Abstract
Changes in the oral microbiome, particularly Fusobacterium nucleatum, are associated with oral squamous cell carcinoma (OSCC). F. nucleatum has been reported to modulate local immunity in cancers. We aimed to assess the association between intratumoral F. nucleatum and clinico-pathological features, relapse, and overall survival (OS) in two independent cohorts of patients with OSCC, and to explore the interplay with immune-related genes. We retrospectively analyzed tissue samples from a first cohort of 122 patients with head and neck squamous cell carcinoma, including 61 OSCC (cohort #1), and a second cohort of 90 additional OSCC (cohort #2). We then performed a sensitivity analysis on the merged cohort of OSCC patients (N = 151). F. nucleatum 16S rRNA gene sequences were quantified using real-time quantitative PCR. The presence of gram-negative bacteria and macrophages was confirmed by LPS and CD163 immunostainings, respectively. F. nucleatum positivity was associated with older age, less alcohol and combined alcohol plus tobacco consumption, and less frequent lymph node invasion. There was a trend for a lower recurrence rate in F. nucleatum-positive cases, with less metastatic relapses compared to F. nucleatum-negative tumors, and significantly longer OS, relapse-free and metastasis-free survival. F. nucleatum status was independently associated with OS in multivariate analysis. Immune-related gene and immunohistochemistry analyses showed that gram-negative bacteria load inversely correlated with M2 macrophages. F. nucleatum-associated OSCC has a specific immune microenvironment, is more frequent in older, non-drinking patients, and associated with a favorable prognosis.
Similar content being viewed by others
Introduction
Head and neck cancers encompass a variety of tumors originating from the oral cavity, hypopharynx, oropharynx, nasopharynx, or larynx1. It is the sixth most common malignancy worldwide, accounting for approximately 5% of all cancer cases and of all cancer deaths1, 2. Squamous cell carcinoma is the most frequent histological type, representing more than 90% of these tumors3. Classical risk factors for head and neck squamous cell carcinoma (HNSCC) include tobacco and alcohol consumption, as well as human papillomavirus (HPV) infection that has been demonstrated to have a favorable prognostic impact and better response to treatments4, 5. Smoking-related HNSCCs demonstrate near universal loss-of-function TP53 mutations, which are associated with shorter survival and resistance to radiotherapy and chemotherapy6. No relevant biomarkers for tailored therapeutic strategies have been identified in HNSCC to date1.
Oral squamous cell carcinoma (OSCC) is the most common subset of HNSCC2. Nearly two-third of cases are attributed to tobacco smoking and alcohol consumption, and unlike cancers of the oropharynx, only a small fraction of OSCC cases (approximately 4%) are related to HPV infection (mostly, HPV16)4, 7.
The human body is inhabited by over 100 trillion microbial cells living in symbiosis with their host8. The term microbiome stands for “the collective genomes and gene products of all microbes residing within an organism”8. Most of these bacteria are located in the gut. Individual bacteria and shifts in microbiome composition are associated with human disease, including cancer9, 10. Hence, the oral bacterial flora plays an essential role in maintaining a healthy oral physiological environment and changes in the oral microbiome have been associated with cancer11. Recently, it has been demonstrated that bacteria are also found within tumoral tissue, so called “intratumoral” microbiota12. Periodontal pathogens including Fusobacterium nucleatum (F. nucleatum) are a risk factor for OSCC, independent from tobacco, alcohol, and HPV11, 13, and their abundance in saliva samples increases with tumor progression14, 15. Little is known about the association between intratumoral F. nucleatum and HNSCC tumor biology16, 17. Moreover, F. nucleatum has been reported to modulate local immunity in other cancers (mainly, colorectal cancers), particularly macrophages and T regulatory cells (Tregs), via Toll like receptor (TLR) 2–4 signaling18,19,20.
In this study, we assessed the association between intratumoral F. nucleatum and clinico-pathological features, relapse, and overall survival (OS) in two independent cohorts of patients with OSCC, and explored the interplay between F. nucleatum and immune-related genes.
Patients and methods
Patients and samples
We retrieved samples from patients with HNSCC who underwent upfront surgery at the Curie Institute between 1990 and 2006. We selected patients with complete clinical, histological, and biological data and long-term follow-up.
We used samples from two independent cohorts: the cohort #1 consisted of 122 patients with HNSCC from various primary sites: oral cavity (OSCC; n = 61), oropharynx (n = 22), hypopharynx (n = 17), and larynx (n = 22). The cohort #2 consisted of 90 OSCC patients. Finally, we grouped the OSCC patients from the first cohort with the second cohort for analysis (merged cohort). The flow chart of patients is displayed in Fig. 1.
The study was conducted in accordance with the ethics principles of the Declaration of Helsinki and GDPR regulations. In accordance with the French regulations, informed consent was obtained from all subjects. This study was approved by the French Ethical Committee (Agreement number D-750602, France) and the ethics committee of the Institut Curie (Agreement number C75-05-18).
Genomic DNA and total RNA extractions
Tumor samples were frozen in liquid nitrogen in a cryotube immediately after surgery and stored at − 80 °C, under temperature control.
