any changes of the gut microbiota caused by our acute or chronic dosing treatments, targeted 16S qPCR was performed for select bacterial groups on fecal samples collected from pens at day −1, day 1, day 3, and day 6. Relative abundances obtained were consistent with previous examinations of the piglet microbiome, showing a microbiome dominated by Bacteroidetes and Firmicutes38,39. Overall, the bacterial groups tended to increase relative to control and pre-treatment samples, likely due to natural microbiome succession. A summary of these significant (P < 0.05) or marginally significant (P < 0.10) bacterial group changes at each time point is shown in Table 2. Both acute and chronic doses of E. coli 62-57nal impacted relative quantities of E. coli populations compared to the control. Additionally, both dosing regimens had comparable impacts on microbiome progression. Pretreatment compared to post-treatment samples of the acute dose had the most significant/marginally significant changes with eight of the ten tested bacterial taxa (Bacteroidetes, Enterococcus, Faecalibacterium, Firmicutes, Lactobacillus, Streptococcus, Fusobacterium, and Universal) showing increased populations. The chronic treatment showed similar but less dramatic changes, with six of ten taxa (Bacteroidetes, Enterococcus, Lactobacillus, Streptococcus, E. coli, and Ruminococcaceae) showing increased levels from pre- to post-treatment. The control group showed only two altered bacterial groups, Enterococcus and E. coli, from pre- to post-treatment. The observed increase for Enterococcus and E. coli within the control treatment is consistent with the previously reported natural post-weaning piglet microbiome maturation which shows an increase in levels of Enterococcus and Enterobacteriaceae at 8 days post-weaning40. The acute dose of E. coli had a slightly more pronounced impact on the gut microbiome maturation than the chronic dose, however both acute and chronic treatments were sufficient to cause a detectable dysbiosis. This dysbiosis indicates incidences of ETEC amongst weaned piglets can significantly disrupt the normal microbiome progression during this transitionary period.
Principal component analysis (PCA) of 16S qPCR results also provides clear evidence of dysbiosis in E. coli-treated groups (Fig. 3A). PCA of pre- vs post-treatment samples indicates that by day 6 the acute treatment clearly clustered away from its pretreatment sample, while the chronic day 6 sample showed an intermediate clustering from its pre-treatment sample. In contrast, the control group remained tightly clustered throughout the trial period. This contrast in clustering suggests the microbiome perturbations are induced by the E. coli challenge and are not merely normal microbiota progression. The observed relative stability of the control microbiome is consistent with other studies, which reported a microbiome shift immediately after weaning and reached relative stability within 10 days after weaning39. These findings indicate that both the single acute dose and the chronic lower dose of E. coli 62-57nal caused varying degrees of a similar dysbiosis. While in this present study the acute dose of pathogenic E. coli provided a more pronounced microbiome defect, the slight alteration caused by the chronic dose may still be more reflective of chronic exposure to pathogens, as piglets may experience.
Clustering of treatments by principal component analysis (PCA) of metagenomic and metabolomic results. By the end of the trial period, both chronic and acute treatments are able to be separated from their respective pretreatment samples using qPCR and metabolomic data while the control group remains relatively constant. (A) PCA comparing pre-treatment and post-treatment samples using 16S qPCR of major bacterial taxonomic groups. Red dots indicate pre-treatment samples, green diamonds indicate post-treatment samples, and green/red ellipses represent 95% confidence regions. (B) PCA comparing pre-treatment and post-treatment samples using identifiable fecal metabolites.
Alterations in the fecal metabolome
To further characterize the differences in disease state caused by acute and chronic E. coli 62-57nal challenge, untargeted metabolomics was performed on fecal samples collected pretreatment (d −1) and days 1, 3 and 6 post-treatment. Metabolite profiles of fecal samples were analyzed by Metaboanalyst41. Analysis of the identifiable metabolites by PCA clearly distinguished between challenge and control groups (Fig. 3B). Similar to the results of microbiome analysis (Fig. 3A), the acute and chronic day 6 samples clearly cluster separately from their pre-treatment samples, while the control samples did not separate. The stability of the control group indicates the natural enzymatic, microbial, and structural maturation of the weaned piglet gut40 was not responsible for the observed shifts in the acute or chronic treatment groups.
As with the 16S qPCR approach, metabolomic comparison of post-treatment samples with their respective pre-treatment samples was more informative when identifying significant changes in individual metabolites. Volcano plots (P-value < 0.10 and > twofold change) were used to identify metabolites that significantly changed following treatment41 (Fig. 4). A full list of metabolites identified by volcano plot is provided in S1 Table. Metabolomics data related to swine, while growing, is fairly limited42,43,44,45,46, particularly metabolomics pertaining to piglets in a diseased stated, most of which are serum or tissue based47,48,49,50. The Human Metabolome Database51 is much more comprehensive, particularly for diseased states, than the Livestock Metabolome Database 52. Given these limitations in existing swine metabolomics data, the metabolites identified in this manner were categorized based on the Human Metabolome Database chemical taxonomy and compared with relevant human diseased states in which fecal metabolomics are more prevalent. Only four identified metabolites were shared between the chronic and control groups, and ten were common to both the chronic and acute treatment groups (Fig. 4). Changes in metabolites in the control group were presumed to be associated with the normal development of the weaned piglet gastrointestinal tract.
Volcano plots comparing changes in identifiable metabolite profiles in pre- and post- treatment fecal samples. Figures show metabolites that significantly changed from day −1 to day 6 within each pathogenic E. coli dosing regimen. Points represent individual metabolites, which are color-coded based on the Human Metabolomics Database chemical taxonomy 51. Dotted lines indicate significance cutoffs of P-value < 0.1 (Y-axis) and > twofold change in abundance (X-axis); points in the upper left and right quadrants of each graph represent metabolites with significant, > twofold changes from pre- to post-treatment. Point shape indicates if a metabolite is shared by the acute and chronic treatments (squares) or chronic and control treatments (triangles); circular points indicate metabolites that were either not detected or had non-significant changes in other treatments. Both of the E. coli-treated groups exhibited elevated levels of metabolites associated with amino acids and reduced metabolites associated with fatty acids compared to the control group.
