Abstract
Enhancer DNA methylation and expression of MYBPHL was studied in multiple myeloma (MM). By bisulfite genomic sequencing, among the three CpGs inside the MYBPHL enhancer, CpG1 was significantly hypomethylated in MM cell lines (6.7–50.0%) than normal plasma cells (37.5–75.0%) (P = 0.007), which was negatively correlated with qPCR-measured MYBPHL expression. In RPMI-8226 and WL-2 cells, bearing the highest CpG1 methylation, 5-azadC caused enhancer demethylation and expression of MYBPHL. In primary samples, higher CpG1 methylation was associated with lower MYBPHL expression. By luciferase assay, luciferase activity was enhanced by MYBPHL enhancer compared with empty vector control, but reduced by site-directed mutagenesis of each CpG. RNA-seq data of newly diagnosed MM patients showed that MYBPHL expression was associated with t(11;14). MOLP-8 cells carrying t(11;14) express the highest levels of MYBPHL, and its knockdown reduced cellular proliferation and increased cell death. Herein, as a proof-of-concept, our data demonstrated that the MYBPHL enhancer, particularly CpG1, was hypomethylated and associated with increased MYBPHL expression in MM, which was implicated in myelomagenesis.
Similar content being viewed by others
Introduction
Multiple myeloma (MM) is characterized by the neoplastic proliferation of clonal plasma cells in the bone marrow1,2. Clinically, MM is defined by the presence of ≥ 10% clonal bone marrow plasma cells associated with features of end-organ damages including hypercalcaemia, renal failure, anaemia, or bone lesions that known as CRAB. These criteria have recently been updated to include the presence of any one of the following biomarkers, clonal bone marrow plasma cells of > 60%, a serum free light chain (SFLC) ratio of ≥ 100, or > 1 focal lesion in magnetic resonance imaging (MRI) studies2. Genetically, MM is a heterogeneous and can be broadly classified into non-hyperdiploid (NHRD) and hyperdiploid (HRD)3,4. NHRD is characterized by a primary translocations involving juxtaposition of the strong immunoglobulin heavy (IgH) chain at 14q32 upregulating one of a number of partner oncogenes including CCND1, FGFR3, MMSET, MAF, CCND3 and MAFB3,4. HRD in contrast is characterized by trisomies of odd-numbered chromosomes3,4. Moreover, both NHRD and HRD may carry secondary translocations involving MYC and other genetic abnormalities, including del(17p), del(13), amp(1q), del(1p), and RAS mutations3,4.
The effective regulation of gene control is essential for the specialized function of individual cells within a multicellular organism. DNA methylation is an important component of this process, which is frequently impacted in cancer5,6. A typical feature of methylation in carcinogenesis is global DNA hypomethylation associated with locus-specific hypermethylation at the promoter region of tumour suppressor genes7,8, features which are also typical of MM. Promoter DNA hypermethylation is associated with the recruitment of histone methyltransferases and histone deacetylases, followed by formation of a compact chromatin configuration leading to gene silencing and such processes are active in MM where methylation of promoter-associated CpG island has been shown to result in the inactivation of the tumour suppressors SHP1 and miR-34b/c9,10.
A further element of gene control is provided by a class of cis-acting DNA sequence termed enhancers that are recognized by combinations of histone modifications, transcription factor occupancy, chromatin accessibility, and enhancer RNA expression11,12,13,14. Emerging evidence has demonstrated that structural genomic changes such as translocation, indel, and mutation, in enhancer regions could result in dysregulated gene expressions and hence disease pathogenesis15. Mechanistically, enhancer DNA is brought into close proximity with its target promoter region via DNA looping mediated by binding with transcription factors, mediator proteins, and activator proteins, resulting in formation of pre-initiation complex for gene transcription16. Major markers include the presence of H3K27ac and H3K4me1 but absence of H3K4me317,18. Recently, enhancer DNA methylation has been implicated in the regulation of gene expression18,19. For example, by integrated analysis of enhancer regions, Illumina 450 K methylation array, and RNA-seq data, methylation of multiple enhancers, including the MYBPHL (myosin binding protein H like) enhancer, were shown to be inversely correlated with the expressions of their target genes in hepatocellular carcinoma (HCC)18.