A tumor fragment of 5–40 mg and 10–50 mg was used for DNA and RNA extraction, respectively. Tumoral cellularity was evaluated on a 4–5 µm cryosection and samples with less than 50% of tumoral cells were excluded from the study. Nucleic acid extraction was performed on macrodissected tumoral zones, according to the following protocols. Total genomic DNA was extracted from with phenol–chloroform after proteinase K digestion, followed by the precipitation of nucleic acids in ethanol. Total RNA was extracted with miRNeasy Mini kit Qiagen following supplier’s recommendations. The quality of RNA was verified by migration on agarose gel. Nucleic acids were quantified using Nanodrop spectrophotometer ND-1000 (ThermoScientific, Wilmington, DE, USA).
In order to rule out external contaminations for F. nucleatum analysis, we included negative controls (buffers/reagents without tumor samples) and the samples were manipulated under a hood and with mask and gloves.
TP53 and PIK3CA mutation status
Information about TP53 mutation status was extracted for the results of a Next Generation Sequencing (NGS) panel and PIK3CA mutation status was determined by High Resolution Melting (HRM) analysis in all samples (cohort #1 and cohort #2).
Fusobacterium nucleatum status analysis by real-time quantitative PCR
Quantitative values were obtained from the cycle number (Cycle Threshold, Ct value) at which the increase in the fluorescence signal associated with exponential growth of PCR products started to be detected. Detection is performed by the laser detector of the ABI Prism 7900HT Sequence Detection System (Applied Biosystems, ThermoFisher Scientific, Waltham, MA), using PE Biosystems analysis software according to the manufacturer’s manuals.
The precise amount of genomic DNA (based on optical density) and its quality (i.e. lack of extensive degradation) are both difficult to assess. We therefore normalized F. nucleatum levels on the basis of JUN contents, used as human diploid control. Results, expressed as N-fold differences in F. nucleatum 16S rRNA gene copy number and termed “Nfn,” were determined as Nfn = 2ΔCtsample, where the ΔCt value of the sample is determined by subtracting the average Ct value of the F. nucleatum 16S rRNA gene from the average Ct value of the diploid control gene. F. nucleatum Ct values lower than 32 were considered as F. nucleatum-positive and F. nucleatum Ct values higher than 32 were considered as F. nucleatum-negative. For the F. nucleatum-positive tumors, the Nfn values were normalized such that a Ct value of 32 was equal to 1, i.e. the smallest accurately quantifiable amount of F. nucleatum 16S rRNA gene copy number.
Primers for F. nucleatum (targeting the 16S rRNA gene DNA sequence, Upper 5′-AGC GTT TGA CAT CTT AGG AAT GA-3′ and Lower 5′-GCC ATG CAC CAC CTG TCT T-3′) and JUN (Genbank accession number NM_002228, Upper 5′-CAC GGC CAA CAT GCT CAG G-3′ and Lower 5′-GCA TGA GTT GGC ACC CAC TGT-3′) were designed with the assistance of Oligo 6.0 computer program (National Biosciences, Plymouth, MN). The primer pair for F. nucleatum was selected to be unique relative to the sequences of closely related family member 16S rRNA genes of the Fusobacterium species. Agarose gel electrophoresis and DNA sequencing were used to verify the specificity of the F. nucleatum PCR amplicon.
Real-time quantitative PCR was performed using the Power SYBR Green master mix (ThermoFisher Scientific, Waltham, MA). The thermal cycling conditions were an initial denaturation step at 95 °C for 10 min, followed by 50 cycles at 95 °C for 15 s and 65 °C/60 °C for 1 min for JUN/F. nucleatum respectively, and then the melt curve steps. The amplifications specificity was confirmed by melting curve analysis.
Immune-related gene expression analysis by real-time quantitative reverse transcription PCR
The immune-related genes were selected in collaboration with immunologists from our group, consistent with our previous publications21, 22.
Immune-related gene expression levels were normalized on the basis of TBP contents (Genbank accession number NM_003194)23. We chose TBP, which encodes the TATA box binding protein, as endogenous control because the prevalence of its transcripts is moderate, and because there are no known TBP retropseudogenes (retropseudogenes lead to co-amplification of contaminating genomic DNA and thus interfere with RT‐PCR, despite the use of primers in separate exons)23. We had previously selected and used the same gene as endogenous control in different cancer types including HNSCC21, 24.
Results, expressed as N-fold differences in target gene expression and termed “Ntarget,” were determined as Ntarget = 2ΔCtsample, where the ΔCt value of the sample is determined by subtracting the average Ct value of the target gene from the average Ct value of the control gene.
Primers for immune-related genes were designed with the assistance of Oligo 6.0 computer program (National Biosciences, Plymouth, MN). We searched the dbEST and nr databases to confirm the total gene specificity of the nucleotide sequences chosen as primers and the absence of single nucleotide polymorphisms25. In particular, the primer pairs were selected to be unique relative to the sequences of closely related family member genes or of the corresponding retropseudogenes25. Agarose gel electrophoresis was used to verify the specificity of PCR amplicons. The nucleotide sequences of the oligonucleotide hybridization primers and the average Ct value are shown in Suppl. Table 1. Sixty-four genes mainly involved in the immune process were selected, in particular 19 checkpoint T cell and tumor cell genes, 16 chemokine genes and 18 immune cell population genes. For each primer pair we performed no-template control, no-RT control (RT negative) and RT control with genomic DNA assays, which produced negligible signals that were usually greater than 40 in Ct value, suggesting that primer-dimer formation and genomic DNA contamination effects were negligible.