Within the chronic and acute treatments, increased levels of amino acid metabolites were identified in the post-treatment samples, including lysine, ornithine, homoserine, histidine, tyramine, beta-alanine, (Fig. 4, S1 Table). The increased levels of amino acids and amino acid metabolites in post-treatment fecal samples suggests the E. coli 62-57nal treatment led to amino acid malabsorption and/or secretion, likely due to disruption of chemical gradients, inflammation, and microbiome perturbations within the gut caused by pathogenic E. coli treatment. Additionally, metabolites associated with bacterial amino acid degradation, 5-aminovaleric acid and putrescine, were found at increased levels within acute and chronic treatment samples. These have been previously associated with ulcerative colitis53,54. The presence of these metabolites is consistent with the model that suggests inflammation caused by E. coli treatment induced amino acid malabsorption and subsequent degradation by the resident microbiota. Increased presence of fecal amino acids is supported by Wu et al. who reported decreased amino acid metabolites in jejunal tissue from ETEC challenged piglets49, supporting malabsorption in small intestine leading to increased presence of amino acids in the feces. This observation of increased fecal amino acid and amino acid metabolite levels also agrees with studies examining fecal metabolite profiles of humans with inflammatory bowel diseases like ulcerative colitis and Crohn’s disease of which chronic diarrhea is a symptom55,56.
Fecal metabolites that were significantly reduced in post-E. coli treatment samples were primarily fatty acid metabolites (Fig. 4), including stearic acid, myristic acid, and arachidic acid. The levels of lipid-soluble alpha-tocopherol (vitamin E) was also reduced in both treatment groups. The observed depletion of fatty acids within the feces may be indicative of immune system activation, which is supported by our observation of increased levels of serum IL-6 and IL-8. Growing evidence on the importance of “immunometabolism” suggests activated macrophage subtypes and chronically activated T-cells demonstrate increased uptake of fatty acids as they rely more on fatty acid oxidation in order to maintain the high energy levels required to mount an immune response57,58. Prior studies examining metabolomic profiles of human inflammatory bowel diseases found pronounced decreases in the levels of short chain fatty acids (SCFA), which are the end products of bacterial fermentation that are absorbed by the large intestine; this presumably signaled a dysbiosis of gut flora55,56. In our current study a reduction in the SCFA metabolites butyrate, alpha-ketoglutarate and fumaric acid were observed. Our metabolomic findings indicate both the chronic and acute E. coli 62-57nal treatments caused sufficient dysbiosis to statistically distinguish pre- and post-treatment samples (Fig. 3B) in large part due to amino acid malabsorption and reduction in fatty acid metabolites (Fig. 4), generating metabolomic profiles resembling those of human inflammatory gastrointestinal disease associated with diarrhea.
Whole genome sequencing
To better understand the gene content of E. coli 62-57nal that may contribute to its pathogenicity, its genome was sequenced. The genome of E. coli 62-57nal was assembled into 378 contigs of greater than 200 bp totaling in 5.6 Mbp length and at an average 45-fold coverage. The entire set of 378 contigs was submitted for sequence typing using SerotypeFinder v1.159 and confirmed to be O138 and H14 as reported previously60. Analysis of the assembled contigs by BLASTx (> 40% identity, E value < 10–5) against a database of known E. coli virulence genes identified a number of genes in E. coli 62-57nal associated with pathogenesis, shown in S2 Table. Major identified virulence factors in E. coli 62-57nal include hemolysin (hlyABCD), Shiga toxin (stx2e), an intimin-like adhesin (fdeC), heat-stable enterotoxin II (stiI), heat-labile enterotoxin IIA (eltAB), a heme uptake system for acquisition of iron (chuUAVYTWSX), and the F18ac+ fimbrial adhesin. Hemolysin (Hly) is an exotoxin that is associated with many pathogenic strains of E. coli61. Hemolysin is primarily thought to have a role in pathogenesis in extra-intestinal infections, such as those of the urinary tract or septicemia and studies have shown that Hly plays little to no role in clinical signs of diarrhea62. However recent data, using both in-vivo murine and in-vitro models, show secretion of Hly can disrupt tight-gap junctions and increase colonic mucosal inflammation63,64. This inflammation from Hly has been proposed as a contributing factor for ulcerative colitis in humans, a chronic inflammation63. Consistent with E. coli 62–57’s original edema isolation source, an Stx2e Shiga toxin was identified on a putative prophage element. The Stx2e subtype is known to be associated with edema disease of swine65, however in our present study we did not observe signs of edema in challenged piglets. The primary contributors to the observed scouring phenotype were most likely the identified heat-stable enterotoxin II (STIIB) and heat-labile enterotoxin IIA (LT-IIA). While acting by different modes, both STIIB and LT-IIA have been shown to cause release of water and electrolytes from host membranes, thereby causing diarrhea or scours11,66 . Heat stable and heat labile enterotoxins are the most common exotoxins that are associated with diarrhea in piglets, present in 72% and 57% of ETEC isolates from piglet scours, respectively67. E. coli 62-57nal also contains a number of virulence factors associated with colonization and survival within the host, including the iron scavenging chu genes and various adhesin genes coding for the proteins AIDA-I autotransporter, F18ac+ fimbrial adhesin, and the intimin-like FdeC61,67,68. Taken together, the presence of these virulence factors explains the observed diarrhea/scouring and colonization phenotype. Furthermore, the enterotoxin mode of action that induces this diarrhea also disrupts the Na + gradient66, which amino acid absorption is dependent upon in the gut69, likely explaining the increased amino acid levels observed in the fecal metabolomes of ETEC-challenged animals.
Conclusion
Suitable piglet models exploring the pathogenesis of the E. coli pathotypes responsible for common post-weaning diseases like PWD and ED are necessary for evaluating alternative treatment options. The model presented here may serve as a detailed model for the evaluation of novel PWD treatments. In this study, weaned piglets were challenged with either a single bolus of 1.5 × 1010 CFU of a pathogenic STIIB + , Stx2e + , and LT-IIA + E. coli strain or average daily doses of 5.4 × 108 CFU of the same strain. Both chronic and acute treatment groups exhibited significant increases in fecal E. coli 62-57nal shedding, intestinal E. coli 62-57nal colonization and serum IL-6 levels compared to controls (Figs. 1, 2). Furthermore, both treatments induced similar levels of dysbiosis as measured by targeted 16S qPCR and untargeted metabolomics (Fig. 3). These alterations of the microbiome and metabolome consisted of significant increases in Bacteroidetes and Enterococcus, amongst others, and increased levels of amino acid metabolites within the feces. These findings imply that high acute doses of inoculum, as are often utilized in studies of E. coli gastrointestinal disease in pigs, are not necessarily required to establish a disease state, and lower levels of inoculum may represent a comparable disease state important for the study of chronic inflammation or chronic exposure to pathogens. Although, as presented here, it is unclear if a single low dose of E. coli would induce similar clinical signs to the high acute dose without repeated dosing. Future work may benefit from an additional treatment group consisting of a single low dose. In this study, fecal calprotectin measurements and histological examination of intestinal sections from challenged animals did not indicate any significant differences between control and treatment groups. Given the lack of blunted villi in this work which was previously associated with PWD, and likely a contributing factor of decreased nutrient uptake, future studies employing the model described here may benefit from an extended trial period, as well as controlled changes in animal nutrition, to achieve blunted villi.