MYBPHL encodes a protein with two immunoglobulin superfamily domains and a fibronectin 3 domain, the normal expression of which is restricted to the heart, however, there have been report of decreased methylation and increased expression in pituitary adenoma20. It is located at chromosome 1p13 a region frequently affected by structural variation and copy number change in MM21. Using gene expression analysis we noted that it is expressed in a subgroup of samples where it is associated with t(11;14). These observations suggest that it may be expressed as part of the malignant process as a consequence of methylation change. We have previously elucidated the role of methylation based control of the expression of this gene in hepatocellular cancer and we hypothesized it may also be important for malignant transformation of plasma cells. Herein, enhancer DNA methylation and expression of MYBPHL was studied in CD138-sorted normal plasma cells, human MM cell lines (HMCLs), and CD138-sorted primary samples of MM. Moreover, to the best of our knowledge, the data presented here is the first to demonstrate a potential oncogenic function of MYBPHL in MM.
Results
Enhancer of MYBPHL is methylated in normal plasma cells but hypomethylated in HMCLs
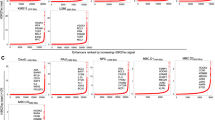
The enhancer DNA of MYBPHL has a genomic size of 348 bp and is embedded in the first intron of MYBPHL, which is localized at 1p13, and has been shown to be associated with three CpG dinucleotides, Fig. 1a. By bisulfite genomic sequencing (BGS), DNA methylation of the MYBPHL enhancer was studied in bisulfite-converted DNA of CD138-sorted normal plasma cells from healthy bone marrow donors (n = 7), human MM cell lines (HMCLs; n = 7), and an enzymatically methylated positive control DNA. The result from methylated positive control DNA showed conversion of all unmethylated non-CpG cytosine residues into uracils (turned into thymidines after PCR), whereas methylated cytosine residues in CpG dinucleotides remained unchanged, indicating complete bisulfite conversion of DNA samples and specificity of BGS, Fig. 1a. Of the three CpG dinucleotides in the MYBPHL enhancer, the mean methylation levels were 64.9% and 56.4% in CD138-sorted normal plasma cells and HMCLs, respectively, Table 1. Particularly, methylation of the first CpG dinucleotide (CpG1) was significantly higher in CD138-sorted normal plasma cells than HMCLs (58.8% vs. 30.4%; P = 0.007). These data suggest that the MYBPHL enhancer was hypomethylated, especially at CpG1, in HMCLs than compared to CD138-sorted normal plasma cells.
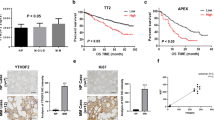
MYBPHL enhancer methylation and MYBPHL expression. (a) Schematic diagram of MYBPHL enhancer. MYBPHL enhancer (chr1:109841804–109842152), localized to 1p13, was retrieved from FANTOM5 Human Enhancer Tracks (http://slidebase.binf.ku.dk/human_enhancers/) and viewed in the UCSC Genome Browser39. Transcription start site (TSS) and the direction of transcription of MYBPHL were indicated by the red elbow arrow. MYBPHL enhancer region was shown overlapping with histone modifications annotating functional enhancer, i.e. presence of H3K27ac and H3K4me1 but absence of H3K4me3. Data from GeneHancer predicted interaction (downward curve) between MYBPHL enhancer to its target MYBPHL promoter40. Distribution of the three CpG dinucleotides on MYBPHL enhancer was depicted using BISMA 41. Bisulfite genomic sequencing (BGS) of the methylated positive control DNA showed conversion of all unmethylated cytosine residues into uracils (turned into thymidines after PCR), whereas methylated cytosine residues in CpG dinucleotides remained unchanged, indicating complete bisulfite conversion of DNA samples and specificity of BGS. (b) In HMCLs, methylation of all three CpG dinucleotides on average or in individual was plotted against MYBPHL expression as detected by qPCR. Data demonstrated that higher MYBPHL enhancer methylation, particularly CpG1, correlated with lower MYBPHL expression. (c) In RPMI-8226 and WL-2 cells, which had high level of MYBPHL enhancer methylation, treatment with 5-azadC led to MYBPHL enhancer demethylation, as evidenced by BGS, with concomitant re-expression of MYBPHL transcript, as detected by qPCR. (d) In primary myeloma samples, higher methylation of MYBPHL enhancer, particularly CpG1, correlated with lower MYBPHL expression. Data of qPCR were mean delta Ct values from three independent experiments with triplicate in each.