RNA was reverse transcribed in a final volume of 20 µL containing 1X RT buffer (50 mM Tris–HCl (pH 8.3); 75 mM KCl; 3 mM MgCl2), 10 mM DTT, 100 units of SuperScript II Reverse Transcriptase with reduced RNase H activity (Life Technologies, Carlsbad, CA), 0.5 mM each dNTP (GE Healthcare, Chicago, IL), 0.15 µg/µL random primers (Life Technologies, Carlsbad, CA), 20 units of RNasin Ribonuclease Inhibitor (Promega, Madison, WI), and 1 µg of total RNA. The thermal cycling conditions were hybridization of random primers at 25 °C for 10 min, cDNA synthesis at 42 °C for 30 min, and reverse transcriptase inactivation at 99 °C for 5 min and cooling at 4 °C.
All of the real-time quantitative PCR reactions were performed using the Power SYBR Green master mix (ThermoFisher Scientific, Waltham, MA) using an ABI Prism 7900HT Sequence Detection System (Applied Biosystems, ThermoFisher Scientific, Waltham, MA). The thermal cycling conditions were an initial denaturation step at 95 °C for 10 min, followed by 50 cycles at 95 °C for 15 s and 65 °C for 1 min, and then the melt curve steps. The amplifications specificity was confirmed by melting curve analysis.
Immunohistochemistry
Immunohistochemistry assay was performed using CD163 (Novocastra Leica, ref.: CD163-L-CE, monoclonal Mouse, clone 10D6, pH9 1/1000, 30 m) and LPS (MyBioSource, ref.: MBS2111237, monoclonal Mouse, clone C8, pH6, 1/500, 60 m) antibodies. Paraffin-embedded tissue blocks, obtained at the time of the initial diagnosis, were retrieved from the archives of the Department of Diagnostic and Theranostic Medicine, Curie Institute. Sections of 3 µm in thickness were cut with a microtome from the paraffin-embedded tissue blocks of OSCCs. Tissue sections were deparaffinized and rehydrated through a series of xylene and ethanol washes. Briefly, key figures included: (1) antigen retrieval in 0.1 mol/L citrate buffer, pH 6 (BioCare, Pacheco, CA, USA) in a pressure cooker (4 min); (2) blocking of endogenous peroxidase activity by immersing sections in 3% hydrogen peroxide in methanol for 15 min and subsequently rinsing them in water and PBS; (3) incubation with primary antibodies against the targeted antigen; (4) immunodetection with a biotin-conjugated secondary antibody formulation that recognizes rabbit and mouse immunoglobulins, followed by peroxidase-labeled streptavidin and linking with a rabbit biotinylated antibody against mouse immunoglobulin G (DAKO SA) and (5) chromogenic revelation with DAB and counterstaining with Mayer’s hematoxylin26. All immunostainings were processed using a Leica BOND RX research automated immunostaining device. A semi quantitative score (intensity × frequency) was used for interpretation of immunostaining with anti-LPS antibody (0 = negative staining, 1 = weak staining, 2 = moderate staining and 3 = strong staining). Semi-quantitative evaluation of the tumoral and non-tumoral cells was performed (0: absence of cells, +: < 5%, ++: 5 to 25%, +++: > 25%). Percentage of macrophage immunostaining with anti-CD163 antibody was also evaluated semi-quantitatively using the following score Histologic Score (HS): HS 0: absence of positive cells; HS 1 (+): 1 < HS < 25%; HS 2 (++): 26 < HS < 50%; HS 3 (+++): HS > 50%.
Statistical methods
Relationships between F. nucleatum and clinical, biological, and pathological parameters were assessed by using the Chi-square, Chi-square with Yates correction or Fisher tests, as appropriate. Spearman rank correlation non-parametric test was used to determine relationships between F. nucleatum and immune-related genes levels. Bonferroni correction was applied to adjust for multiple tests. Differences were considered significant at confidence levels greater than 95% (p < 0.05).
Survival endpoints were defined according to the DATECAN consensus27. OS was determined from the time of initial diagnosis to the time of death, regardless of the cause of death. Relapse-free survival (RFS) was determined from the time of initial diagnosis to the time of relapse (locoregional and/or metastatic) or death, whichever occurred first, regardless of the cause of death. Metastasis-free survival (MFS) was determined from the time of initial diagnosis to the time of metastatic relapse or death, whichever occurred first, regardless of the cause of death. In the absence of event, patients were censored at the date of last follow-up. Survival distributions were estimated by the Kaplan–Meier method, and the significance of differences between survival rates were ascertained with the log-rank test. The multivariate Cox proportional hazards regression model was used to assess the prognostic significance of F. nucleatum and clinical markers on OS; parameters with p values < 0.05 in univariate analysis were entered into the final multivariable Cox regression model, after considering redundancy between variables. The results are presented as hazard ratios (HR) and 95% confidence intervals (95%CIs).