In addition to a model for PWD, the relatively mild disease state presented here may also be adapted and appropriate for modeling of chronic human enteropathies in which an underlying dysbiosis is a presumed cause like inflammatory bowel disease70 or the understudied environmental enteric disfunction36. Rodent models have often been used as disease surrogates, however these models are at a disadvantage when it comes to accurately representing human diseases and syndromes. Swine more accurately resemble humans in anatomy, genetics, and physiology, making them a more appropriate model for human biomedical research71. For example, many of the differential metabolites identified in this study (Fig. 4) and the increases in inflammatory cytokines (Fig. 2) reflect those found in human chronic inflammatory disorders70,72. However, if modeling of the villous blunting associated with disorders like environmental enteric disfunction is desired, increase in trial length and additional malnutrition may be required, as mentioned previously.
Materials and methods
Ethics declaration
All procedures involving animals and their care were approved and monitored by the Animal Care and Use Committee of the USDA Southern Plains Agricultural Research Center (SPARC) and the Texas A&M University Institutional Animal Care and Use Committee. All approved methods were carried out in accordance with Animal Care and Use Committee of the USDA Southern Plains Agricultural Research Center (SPARC) and the Texas A&M University Institutional Animal Care and Use Committee. Materials and Methods, and Results sections are reported and prepared in accordance with ARRIVE guidelines.
Bacterial strains and culture conditions
Escherichia coli strain NCDC 62–57 (O138:K81(B):H14) was originally isolated from swine showing clinical signs of porcine edema disease60. This strain was obtained from the ATCC (ATCC 23,545). A spontaneous nalidixic acid-resistant mutant of this strain was isolated and used throughout this work, and will be referred to as strain 62-57nal. This mutation was stable through multiple consecutive transfers. The bacterium was routinely cultured in LB broth (Bacto tryptone 10 g/L, Bacto yeast extract 5 g/L, NaCl 10 g/L) or on LB agar plates (LB broth plus 15 g/L Bacto agar) aerobically at 37 °C. Samples obtained from animals challenged with E. coli 62-57nal were plated on MacConkey agar (Becton–Dickinson) amended with 50 µg/ml nalidixic acid (MacConkey + nal).
Whole genome sequencing of E. coli 62-57nal
Total DNA was extracted from an overnight culture of E. coli 62-57nal using the Qiagen DNeasy Blood and Tissue Kit following the manufacturer’s specifications for bacterial cells (Qiagen, Cat No. 69504). Isolated genomic DNA was sequenced on the Illumina MiSeq platform using Illumina V2 500 cycle reagent chemistry generating paired-end 250 bp reads. Reads were quality controlled using FastQC (bioinformatics.babraham.ac.uk), FastX Toolkit (hannonlab.cshl.edu), and assembled using SPAdes 3.5.073 at k-mer settings of 21,33,55. Contigs < 200 bp or with aberrantly low coverage (< eightfold) were filtered from the assembly to yield 378 contigs totaling 5.6 Mbp with ~ 45-fold average coverage. The resulting contigs were deposited to Genbank under Bioproject/Accession (PRDF00000000), and underwent automated annotation using the NCBI Prokaryotic Genome Annotation Pipeline74.
Putative virulence factors were identified based on homology using BLASTx of WGS contigs with a custom database of E. coli virulence factors containing E. coli-associated proteins contained in mVirDB75 and characterized E. coli virulence factors61. As a control, the genome of the non-pathogenic lab strain of E. coli MG1655 (Accession: GCF_000005845.2) was analyzed against the same database. Hits in common from MG1655 and 62-57nal were excluded based on the presumption that they were part of the non-pathogenic E. coli gene repertoire. Protein sequences identified in this initial screening were extracted from the 62-57nal genome and manually investigated using BLASTp and InterProScan to confirm conserved domains were intact and putative gene products were approximately full length76,77. The supplied O- and H- antigen serotype provided by ATCC were also confirmed bioinformatically using the SerotypeFinder v1.1 tool located at the Center for Genomic Epidemiology website59.
Weaned piglet challenge model of E. coli 62-57nal
Animals and facilities
A group of 24 weaned barrows approximately 21 days of age (Landrace × Large White, initial mean BW 6.35 kg) were collected on the day of their weaning and housed at SPARC in College Station, TX. Pigs were randomly assigned to pens (4.634 m2) with 2 barrows per pen that had solid concrete flooring and was equipped with a nipple waterer, rubber mat, and feeder. Genetic susceptibility to F18ac + ETEC infection was not determined in the present study. Pigs were provided ad libitum access to water and feed; the diet was a standard phase 1 nursery pig pelleted diet (S3 Table) formulated to meet or exceed the National Research Council (2012) recommended requirements of nutrients.