Enhancer DNA methylation negatively correlated with expression of MYBPHL.
To investigate the relationship of enhancer DNA methylation and expression of MYBPHL, MYBPHL expression was measured in HMCLs (n = 7). A higher level of MYBPHL expression was demonstrated by Taqman-based qPCR, in HMCLs that were associated with lower MYBPHL enhancer methylation, Fig. 1b. However, among the three CpG dinucleotides, higher methylation of CpG1, but not CpG2 or CpG3, was demonstrated in HMCLs with lower MYBPHL expression. This result implicates this region as being the most potent region in controlling expression of this gene.
To validate the inverse relationship between enhancer DNA methylation and expression of MYBPHL, RPMI-8226 and WL-2 cells, which had highest CpG1 methylation, were treated with a hypomethylation agent, 5-azadC. Upon treatment with 5-azadC, both RPMI-8226 and WL-2 cells showed demethylation of the MYBPHL enhancer, as demonstrated by BGS, and this finding was associated with re-expression of MYBPHL transcripts by qPCR, Fig. 1c. In primary MM samples at relapse (n = 12), MYBPHL expression in CD138-sorted MM plasma cells was examined by qPCR. The result of this showed that MYBPHL enhancer methylation in six samples with paired CD138-sorted DNA available by BGS correlated with lower levels of expression of MYBPHL. Similarly, in primary samples, higher methylation of MYBPHL enhancer, especially CpG1 rather than CpG2 or CpG3, was correlated with lower MYBPHL expression, Fig. 1d.
To confirm the role of enhancer CpG dinucleotides on MYBPHL expression, the full-length MYBPHL enhancer was cloned and a comparison of constructs with site-directed mutagenesis at each of the three CpG dinucleotides by luciferase reporter assays was carried out. The results showed that luciferase activity was increased upon insertion of the MYBPHL enhancer in comparison with an empty vector control, demonstrating the transcriptional activity of the MYBPHL enhancer, Fig. 2. Moreover, by mutation of individual CpG into ApG, luciferase activity was significantly decreased as compared with wild-type MYBPHL enhancer, thereby functionally important for the MYBPHL enhancer on gene expression, Fig. 2.
Enhancer-associated CpG dinucleotides and enhancer activity. By luciferase reporter assay, luciferase activity was increased by the presence of wild-type MYBPHL enhancer DNA, as compared with empty vector control. By site-directed mutagenesis of individual CpG into ApG, luciferase activity was significantly reduced, as compared with that of wild-type (***P ≤ 0.001). Data were mean ± s.d. of firefly/renilla from three independent experiments with triplicate in each.
MYBPHL is expressed in newly diagnosed MM patients.
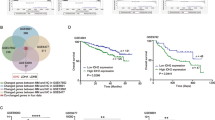
Using the RNA-seq data from the CoMMpass study, we examined the expression of MYBPHL in newly diagnosed MM, and showed that it is either not expressed in 46% of patients (342/734) or expressed at low levels. However, in 9 of 734 cases, its level was expressed at greater than 5, Fig. 3a. Using, sequencing defined subgroups, we analyzed the overall expression of MYBPHL and showed that it varied among cytogenetic subgroups with higher expression levels seen in the CCND1 and MAFA groups [t(11;14) and t(8;14) respectively], Fig. 3b–d.