Results
Association of Fusobacterium nucleatum with clinico-pathological features and overall survival (OS) in the cohort #1
The first cohort (cohort #1) comprised 122 patients with untreated HNSCC from various primary sites, with a majority of tumors arising from the oral cavity (OSCC, n = 61, 50%). Patient characteristics are listed in Suppl. Table 2. Median age was 57 years (range 22–78). Most patients were male (76.2%), with alcohol (64.8%) or tobacco (71.3%) consumption or both (59.1%). Nine patients (7.4%) were HPV-positive. Pathological staging showed a high proportion of UICC stage IV tumors (50.8%). Seventy-two (59%) harbored a mutation in TP53 gene and 15 (12.3%) in PIK3CA gene.
Among the 122 HNSCC samples tested, 35 (28.7%) were scored F. nucleatum-negative and 87 (71.3%) were scored F. nucleatum-positive. Among the 87 F. nucleatum-positive tumors, major differences of Nfn values (determined as described in “Patients and Methods”) were observed, ranging from 1.0 to 1613 (Suppl. Fig. 1).
Association of patient characteristics with F. nucleatum status is displayed in Suppl. Table 2. F. nucleatum positivity was associated with an enrichment in female (28.7% vs. 11.4%, p = 0.042), non-drinking patients (42.4% vs. 13.6%, p = 0.014), with more tumors from the oral cavity and oropharynx (55.2% vs. 37.1%, and 21.8% vs. 8.6%, respectively) and less tumors from the hypopharynx (5.7% vs. 34.3%, p = 0.0003).
Alcohol (p = 0.019) and tobacco (p = 0.017) consumption, pT (p = 0.021) and pN (p = 0.045) and UICC (p = 0.013) stage, absence of HPV infection (p = 0.0061), presence of TP53 mutation (p = 0.0007), and distant (metastatic) relapse (p = 0.0085) were significantly associated with shorter OS in univariate analysis (Suppl. Table 3).
In this overall HNSCC population, there was a non-significant trend toward longer OS in patients with F. nucleatum-positive tumors (HR: 0.64, p = 0.092) (Fig. 2A). We analyzed the association of F. nucleatum status with OS according to tumor primary site (Fig. 2B,C and Suppl. Fig. 2). We observed that this trend was mainly driven by the oral cavity subgroup (HR: 0.51, p = 0.089) (Fig. 2B), while there was no significant association with OS in patients with HNSCC tumors from other primary sites (Fig. 2C and Suppl. Fig. 2).
Overall survival (OS) curves according to F. nucleatum status (A) in the overall head and neck squamous cell carcinoma (HNSCC) patient population (N = 122), (B) in the oral cavity tumor (OSCC) subgroup (N = 61) of cohort #1, (C) in the non-OSCC subgroup (N = 61) of cohort #1, (D) in cohort #2 (N = 90), and (E) in the merged cohort (N = 151).
Therefore, we decided to focus on the subset of patients with oral cavity tumors for the next steps. In this subgroup of 61 OSCC patients from cohort #1, F. nucleatum positivity (78.7%) was associated with lower pN stage (p = 0.043) and a non-significant trend for more non-drinkers (p = 0.092) compared to F. nucleatum-negative tumors (Suppl. Table 4). Among the clinical, biological and pathological parameters listed above, TP53 mutation (HR: 2.76, p = 0.0093) and UICC stage (HR: 2.79, p = 0.043 for stage II and HR: 2.70, p = 0.049 for stage IV) were significantly associated with OS (Table 1).
Association of Fusobacterium nucleatum with clinico-pathological features and OS in the cohort #2
To further explore the association of F. nucleatum with OS in patients with OSCC, we used an independent cohort composed of 90 patients with HNSCC tumors exclusively from this primary site (cohort #2). Patient characteristics and association with F. nucleatum status as displayed in Suppl. Table 5. F. nucleatum was detected in 76 (84.4%) tumors. Most OSCC patients were men (62.2%), ≥ 56 years (76.7%), non-drinkers (70.4%) but smokers (51.2%), with HPV-negative (96.7%), advanced (mainly, stage IV: 51.1%) tumors. F. nucleatum status was associated with patient age (p = 0.0036), pN stage (p = 0.028) and tumor grade (p = 0.040). Advanced UICC stage (HR: 2.44, p = 0.030 for stage IV), high pN stage (HR: 2.42, p = 0.045 for pN1, and HR: 3.03, p = 0.0010 for pN3), and grade (HR: 3.39, p = 0.031 for grade III, and tumor relapse (HR: 5.16, p < 0.0001) were associated with poor OS (Table 1). Similarly to the cohort #1, we observed a non-significant trend toward longer OS in the F. nucleatum-positive group (HR: 0.54, p = 0.066) (Fig. 2D).