E. coli 62-57nal challenge trial
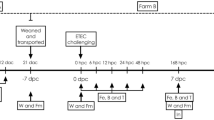
The pigs were held 2 d prior to the start of treatment in order to be pre-screened for endogenous enterotoxigenic E. coli (ETEC). Pigs were randomly allotted to one of three treatments: Non-challenged control, acute challenge (a single dose of 1.5 × 1010 CFU), and chronic challenge (an average daily dose of 5.4 × 108 CFU). Chronic daily doses ranged from 1.9 × 108 to 1.6 × 109 CFU with a median of 3.4 × 108 CFU. After random assignment of piglets to pens and treatments, researchers were unblinded in administration of the different treatments, and samples from each treatment were handled and collected uniformly. Each treatment had a total of 4 pens, 2 pigs per pen, for n = 8 pigs total. All pigs were housed in the same barn, with pens separated by empty pens to prevent cross-contamination between treatments. Additionally, disinfectant foot baths, routine cleaning of pens, disinfection of sampling equipment between treatment groups were used to prevent cross-contamination, and pig/pen sampling and dosing was performed in an order to reduce chance of contamination. Pens were routinely cleaned to further reduce sources of potential cross-contamination. E. coli 62-57nal was cultured overnight (16–18 h) in Tryptic Soy Broth (TSB; B-D Bacto) at 37 ºC with aeration. The acute treatment received a single dose of 6 ml of overnight E. coli culture on d 1 and the chronic treatment received a daily dosage of 6 ml of a tenfold dilution in phosphate buffered saline (PBS, Corning Cellgro) starting on d 1 until the termination of the trial. Pigs and feeders were weighed on d 0, 1, 3, and 6 for calculation of average daily gain (ADG), average daily feed intake (ADFI), and gain to feed (G:F). The d 1 collection of weight data, feces, and blood was approximately 12 h after the initial E. coli dose was administered. All animals were humanely euthanized and necropsied on d 7 for collection of intestinal scrapings and sections which were used for analysis of E. coli colonization and determination of villi blunting, respectively.
Pre-screening of animals for endogenous phage and ETEC
Prior to the trial initiation (d −1), pigs were screened for endogenous pathogenic E. coli, E. coli phages, and enteric bacteria capable of growing in the presence of 50 µg/ml nalidixic acid. Briefly, approximately 1 g of feces from each pen was mixed with 9 ml PBS, vortexed until homogenous, serially diluted (tenfold increments), and plated onto both MacConkey agar plates and MacConkey + nal. A chloroformed aliquot of the first sample dilution was plated onto Tryptic Soy Agar (TSA) plates overlaid with 0.5% top agar (5 g Tryptone, 5 g NaCl, 500 ml dH2O, 0.5% w/v Agar) inoculated with 100 μl from an overnight culture of E. coli 62-57nal. MacConkey + nal plates were screened for breakthrough colonies and TSA plates were screened for plaque formation or zones of clearing to determine phage presence. After overnight incubation, 3 colonies were selected from MacConkey plates from each pen and mixed with 150 µl Tris EDTA (TE) buffer. Each sample was boiled for 10 min then centrifuged at 8,000×g for 2 min. These colonies were screened via multiplex PCR for: Universal stress protein A (uspA), heat-labile toxin (LTI), heat-stable enterotoxin I (STI), heat-stable enterotoxin II (STII), Shiga toxin type 1 (Stx1) and Shiga toxin type 2 (Stx2) using previously established and validated primers78,79. Positive bands of appropriate size were confirmed using individual primer sets and resultant PCR products were visualized on a 1.5% agarose gel with gel red (Biotium).
Fecal collection and determination of E. coli 62-57nal and fecal calprotectin within feces
A representative fecal sample was collected from each pen on d 1, 3, and 6 to determine fecal E. coli populations. The samples were collected in individual, sterile 50 ml conical tubes and transported on wet ice to the laboratory. These fecal samples were processed and diluted in the same manner as described above, and were also spot plated (20 µl) on MacConkey agar containing 50 µg/ml nalidixic acid which were incubated for 10–12 h at 37 °C to avoid colonies merging within the spots. Fecal calprotectin was determined by a commercially available porcine ELISA kit (MyBioSource, San Diego, CA) with a minimum detection limit of 6.25 ng/ml with an intra-assay CV of less than 15%.
Blood sampling and serum analysis
Blood samples were collected from 2 pigs per pen on d 6. Blood was collected from the cranial vena cava via a 20 gauge needle and a 10 mL serum vacutainer tube (BD, with clot activator and gel for serum separation). Tubes were inverted and allowed a minimum of 30 min to clot. Samples were centrifuged at 1,600×g for 10 min at 2 °C, and the separated serum samples were stored at −80 °C until analysis was performed. Serum concentrations of interleukin 6 (IL-6) and interleukin 8 (IL-8) were determined via porcine ELISA kits (R&D Systems, Minneapolis, MN). The minimum detection for IL-6 was 18.8 pg/ml and 62.5 pg/ml for IL-8. Assays were conducted as outlined by the manufacturer.
Intestinal sampling and histology
Pigs were humanely euthanized, necropsied, and had samples collected for intestinal histology and E. coli populations from the duodenum, jejunum, ileum, cecum, and colon. Segments of the small intestine (duodenum, jejunum, and ileum) and large (cecum and colon) intestine were tied off with plastic zip ties to prevent cross contamination. Adherent bacterial samples were collected by rinsing the intestinal mucosal surface with sterile PBS, scraping a 2–3 cm section of the surface with a glass microscope slide, and then placing the sample into sterile 15 ml conical tubes containing 4.25 g of 2 mm glass beads (Walter Stern Inc.) and 8 ml of sterile PBS. Tissue scraping samples were vortexed for 5 min at 3000 rpm on a platform vortex mixer to homogenize samples. Cecum and distal colon contents were also collected and homogenized by thoroughly vortexing 0.5–5 g sample in 25 ml of PBS. Sample homogenates were serially diluted (tenfold increments) in PBS and spot-plated (20 ml) to MacConkey agar with 50 µg/ml nalidixic acid. All bacterial population estimates were normalized to the initial sample wet weight.
At the time of the necropsy, histopathologists were blinded for both tissue sample collection and examination of fixed tissue for adherent bacteria as well as evidence of tissue inflammation. For histology, the distal end of each intestinal segment (duodenum, jejunum, ileum, cecum and colon) directly adjacent to the section used for bacterial sampling was collected. Samples were positioned onto a 5.08 cm × 5.08 cm cardboard sections and secured with small clips to prevent tissue curling. Consecutive tissue samples were fixed in Carnoy’s solution (60% ethanol, 30% chloroform, 10% glacial acetic acid) at a 20:1 ratio for 30–45 min and 10% neutral buffered formalin (VWR scientific) at a 10:1 ratio for 24 h, followed by storage in 70% Ethanol until further processing. Tissues were trimmed into longitudinal sections of 5 mm width and embedded into paraffin using standard procedures80. After processing, 4 µm sections were placed on slides and stained with hematoxylin and eosin and evaluated histologically in a treatment blinded fashion. The slides were analyzed by a board-certified pathologist for rod attachment, presence of inflammation, and morphological changes (i.e., villous blunting, epithelial erosion). A total of 12 pigs, 4 from each treatment, were randomly selected for the evaluation of villus length, and well-oriented and intact villi were measured from the lamina muscularis mucosae layer to villus tip.