Expression and function of MYBPHL in MM cells. (a) Expression (RPKM) of MYBPHL by RNA-seq in 734 newly diagnosed MM patients. (b) Pattern of MYBPHL expression (RPKM) in different sequencing defined translocation subgroups. (c, d) Expression (RPKM) of MYBPHL is higher among the t(11;14) and t(8;14) patients. (e) Effect of MYBPHL knockdown in MM cells. In t(11;14)-bearing MOLP-8 cells, which had highest MYBPHL expression, MYBPHL-specific siRNAs were transfected, incubated for 48 h, followed by subsequent assays. By qPCR, MYBPHL expression was measured and compared with those transfected with scrambled negative control. (f) By MTS assay, cellular proliferation in response to MYBPHL knockdown was determined as compared with scrambled negative control. (g) By trypan blue staining, number of dead cells subjected to MYBPHL knockdown was examined as compared with scrambled negative control. Data were mean ± s.d. of three independent experiments with triplicate in each. RPKM reads per kilobase per million mapped reads.
MYBPHL is associated with oncogenic function in MM cells
The function of MYBPHL in MM cells was studied by siRNA-mediated knockdown. In MOLP-8 cells [carrying the t(11;14)(q13;q32)], in which MYBPHL expression was the highest among the HMCLs used, transfection with MYBPHL-specific siRNAs led to downregulation of MYBPHL expression by qPCR, Fig. 3e, or by Western blot, Supplementary Fig. 1, as compared cells transfected with scrambled negative control. By MTS assay, knockdown of MYBPHL led to inhibition of cellular proliferation by ~ 30%, Fig. 3f, as compared with scrambled negative control. Conversely, by trypan blue staining, knockdown of MYBPHL increased cell death by ~ 30%, Fig. 3g, as compared with scrambled negative control. Therefore, these data suggested a pro-proliferative role of MYBPHL in MM cells.
Discussion
We explored a potential role for MYBPHL as an oncogene by siRNA knockdown in plasma cell lines overexpressing it. We demonstrated that knockdown of MYBPHL leads to decreased cellular proliferation and increased cell death, suggesting a potential novel role for MYBPHL in myelomagenesis in addition to its known functions in heart diseases22,23. We showed that MYBPHL is hypomethylated in MM cells in comparison with CD138-sorted normal plasma cells suggesting that it may function as an oncogene that is regulated by demethylation. Global DNA hypomethylation has been shown to be a hallmark of MM and increases as the disease progresses24,25. Such widespread DNA hypomethylation is linked to chromatin instability and transcriptional activation of oncogenes presumably via demethylation of promoter-associated CpG islands26. The MYBPHL enhancer is embedded in its first intron, about 7.5 kb downstream of the transcription start site. In the context of a potential oncogenic role for this gene, a recent genome-wide methylation study in MM has demonstrated that DNA hypermethylation is more likely to be associated with intronic enhancers than gene promoters and correlated with reduced enhancer activity and downregulation of their associated genes, e.g. SLC15A4, PVT1, and NCOR227. Our data show that the expression of MYBPHL was repressed by hypermethylation at its intronic enhancer and hypomethylation was associated with upregulation.
Exploring the methylation based control of gene expression at this locus further we showed that different CpG dinucleotides at the same intronic enhancer had a differential influence in with the regulation of expression of the associated gene. Of the three CpG dinucleotides, CpG1 was mainly associated with the expression of MYBPHL in both HMCLs and primary samples of MM, therefore, these data not only rekindled the role of single-site CpG methylation in the regulation of gene expression, but also demonstrated that regulation by single-site CpG methylation was not restricted to the promoter region28,29. On the other hand, as the luciferase assay did not correspondingly indicate that C- > A mutation at CpG1 affected the enhancer activity more than CpG2 or CpG3, thereby suggesting that methylation of CpG1 might also affect something other than the enhancer activity, such as interaction with the target promoter. Even though enhancer landscapes are considered to be cell-specific13,30, we clearly showed that the interaction of the MYBPHL enhancer-promoter pair in myeloma plasma cells is consistent with the previous report in HCC18.
In MM, as emerging evidences have shown that, in contrast to normal cells, cancer cells were more sensitive to enhancer-selective inhibitors, such as inhibitors targeting BET-bromodomain protein 4 (BRD4) and cyclin-dependent kinase 7 (CDK7)31,32,33, these data further support to the concept of incorporating enhancer-targeting inhibitors in the treatment of MM. Moreover, by the advent of genome-wide technologies interrogating CpG dinucleotides, in particular those residing outside of promoter regions, which had not been covered before, our data indicate that further studies of DNA methylation at “nascent” CpG dinucleotides may give new insights into the regulation of gene expression in cancer and may provide either potential therapeutic interventions or the identification of methylation based biomarkers with clinical significance34.