Sensitivity analysis for the association of Fusobacterium nucleatum with OS in the merged cohort
In order to increase the study power, we performed a sensitivity analysis after grouping the two cohorts of patients with OSCC tumors (merged cohort, n = 151). Patients characteristics are described in Table 2. As previously observed, patients were mostly men (61.6%), ≥ 56 years (70.2%), non-drinkers (61.2%) but smokers (55.1%), with HPV-negative (97.4%), pN0 (53.6%), UICC stage IV (42.4%) tumors. In the merged cohort, 124 (82.1%) samples were positive for F. nucleatum (distribution of Nfn value displayed in Suppl. Figure 1), which was associated with older age (≥ 56 years: 74.2% vs. 51.9%, p = 0.021), fewer drinkers (34.9% vs. 60.0%, p = 0.034), and less frequent lymph node involvement (more pN0 tumors, 58.1% vs 33.3%, p = 0.0016) (Table 2). Positive pN status (p = 0.0040) and higher UICC stage (stage ≥ II, p = 0.0036), presence of TP53 mutation (HR: 2.13, p = 0.0033), and tumor relapse (HR: 2.75, p < 0.0001) were prognostic indicators (Table 1). In the merged cohort, the outcome of the patients with a F. nucleatum-positive tumor was significantly better than that of the 27 patients with a F. nucleatum-negative tumor in term of OS (HR: 0.51, p = 0.0094; 5-year OS 60.5% vs. 37.7%; 10-year OS 47.9% vs. 18.8%) (Fig. 2E).
Relapse-free (RFS) and metastasis-free survival (MFS) in the merged cohort
One hundred and fourty two patients were evaluable for disease relapse. There was a non-significant trend for less frequent disease recurrence in patients with F. nucleatum-positive tumors (35.5% vs. 51.8%, p = 0.11), with more locoregional (70.5% vs. 53.3%, p = 0.041) vs metastatic relapse compared to F. nucleatum-negative tumors (Table 2).
RFS and MFS curves according to F. nucleatum status are displayed in Fig. 3. F. nucleatum-positive tumors were associated with significantly longer RFS (median: 7.06 vs. 2.11 months, p = 0.0091) and MFS (9.71 vs. 3.54 months, p = 0.0016) compared to F. nucleatum-negative tumors.
Multivariate analysis of OS predictors and stratification according to pT and pN stage
Using a Cox proportional hazards model, we also assessed the prognostic value for OS of parameters that were significant in univariate analysis, i.e. pT, pN, UICC stage, TP53 mutation (Table 1) and F. nucleatum status (Fig. 2E). As pT/pN and UICC were redundant variables, the model was built with UICC stage, TP53 and F. nucleatum. The prognostic significance of these three parameters was maintained in Cox multivariate regression analysis (Table 3).
In addition, we assessed the prognostic value of F. nucleatum on OS after stratification on pT and pN as main prognostic factors in OSCC. Interestingly, the combinations of pT1/T2 and F. nucleatum positivity (n = 49) and pN0 and F. nucleatum positivity (n = 72) identify a subgroup of patients with a very favorable survival (Fig. 4).
Relationship between Fusobacterium nucleatum load and mRNA expression of immune-related genes
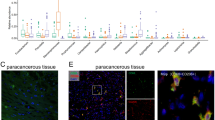
To explore the underlying mechanisms of the favorable prognosis of F. nucleatum-positive tumors, as several studies recently suggested that F. nucleatum modulates the local immunity of various cancers (particularly, macrophages and Tregs)18,19,20, we tested possible association between F. nucleatum load and expression of various immune-related genes in 115 evaluable F. nucleatum-positive samples from the merged cohort (Suppl. Table 6). We observed a significant negative association between F. nucleatum load and markers of B lymphocytes (CD20, p = 0.027), T helper lymphocytes (CD4, p = 0.013), M2 macrophages (CD163, p = 0.020), and fibroblasts (PDGFRß, p = 0.0067). Toll-like receptor (TLR) 4 (p = 0.020) and OX40 ligand (TNFSF4) (p = 0.0067) expressions were significantly decreased in tumors with high F. nucleatum load. Notably, TNFSF9 receptor (TNFRSF9) expression was decreased (p = 0.0067) while the expression of its ligand (TNFSF9) increased with F. nucleatum load, along with the pro-inflammatory cytokine IL-1ß (p = 0.020). Correlations between F. nucleatum load and TNFSF4, TNFSF9, IL1B, and CD163 expression levels are displayed in Supplementary Fig. 3. There was no association with cell proliferation and APOBEC genes.
Validation by LPS and CD163 immunostainings
In order to validate the interplay between bacteria and immune cells, particularly macrophages, we performed immunostainings to detect bacteria using an anti-lipopolysaccharide (LPS) antibody and an anti-CD163 labelling M2 macrophages. We show that, consistent with our PCR results, tumors with high (or low) CD163 and F. nucleatum RNA levels display high (or low) CD163 and LPS expressions, respectively, with an inverse correlation between CD163 and LPS expression levels (Fig. 5 and Suppl. Table 7). LPS staining was located mainly in the cytoplasm of tumor and macrophage cells and more rarely in the form of extracellular bacterial vesicles (Suppl. Fig. 4).