Analysis of fecal metabolites
Fecal samples collected from the floors of pens on d −1, 1, 3 and 6 were lyophilized and sent to the West Coast Metabolomics Center at the University of California Davis for untargeted metabolomic analysis. Untargeted GC-TOF profiling was performed following previously published parameters81. The resultant dataset is available on Metabolomics Workbench 82, under study number ST001041.
16S qPCR
For the purposes of quantifying select bacterial populations, microbial DNA was extracted from 50 mg of lyophilized feces using the Zymo Quick-DNA Fecal/Soil Microbe Kits following the manufacturer's instructions. Five ng of DNA was used to amplify 16S regions of select bacterial groups using Biorad SsoFast EvaGreen Supermix using reaction conditions described previously83 and family/genus/species specific primers described previously83,84. qPCR data is reported as log10 of starting 16S copy number per 5 ng of DNA isolated. Specific primer sets used were for Universal, Faecalibacterium, Streptococcus, E. coli, Fusobacterium, Firmicutes, Bacteroidetes, Lactobacillus, Ruminoccaceae, and Enterococcus.
Statistical analysis
Growth performance along with cytokine and intestinal bacterial population data were analyzed using the PROC MIXED procedure in SAS 9.3 (SAS Inst. Inc., Cary, NC). The model fixed effect was treatment with pen set as a random effect for growth performance, cytokine and intestinal bacterial population data. Fecal samples were collected on a pen basis, therefore pen was not included as a random effect. Calprotectin levels and fecal colony counts were analyzed as repeated measures using the PROC GLIMMIX procedure. Treatment, day, and treatment × day served as fixed effects. Day of collection also served as the repeated measure with pen as the subject. Metabolite data was normalized to sum, mean-centered and divided by the standard deviation of each variable, and analyzed for significant or trending metabolites between treatments using MetaboAnalyst version 4.541. Comparison of qPCR LOGSQ values was carried out using JMP Version 13 (SAS Inst. Inc., Cary, NC.). qPCR treatment means were compared pairwise on a per time point basis; many of the datasets did not pass the Shapiro–Wilk test for normality and were of small sample size, therefore treatments were compared using the nonparametric Wilcoxon Exact Test. qPCR data was also considered using multivariate methods on a pre/post treatment basis using principal component analysis (PCA). Results were considered significant at P ≤ 0.05 and marginally significant between P > 0.05 and P ≤ 0.10.
Data availability
Data supporting the findings of this study are available within this article and/or supplementary materials, additional supplements can be made available from corresponding author upon reasonable request. Data pertaining to resultant metabolomic analysis is available at Metabolomics Workbench, under study number ST001041. Genomic sequencing data available from Genbank under Bioproject (PRDF00000000).
References
Rhouma, M., Fairbrother, J. M., Beaudry, F. & Letellier, A. Post weaning diarrhea in pigs: risk factors and non-colistin-based control strategies. Acta Vet. Scand. 59, 31. https://doi.org/10.1186/s13028-017-0299-7 (2017).
Fairbrother, J. M. & Nadeau, É. in Diseases of Swine 807–834 (2019).
Fairbrother, J. M., Nadeau, É. & Gyles, C. L. Escherichia coli in postweaning diarrhea in pigs: an update on bacterial types, pathogenesis, and prevention strategies. Anim. Health Res. Rev. 6, 17–39. https://doi.org/10.1079/ahr2005105 (2007).
Imberechts, H., De Greve, H. & Lintermans, P. The pathogenesis of edema disease in pigs. A review. Vet. Microbiol. 31, 221–233. https://doi.org/10.1016/0378-1135(92)90080-D (1992).
Frydendahl, K. Prevalence of serogroups and virulence genes in Escherichia coli associated with postweaning diarrhoea and edema disease in pigs and a comparison of diagnostic approaches. Vet. Microbiol. 85, 169–182. https://doi.org/10.1016/S0378-1135(01)00504-1 (2002).
Jones, G. W. & Rutter, J. M. Role of the K88 antigen in the pathogenesis of neonatal diarrhea caused by Escherichia coli in piglets. Infect. Immun. 6, 918–927 (1972).
Isaacson, R. E. Enteric bacterial pathogens, villus atrophy and microbial growth. Veterinary Quarterly 20, 68–72. https://doi.org/10.1080/01652176.1998.9694973 (1998).
Berberov, E. M. et al. Relative importance of heat-labile enterotoxin in the causation of severe diarrheal disease in the gnotobiotic piglet model by a strain of enterotoxigenic Escherichia coli That produces multiple enterotoxins. Infect. Immun. 72, 3914–3924. https://doi.org/10.1128/IAI.72.7.3914-3924.2004 (2004).
Wellock, I. J., Fortomaris, P. D., Houdijk, J. G. M. & Kyriazakis, I. Effects of dietary protein supply, weaning age and experimental enterotoxigenic Escherichia coli infection on newly weaned pigs: Performance. Animal 2, 825–833. https://doi.org/10.1017/S1751731108001559 (2008).
Wassenaar, T. M. & Gunzer, F. The prediction of virulence based on presence of virulence genes in E. coli may not always be accurate. Gut Pathogens 7, 15. https://doi.org/10.1186/s13099-015-0062-4 (2015).
Zhang, W. et al. Significance of heat-stable and heat-labile enterotoxins in porcine colibacillosis in an additive model for pathogenicity studies. Infect Immun 74, 3107–3114. https://doi.org/10.1128/IAI.01338-05 (2006).
Bhandari, S. K., Xu, B., Nyachoti, C. M., Giesting, D. W. & Krause, D. O. Evaluation of alternatives to antibiotics using an Escherichia coli K88+ model of piglet diarrhea: Effects on gut microbial ecology1. J. Anim. Sci. 86, 836–847. https://doi.org/10.2527/jas.2006-822 (2008).
Becker, S. L. et al. Effects of an F18 enterotoxigenic Escherichia coli challenge on growth performance, immunological status, and gastrointestinal structure of weaned pigs and the potential protective effect of direct-fed microbial blends. J. Anim. Sci. 98, skaa113. https://doi.org/10.1093/jas/skaa113 (2020).