Representative pictures of infiltrating squamous cell carcinomas of the oropharynx: hematoxylin–eosin–safran (HES) stained slides, CD163 and lipopolysaccharide (LPS) immunostainings in a CD163 high/LPS low case (CD163: Histologic Score [HS] 3 (> 50%), LPS: slight expression; F. nucleatum negative) (A) and in a CD163 low/LPS high case (CD163: HS 1 (< 25%), LPS: strong expression; F. nucleatum positive) (B) (scale bar = 200 µM; original magnification: ×50).
Discussion
In this work, we assessed the association between intratumoral F. nucleatum and clinico-pathological features and OS, RFS and MFS in two independent cohorts of patients with OSCC, and explored the interplay between F. nucleatum and well-known immune-related genes. Overall, we showed that F. nucleatum identified a subgroup of OSCC, more frequent in older, non-drinking patients, and associated with less frequent lymph node invasion and distant relapse, and favorable OS (independent predictor), RFS and MFS outcomes in the merged cohort. Other independent prognostic indicators included UICC stage and TP53 mutational status, as previously reported6. Interestingly, the association of low pT or pN stage with F. nucleatum positivity allowed the identification of a patient subgroup with remarkably good prognosis.
The prevalence of F. nucleatum positivity in our study was higher than previously described (82% vs. 16% in a recently published meta-analysis), which may be explained by the high sensitivity of our real-time quantitative PCR assay and the selection of OSCC tumors only (in which F. nucleatum are enriched)28.
The positive correlation between F. nucleatum and survival was unexpected as this bacteria is usually associated with poor prognosis in cancers, particularly in colorectal cancer29,30,31,32. Based on several studies suggesting that F. nucleatum modulates the local immunity of cancers18,19,20, we assessed the association between F. nucleatum loads and expression of various immune-related genes to explore the underlying mechanisms of the favorable prognosis of F. nucleatum positive tumors. We showed that tumors with high F. nucleatum loads displayed low RNA levels of M2 macrophages (CD163), CD4 lymphocytes, fibroblasts (PDGFRß), TLR4, OX40 ligand (TNFSF4), and TNFRSF9, and high levels of TNFSF9 and IL-1ß. These results are consistent with our previously published work in which we identified high OX40 ligand and high PDGFRß as factors associated with poor survival21. Immunohistochemistry analyses confirmed that gram-negative (LPS-positive) bacteria (including F. nucleatum) load inversely correlated with CD163-positive cells.
Data regarding the effects of F. nucleatum on inflammation are conflicting. In most studies, F. nucleatum infection was shown to expand myeloid-derived immune cells and Tregs and promote M2 polarization of macrophages, inducing pro-tumoral inflammation, which inhibit T-cell proliferation and induce T-cell apoptosis33, 34. It also directly inhibits cytotoxic T-cell activities via proteins such as Fap235, 36. TLRs are a family of receptors involved in the detection of microbial agents to induce activation of inflammatory and antimicrobial innate immune responses37. TLR4 is a receptor expressed at the surface of macrophages and tumor cells and has been involved in these pro-inflammatory/immunosuppressive activities of F. nucleatum38, 39. As an apparent paradox, in our study we observed that high F. nucleatum loads were associated with low levels of TLR4 and M2 macrophages. Interestingly, in a mice model of intestinal inflammation, TLR2/TLR4 knock-out induced increased colonization of F. nucleatum and production of pro-inflammatory cytokines including IL-1ß (as observed in our study)18.
On another hand, another work suggested that F. nucleatum enhanced the TNFSF9/IL-1ß signaling inducing M1 macrophage polarization19. This is consistent with the positive correlation that we observed between F. nucleatum loads and expression levels of TNFSF9 and IL-1ß cytokines. The lack of effect on M1 polarization could be explained by the concomitant decrease in TNFSF9 receptor.
It is noteworthy that inflammation and M2 infiltrates were found to be associated with poor prognosis in HNSCC40,41,42.
Taken together, these data suggest that in OSCC F. nucleatum may be associated with “permissive” tumor microenvironment, insensitive to pro-inflammatory signals, with low TLR4 signaling and low recruitment of M2, resulting in favorable clinical outcomes. Of note, defects in TLR functions have been associated with ageing, which may partially account for the higher proportion of older patients in the F. nucleatum positive group43, 44.
This work provides a new insight into the prognostic role of intratumoral F. nucleatum in OSCC patients and opens new avenues regarding the biological interplay between this bacteria and OSCC tumor immune microenvironment. Yet our study has some limitations. First, the small sample size did not allow reaching statistical significance in each individual cohort and merging the two cohorts was necessary to obtain sufficient statistical power for OS. In addition, the immune-related gene analysis was based on selected genes, which are not fully specific of each immune cell subtypes. Overall, these results would require further validation in larger prospective cohorts from randomized clinical trials and using more comprehensive methods such as RNA sequencing. Moreover, saliva samples and normal oral tissue were not available for analysis in our study; it could be of interest to assess the correlation between intratumoral F. nucleatum expression and saliva/normal tissue levels in further studies.