Madec, F. et al. Experimental models of porcine post-weaning colibacillosis and their relationship to post-weaning diarrhoea and digestive disorders as encountered in the field. Vet. Microbiol. 72, 295–310. https://doi.org/10.1016/S0378-1135(99)00202-3 (2000).
Casewell, M., Friis, C., Marco, E., McMullin, P. & Phillips, I. The European ban on growth-promoting antibiotics and emerging consequences for human and animal health. J. Antimicrob. Chemother. 52, 159–161. https://doi.org/10.1093/jac/dkg313 (2003).
Owusu-Asiedu, A., Nyachoti, C. M. & Marquardt, R. R. Response of early-weaned pigs to an enterotoxigenic Escherichia coli (K88) challenge when fed diets containing spray-dried porcine plasma or pea protein isolate plus egg yolk antibody, zinc oxide, fumaric acid, or antibiotic. J. Anim. Sci. 81, 1790–1798. https://doi.org/10.2527/2003.8171790x (2003).
Jones, P. H., Roe, J. M. & Miller, B. G. Effects of stressors on immune parameters and on the faecal shedding of enterotoxigenic Escherichia coli in piglets following experimental inoculation. Res. Vet. Sci. 70, 9–17. https://doi.org/10.1053/rvsc.2000.0436 (2001).
Frydendahl, K., Kåre Jensen, T., Strodl Andersen, J., Fredholm, M. & Evans, G. Association between the porcine Escherichia coli F18 receptor genotype and phenotype and susceptibility to colonisation and postweaning diarrhoea caused by E. coli O138:F18. Vet. Microbiol. 93, 39–51. https://doi.org/10.1016/s0378-1135(02)00348-6 (2003).
Nadeau, É. et al. Efficacy of a single oral dose of a live bivalent E. coli vaccine against post-weaning diarrhea due to F4 and F18-positive enterotoxigenic E. coli. Vet. J. 226, 32–39. https://doi.org/10.1016/j.tvjl.2017.07.004 (2017).
Arango Duque, G. & Descoteaux, A. Macrophage Cytokines: Involvement in Immunity and Infectious Diseases. Front. Immunol. 5, 491. https://doi.org/10.3389/fimmu.2014.00491 (2014).
Grimm, M. C., Elsbury, S. K., Pavli, P. & Doe, W. F. Interleukin 8: cells of origin in inflammatory bowel disease. Gut 38, 90–98. https://doi.org/10.1136/gut.38.1.90 (1996).
Gabay, C. Interleukin-6 and chronic inflammation. Arthritis Res Ther 8(Suppl 2), S3–S3. https://doi.org/10.1186/ar1917 (2006).
Scheller, J., Chalaris, A., Schmidt-Arras, D. & Rose-John, S. The pro- and anti-inflammatory properties of the cytokine interleukin-6. Biochim. Biophys. Acta Mol. Cell Res. 878–888, 2011. https://doi.org/10.1016/j.bbamcr.2011.01.034 (1813).
Guerrant, R. L., DeBoer, M. D., Moore, S. R., Scharf, R. J. & Lima, A. A. M. The impoverished gut—a triple burden of diarrhoea, stunting and chronic disease. Nat. Rev. Gastroenterol. Hepatol. 10, 220–229. https://doi.org/10.1038/nrgastro.2012.239 (2013).
Nyachoti, C. M., Kiarie, E., Bhandari, S. K., Zhang, G. & Krause, D. O. Weaned pig responses to Escherichia coli K88 oral challenge when receiving a lysozyme supplement. J Anim Sci 90, 252–260. https://doi.org/10.2527/jas.2010-3596 (2012).
Fossum, C., Wattrang, E., Fuxler, L., Thorleif Jensen, K. & Wallgren, P. Evaluation of various cytokines (IL-6, IFN-α, IFN-γ, TNF-α) as markers for acute bacterial infection in swine - a possible role for serum interleukin-6. Vet. Immunol. Immunopathol. 64, 161–172. https://doi.org/10.1016/S0165-2427(98)00126-3 (1998).
Lee, C. Y., Kim, S. J., Park, B. C. & Han, J. H. Effects of dietary supplementation of bacteriophages against enterotoxigenic Escherichia coli (ETEC) K88 on clinical symptoms of post-weaning pigs challenged with the ETEC pathogen. J. Anim. Physiol. Anim. Nutr. (Berl.) 101, 88–95. https://doi.org/10.1111/jpn.12513 (2017).
McLamb, B. L., Gibson, A. J., Overman, E. L., Stahl, C. & Moeser, A. J. Early weaning stress in pigs impairs innate mucosal immune responses to enterotoxigenic E. coli challenge and exacerbates intestinal injury and clinical disease. PLoS ONE 8, e59838. https://doi.org/10.1371/journal.pone.0059838 (2013).
Brown, E. M. et al. Diet and specific microbial exposure trigger features of environmental enteropathy in a novel murine model. Nat. Commun. 6, 7806. https://doi.org/10.1038/ncomms8806 (2015).
Roseth, A. G., Aadland, E., Jahnsen, J. & Raknerud, N. Assessment of disease activity in ulcerative colitis by faecal calprotectin, a novel granulocyte marker protein. Digestion 58, 176–180. https://doi.org/10.1159/000201441 (1997).
Costa, F. et al. Role of faecal calprotectin as non-invasive marker of intestinal inflammation. Dig. Liver Dis. 35, 642–647 (2003).
Barbosa, J. A. et al. Experimental infectious challenge in pigs leads to elevated fecal calprotectin levels following colitis, but not enteritis. Porcine Health Manag. 7, 48. https://doi.org/10.1186/s40813-021-00228-9 (2021).
Splichal, I., Fagerhol, M. K., Trebichavsky, I., Splichalova, A. & Schulze, J. The effect of intestinal colonization of germ-free pigs with Escherichia coli on calprotectin levels in plasma, intestinal and bronchoalveolar lavages. Immunobiology 209, 681–687. https://doi.org/10.1016/j.imbio.2004.09.009 (2005).
Xiao, D. et al. Effects of chitosan on intestinal inflammation in weaned pigs challenged by enterotoxigenic Escherichia coli. PLoS ONE 9, e104192. https://doi.org/10.1371/journal.pone.0104192 (2014).
Gao, Y. et al. Changes in gut microbial populations, intestinal morphology, expression of tight junction proteins, and cytokine production between two pig breeds after challenge with Escherichia coli K88: A comparative study1. J. Anim. Sci. 91, 5614–5625. https://doi.org/10.2527/jas.2013-6528 (2013).
Watanabe, K. & Petri, W. A. Jr. Environmental enteropathy: Elusive but significant subclinical abnormalities in developing countries. EBioMedicine 10, 25–32. https://doi.org/10.1016/j.ebiom.2016.07.030 (2016).
McCracken, B. A., Spurlock, M. E., Roos, M. A., Zuckermann, F. A. & Gaskins, H. R. Weaning anorexia may contribute to local inflammation in the piglet small intestine. J. Nutr. 129, 613–619. https://doi.org/10.1093/jn/129.3.613 (1999).
Dou, S. et al. Characterisation of early-life fecal microbiota in susceptible and healthy pigs to post-weaning diarrhoea. PLoS ONE 12, e0169851. https://doi.org/10.1371/journal.pone.0169851 (2017).
Chen, L. et al. The maturing development of gut microbiota in commercial piglets during the weaning transition. Front. Microbiol. 8, 1688. https://doi.org/10.3389/fmicb.2017.01688 (2017).
Montagne, L. et al. Main intestinal markers associated with the changes in gut architecture and function in piglets after weaning. Br. J. Nutr. 97, 45–57. https://doi.org/10.1017/S000711450720580X (2007).
Xia, J. & Wishart, D. S. Using MetaboAnalyst 3.0 for comprehensive metabolomics data analysis. Curr Protoc. Bioinform. 55, 141011–141091. https://doi.org/10.1002/cpbi.11 (2016).
Gong, W. et al. Serum Metabolomic Profiling of Piglets Infected with Virulent Classical Swine Fever Virus. Front. Microbiol. 8, 731. https://doi.org/10.3389/fmicb.2017.00731 (2017).
Li, Y. et al. Intestinal microbiome-metabolome responses to essential oils in piglets. Front. Microbiol. 9, 1988. https://doi.org/10.3389/fmicb.2018.01988 (2018).
Carmelo, V. A. O., Banerjee, P., da Silva Diniz, W. J. & Kadarmideen, H. N. Metabolomic networks and pathways associated with feed efficiency and related-traits in Duroc and Landrace pigs. Sci. Rep. 10, 255. https://doi.org/10.1038/s41598-019-57182-4 (2020).
Banerjee, P., Carmelo, V. A. O. & Kadarmideen, H. N. Integrative analysis of metabolomic and transcriptomic profiles uncovers biological pathways of feed efficiency in pigs. Metabolites 10, 275. https://doi.org/10.3390/metabo10070275 (2020).
Chang, H.-W. et al. Gut microbiome contributions to altered metabolism in a pig model of undernutrition. Proc. Natl. Acad. Sci. U.S.A. 118, e2024446118. https://doi.org/10.1073/pnas.2024446118 (2021).
He, Y., Liu, Y. & Ji, P. Metabolomic profile of weaned pigs challenged with E. coli and supplemented with Carbadox or Bacillus subtilis. Metabolites 11, 2008. https://doi.org/10.3390/metabo11020081 (2021).
Sugiharto, S., Hedemann, M. S. & Lauridsen, C. Plasma metabolomic profiles and immune responses of piglets after weaning and challenge with E. coli. J. Anim. Sci. Biotechnol. 5, 17. https://doi.org/10.1186/2049-1891-5-17 (2014).
Wu, H. et al. Jejunal metabolic responses to Escherichia coli infection in piglets. Front. Microbiol. 9, 02465. https://doi.org/10.3389/fmicb.2018.02465 (2018).
Ren, W. et al. Metabolomics study of metabolic variations in enterotoxigenic Escherichia coli-infected piglets. RSC Adv. 5, 59550–59555. https://doi.org/10.1039/C5RA09513A (2015).
Wishart, D. S. et al. HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Res. 46, D608–D617. https://doi.org/10.1093/nar/gkx1089 (2018).
Goldansaz, S. A. et al. Livestock metabolomics and the livestock metabolome: A systematic review. PLoS ONE 12, e0177675. https://doi.org/10.1371/journal.pone.0177675 (2017).
Fothergill, J. C. & Guest, J. R. Catabolism of L-lysine by Pseudomonas aeruginosa. J. Gen. Microbiol. 99, 139–155. https://doi.org/10.1099/00221287-99-1-139 (1977).
Le Gall, G. et al. Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J. Proteome Res. 10, 4208–4218. https://doi.org/10.1021/pr2003598 (2011).
Jansson, J. et al. Metabolomics reveals metabolic biomarkers of Crohn’s disease. PLoS ONE 4, e6386. https://doi.org/10.1371/journal.pone.0006386 (2009).
Bjerrum, J. T. et al. Metabonomics of human fecal extracts characterize ulcerative colitis, Crohn’s disease and healthy individuals. Metabolomics 11, 122–133. https://doi.org/10.1007/s11306-014-0677-3 (2015).
Byersdorfer, C. A. The role of Fatty Acid oxidation in the metabolic reprograming of activated t-cells. Front. Immunol. 5, 641. https://doi.org/10.3389/fimmu.2014.00641 (2014).
Namgaladze, D. & Brune, B. Macrophage fatty acid oxidation and its roles in macrophage polarization and fatty acid-induced inflammation. Biochim. Biophys. Acta 1861, 1796–1807. https://doi.org/10.1016/j.bbalip.2016.09.002 (2016).
Joensen, K. G., Tetzschner, A. M. M., Iguchi, A., Aarestrup, F. M. & Scheutz, F. Rapid and easy in silico serotyping of Escherichia coli isolates by use of whole-genome sequencing data. J. Clin. Microbiol. 53, 2410–2426. https://doi.org/10.1128/jcm.00008-15 (2015).
Ewingwh, T. H. & Davis, B. R. Escherichia coli serotypes associated with edema disease of swine. Cornell Vet. 48, 201–206 (1958).
Mainil, J. Escherichia coli virulence factors. Vet. Immunol. Immunopathol. 152, 2–12. https://doi.org/10.1016/j.vetimm.2012.09.032 (2013).
Moxley, R. A. et al. Pathogenicity of an enterotoxigenic Escherichia coli hemolysin (hlyA) mutant in gnotobiotic piglets. Infect. Immun. 66, 5031–5035 (1998).
Mirsepasi-Lauridsen, H. C. et al. Secretion of alpha-hemolysin by Escherichia coli disrupts tight junctions in ulcerative colitis patients. Clin. Transl. Gastroenterol. 7, e149. https://doi.org/10.1038/ctg.2016.3 (2016).