In conclusion, we highlight a unique association between F. nucleatum and OSCC patient survival and tumor immune microenvironment. This can give a rationale for further exploration of the role of F. nucleatum in OSCC carcinogenesis and response to treatment, particularly immune therapy.
Abbreviations
- 95%CI:
-
95% Confidence interval
- F. nucleatum :
-
Fusobacterium nucleatum
- HNSCC:
-
Head and neck squamous cell carcinoma
- HPV:
-
Human papillomavirus
- HR:
-
Hazard ratios
- HRM:
-
High resolution melting
- HS:
-
Histologic score
- LPS:
-
Lipopolysaccharide
- MFS:
-
Metastasis-free survival
- NGS:
-
Next generation sequencing
- OS:
-
Overall survival
- OSCC:
-
Oral squamous cell carcinoma
- PCR:
-
Polymerase chain reaction
- RFS:
-
Relapse-free survival
- Treg:
-
T regulatory cell
- TLR:
-
Toll-like receptor
- UICC:
-
Union for international cancer control
References
Chow, L. Q. M. Head and neck cancer. N. Engl. J. Med. 382, 60–72 (2020).
Bray, F. et al. Global cancer statistics 2018: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Vigneswaran, N. & Williams, M. D. Epidemiologic trends in head and neck cancer and aids in diagnosis. Oral Maxillofac. Surg. Clin. N. Am. 26, 123–141 (2014).
Hashibe, M. et al. Interaction between tobacco and alcohol use and the risk of head and neck cancer: pooled analysis in the international head and neck cancer epidemiology consortium. Cancer Epidemiol. Biomark. Prev. 18, 541–550 (2009).
Psyrri, A., Rampias, T. & Vermorken, J. B. The current and future impact of human papillomavirus on treatment of squamous cell carcinoma of the head and neck. Ann. Oncol. 25, 2101–2115 (2014).
Leemans, C. R., Snijders, P. J. F. & Brakenhoff, R. H. The molecular landscape of head and neck cancer. Nat. Rev. Cancer 18, 269–282 (2018).
De Sanjose, S. et al. Burden of human papillomavirus (HPV)-related cancers attributable to HPVs 6/11/16/18/31/33/45/52 and 58. JNCI Cancer Spectr 2, pky045 (2018).
Turnbaugh, P. J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Zitvogel, L. et al. Cancer and the gut microbiota: an unexpected link. Sci. Transl. Med. 7, 271ps271 (2015).
Routy, B. et al. The gut microbiota influences anticancer immunosurveillance and general health. Nat. Rev. Clin. Oncol. 15, 382–396 (2018).
Zhao, H. et al. Variations in oral microbiota associated with oral cancer. Sci. Rep. 7, 11773 (2017).
Nejman, D. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science 368, 973–980 (2020).
Ganly, I. et al. Periodontal pathogens are a risk factor of oral cavity squamous cell carcinoma, independent of tobacco and alcohol and human papillomavirus. Int. J. Cancer 145, 775–784 (2019).
Yang, C. Y. et al. Oral microbiota community dynamics associated with oral squamous cell carcinoma staging. Front. Microbiol. 9, 862 (2018).
Zhang, Z. et al. Compositional and functional analysis of the microbiome in tissue and saliva of oral squamous cell carcinoma. Front. Microbiol. 10, 1439 (2019).
Rodriguez, R. M. et al. The landscape of bacterial presence in tumor and adjacent normal tissue across 9 major cancer types using TCGA exome sequencing. Comput. Struct. Biotechnol. J. 18, 631–641 (2020).
Chattopadhyay, I., Verma, M. & Panda, M. Role of oral microbiome signatures in diagnosis and prognosis of oral cancer. Technol. Cancer Res. Treat. 18, 1533033819867354 (2019).
Jia, Y. P. et al. TLR2/TLR4 activation induces tregs and suppresses intestinal inflammation caused by Fusobacterium nucleatum in vivo. PLoS ONE 12, e0186179 (2017).
Wu, J. et al. Autoinducer-2 of Fusobacterium nucleatum promotes macrophage M1 polarization via TNFSF9/IL-1beta signaling. Int. Immunopharmacol. 74, 105724 (2019).
Toussi, D. N., Liu, X. & Massari, P. The foma porin from Fusobacterium nucleatum is a toll-like receptor 2 agonist with immune adjuvant activity. Clin. Vaccine Immunol. 19, 1093–1101 (2012).
Lecerf, C. et al. Immune gene expression in head and neck squamous cell carcinoma patients. Eur. J. Cancer 121, 210–223 (2019).
Hoffmann, C. et al. MMP2 as an independent prognostic stratifier in oral cavity cancers. Oncoimmunology 9, 1754094 (2020).
Meseure, D. et al. Biopathological significance of PIWI-piRNA pathway deregulation in invasive breast carcinomas. Cancers (Basel) 12, 2833 (2020).
De Cremoux, P. et al. Inter-laboratory quality control for hormone-dependent gene expression in human breast tumors using real-time reverse transcription-polymerase chain reaction. Endocr. Relat. Cancer 11, 489–495 (2004).
Vacher, S. et al. Genomic instability signature of palindromic non-coding somatic mutations in bladder cancer. Cancers (Basel) 12, 2882 (2020).