Bucker, R. et al. alpha-Haemolysin of Escherichia coli in IBD: A potentiator of inflammatory activity in the colon. Gut 63, 1893–1901. https://doi.org/10.1136/gutjnl-2013-306099 (2014).
Moxley, R. A. Edema disease. Vet. Clin. North Am Food Anim. Pract. 16, 175–185. https://doi.org/10.1016/s0749-0720(15)30142-0 (2000).
Cai, Y., Martínez, Y., Li Qiang Yao, Y. & Yu, L. Review of enterotoxigenic Escherichia coli enterotoxins. Vol. 4 (2016).
Zhang, W., Zhao, M., Ruesch, L., Omot, A. & Francis, D. Prevalence of virulence genes in Escherichia coli strains recently isolated from young pigs with diarrhea in the US. Vet. Microbiol. 123, 145–152. https://doi.org/10.1016/j.vetmic.2007.02.018 (2007).
Mills, M. & Payne, S. M. Genetics and regulation of heme iron transport in Shigella dysenteriae and detection of an analogous system in Escherichia coli O157:H7. J. Bacteriol. 177, 3004–3009 (1995).
Field, M. Intestinal ion transport and the pathophysiology of diarrhea. J. Clin. Investig. 111, 931–943. https://doi.org/10.1172/jci200318326 (2003).
Guan, Q. A comprehensive review and update on the pathogenesis of inflammatory bowel disease. J. Immunol. Res. 2019, 7247238. https://doi.org/10.1155/2019/7247238 (2019).
Jiminez, J. A., Uwiera, T. C., Douglas Inglis, G. & Uwiera, R. R. Animal models to study acute and chronic intestinal inflammation in mammals. Gut Pathog. 7, 29. https://doi.org/10.1186/s13099-015-0076-y (2015).
Walana, W. et al. IL-8 antagonist, CXCL8(3–72)K11R/G31P coupled with probiotic exhibit variably enhanced therapeutic potential in ameliorating ulcerative colitis. Biomed. Pharmacother. 103, 253–261. https://doi.org/10.1016/j.biopha.2018.04.008 (2018).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Tatusova, T. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 44, 6614–6624. https://doi.org/10.1093/nar/gkw569 (2016).
Chen, L., Zheng, D., Liu, B., Yang, J. & Jin, Q. VFDB 2016: hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res. 44, D694–D697. https://doi.org/10.1093/nar/gkv1239 (2016).
Camacho, C. et al. BLAST+: Architecture and applications. BMC Bioinform. 10, 421. https://doi.org/10.1186/1471-2105-10-421 (2009).
Finn, R. D. et al. InterPro in 2017—Beyond protein family and domain annotations. Nucleic Acids Res. 45, D190–D199. https://doi.org/10.1093/nar/gkw1107 (2017).
Fagan, P. K., Hornitzky, M. A., Bettelheim, K. A. & Djordjevic, S. P. Detection of Shiga-Like Toxin (stx1 and stx2), Intimin (eaeA), and Enterohemorrhagic Escherichia coli (EHEC) Hemolysin (EHEC hlyA) Genes in Animal Feces by Multiplex PCR. Appl. Environ. Microbiol. 65, 868–872 (1999).
Osek, J. Multiplex polymerase chain reaction assay for identification of enterotoxigenic Escherichia coli strains. J. Vet. Diagn. Invest. 13, 308–311 (2001).
Blick, A. K. et al. Comparison of 2 fixatives in the porcine colon for in situ microbiota studies. J. Anim. Sci. 97, 4803–4809. https://doi.org/10.1093/jas/skz325 (2019).
Fiehn, O. et al. Quality control for plant metabolomics: reporting MSI-compliant studies. Plant J. 53, 691–704. https://doi.org/10.1111/j.1365-313X.2007.03387.x (2008).
Sud, M. et al. Metabolomics Workbench: An international repository for metabolomics data and metadata, metabolite standards, protocols, tutorials and training, and analysis tools. Nucleic Acids Res. 44, D463-470. https://doi.org/10.1093/nar/gkv1042 (2016).
Suchodolski, J. S. et al. The fecal microbiome in dogs with acute diarrhea and idiopathic inflammatory bowel disease. PLoS ONE 7, e51907. https://doi.org/10.1371/journal.pone.0051907 (2012).
Walter, J. et al. Detection of Lactobacillus, Pediococcus, Leuconostoc, and Weissella species in human feces by using group-specific PCR primers and denaturing gradient gel electrophoresis. Appl. Environ. Microbiol. 67, 2578–2585. https://doi.org/10.1128/AEM.67.6.2578-2585.2001 (2001).
Acknowledgements
This work was supported, in whole or in part, by the Bill & Melinda Gates Foundation [OPP1139800]. Under the grant conditions of the Foundation, a Creative Commons Attribution 4.0 Generic License has already been assigned to the Author Accepted Manuscript version that might arise from this submission. This work was additionally supported by Texas A&M AgriLife Research. The authors would like to thank Amanda Blake and So Young Park for assistance with qPCR. We would also like to thank Justin Leavitt, Jacob Chamblee, Jacob Lancaster, Lauren Lessor, Ruoyan Luo, Shayna Smith, Caitlin Older, Jeann Leal de Araujo, Kara Dunmire, Logan Joiner, and Lily Hernandez for their assistance with animal necropsy and sample preparation.
Author information
Authors and Affiliations
Contributions
All authors assisted in necropsy at the end of the trial period, the resulting data acquisition, and reviewed the manuscript. Specifically, J.X.B. and S.S. (co-first authors) prepared the manuscript, performed experiments, analyzed data, and assisted in experimental design. A.M.K provided additional support in experiments and orchestration of the piglet trial. C.P. and J.J.G. conceptual and experimental design. J.S.S. resources and guidance on qPCR experiments and metabolomics. K.G. and T.C. facilities, resources, and animal housing. R.R.R, P.R.G. and A.K.B. clinical histopathology and necropsy of piglets.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boeckman, J.X., Sprayberry, S., Korn, A.M. et al. Effect of chronic and acute enterotoxigenic E. coli challenge on growth performance, intestinal inflammation, microbiome, and metabolome of weaned piglets. Sci Rep 12, 5024 (2022). https://doi.org/10.1038/s41598-022-08446-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-08446-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.