Sablin, M. P. et al. Identification of new candidate therapeutic target genes in head and neck squamous cell carcinomas. Oncotarget 7, 47418–47430 (2016).
Bellera, C. A. et al. Protocol of the definition for the assessment of time-to-event endpoints in cancer trials (datecan) project: formal consensus method for the development of guidelines for standardised time-to-event endpoints’ definitions in cancer clinical trials. Eur. J. Cancer 49, 769–781 (2013).
Bronzato, J. D. et al. Detection of fusobacterium in oral and head and neck cancer samples: a systematic review and meta-analysis. Arch. Oral. Biol. 112, 104669 (2020).
Xiao, L. et al. The effect of periodontal bacteria infection on incidence and prognosis of cancer: a systematic review and meta-analysis. Medicine (Baltimore) 99, e19698 (2020).
Gethings-Behncke, C. et al. Fusobacterium nucleatum in the colorectum and its association with cancer risk and survival: a systematic review and meta-analysis. Cancer Epidemiol. Biomark. Prev. 29, 539–548 (2020).
Sun, C. H. et al. The role of Fusobacterium nucleatum in colorectal cancer: from carcinogenesis to clinical management. Chronic. Dis. Transl. med. 5, 178–187 (2019).
Lee, S. A. et al. Global investigations of Fusobacterium nucleatum in human colorectal cancer. Front. Oncol. 9, 566 (2019).
Wu, J., Li, Q. & Fu, X. Fusobacterium nucleatum contributes to the carcinogenesis of colorectal cancer by inducing inflammation and suppressing host immunity. Transl. Oncol. 12, 846–851 (2019).
Nosho, K. et al. Association of Fusobacterium nucleatum with immunity and molecular alterations in colorectal cancer. World J. Gastroenterol. 22, 557–566 (2016).
Gur, C. et al. Binding of the fap2 protein of Fusobacterium nucleatum to human inhibitory receptor tigit protects tumors from immune cell attack. Immunity 42, 344–355 (2015).
Gur, C. et al. Fusobacterium nucleatum supresses anti-tumor immunity by activating ceacam1. Oncoimmunology 8, e1581531 (2019).
Medzhitov, R. Toll-like receptors and innate immunity. Nat. rev. Immunol. 1, 135–145 (2001).
Chen, T. et al. Fusobacterium nucleatum promotes m2 polarization of macrophages in the microenvironment of colorectal tumours via a tlr4-dependent mechanism. Cancer Immunol. Immunother. 67, 1635–1646 (2018).
Yang, Y. et al. Fusobacterium nucleatum increases proliferation of colorectal cancer cells and tumor development in mice by activating toll-like receptor 4 signaling to nuclear factor-kappab, and up-regulating expression of microrna-21. Gastroenterology 152, 851–866 (2017).
Badoual, C. et al. Prognostic value of tumor-infiltrating cd4+ t-cell subpopulations in head and neck cancers. Clin. Cancer Res. 12, 465–472 (2006).
Troiano, G. et al. Prognostic significance of cd68(+) and cd163(+) tumor associated macrophages in head and neck squamous cell carcinoma: a systematic review and meta-analysis. Oral Oncol. 93, 66–75 (2019).
Kumar, A. T. et al. Prognostic significance of tumor-associated macrophage content in head and neck squamous cell carcinoma: a meta-analysis. Front. Oncol. 9, 656 (2019).
Van Duin, D. et al. Age-associated defect in human tlr-1/2 function. J. Immunol. 178, 970–975 (2007).
Panda, A. et al. Age-associated decrease in tlr function in primary human dendritic cells predicts influenza vaccine response. J. Immunol. 184, 2518–2527 (2010).
Author information
Authors and Affiliations
Contributions
Conceived and designed the analysis: C.N., I.B.; Collected the data: D.M., R.L., C.L., C.D., E.J., J.K., O.M., V.C., C.H., M.L., N.B., N.B., E.B., E.P., M.K., C.L.T.; Performed the analysis: M.M., S.V., A.S., D.M., R.L., E.J., J.K.; Interpreted the data: C.N., M.H., D.M., M.K., C.L.T., I.B.; Wrote the paper: C.N., S.V., I.B.; Review and validate the final manuscript: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Neuzillet, C., Marchais, M., Vacher, S. et al. Prognostic value of intratumoral Fusobacterium nucleatum and association with immune-related gene expression in oral squamous cell carcinoma patients. Sci Rep 11, 7870 (2021). https://doi.org/10.1038/s41598-021-86816-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-86816-9
This article is cited by
-
Prognostic impact of oral microbiome on survival of malignancies: a systematic review and meta-analysis
Systematic Reviews (2024)
-
Intratumoural microbiota: a new frontier in cancer development and therapy
Signal Transduction and Targeted Therapy (2024)
-
Intratumoural microbiota: from theory to clinical application
Cell Communication and Signaling (2023)
-
Intratumoral microbiota: roles in cancer initiation, development and therapeutic efficacy
Signal Transduction and Targeted Therapy (2023)
-
Fusobacterium nucleatum and oral cancer: a critical review
BMC Cancer (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.