Of note, these data could be strengthened by correlating MYBPHL methylation and expression in normal plasma cells. However, CD138-sorted normal plasma cells had been exhausted for DNA extraction, whereby a higher level of MYBPHL methylation was demonstrated in normal plasma cells than HMCLs (Table 1). In fact, in contrast to solid cancers, in which normal controls are readily available adjacent to tumour cells, normal plasma cells as a control for MM plasma cells are much more difficult to obtain. As normal plasma cells only constitute < 1% of mononuclear cells in the bone marrow, collection of sufficient CD138-sorted normal plasma cells for both DNA and RNA extraction is extremely difficult. Moreover, in the past, normal bone marrow plasma cells were mainly derived from bone marrow stem cells collected by bone marrow puncture of healthy donors under general anesthesia, whereby normal plasma cells could be isolated. In the last two decades, haematopoietic stem cells were instead collected by apheresis after mobilization of stem cells into the peripheral blood, hence precluding isolation of normal marrow plasma cells. In the future, similar to a recent publication in MM35, removed hip in patients receiving total hip replacement could serve as an alternative source of normal plasma cells for both DNA and RNA extraction. Lastly, the current study serves as a proof-of-principle pilot study, conducive to future enhancer methylation studies in MM that will include more primary samples.
Methods
Patient samples
Twelve patients with relapsed/refractory multiple myeloma (RRMM), including “relapsed” or “relapsed-and-refractory myeloma” cases were included for study of MYBPHL expression. Definition of relapsed myeloma is consistent with previously described36. This study was approved by the Institutional Review Board of Queen Mary Hospital. Written informed consent was obtained from all participants involved in the study. Normal bone marrow plasma cells were obtained from healthy marrow donors for bone marrow transplantation. All study methods were performed in accordance with relevant guidelines and regulations.
Cell culture
Human MM cell lines (HMCLs) KMS-12-PE, MOLP-8, and U-266 were purchased from Deutsche Sammlung von Mikroorganismen und Zellkulturen (Braunschweig, Germany). NCI-H929 was purchased from American Type Culture Collection (Manassas, VA, USA). LP-1 and RPMI-8226 were kindly provided by Prof. Robert Orlowski (Department of Lymphoma/Myeloma, Division of Cancer Medicine, The University of Texas MD Anderson Cancer Center, Houston, TX, USA). WL-2 was kindly provided by Prof. Andrew Zannettino (Myeloma Research Programme, The University of Adelaide, Australia). Cell lines were cultured in RPMI-1640 medium (IMDM for LP-1), supplemented with 10% fetal bovine serum, 50 U/mL of penicillin and 50 μg/mL streptomycin, in a humidified atmosphere of 5% CO2 at 37 °C. All cell culture reagents were purchased from Thermo Fisher Scientific (Waltham, MA, USA).
Bisulfite genomic sequencing
Bisulfite-treated DNA was used as template. Enhancer DNA of MYBPHL was amplified and cloned using TOPO TA Cloning Kit (Thermo Fisher Scientific), followed by sequencing, according to the manufacturer's instructions. Primers used were forward 5′-GGG GGT TTG GTT GAG ATA GAT AT-3′ and reverse 5′-ACC CAA ATA AAA ACA TAA ACC ACA C-3′ 18. PCR conditions for MgCl2/Tm/cycles were 2.0 mM/56 °C/35X. Methylated Human Control (Promega) was used as positive control for DNA methylation.
Hypomethylation treatment
Cells were cultured and treated with a hypomethylating agent, 5-aza-2′-deoxycytidine (5-azadC; Sigma-Aldrich, St. Louis, MO, USA) as previously described10.
Knockdown of MYBPHL
MYBPHL-specific siRNAs (Assay ID: s50998, s50999, s51000) and Cy3-labeled scrambled negative control siRNA (Cat. no.: AM4621) were purchased from Thermo Fisher Scientific (Waltham, MA, USA). Briefly, MOLP-8 cells were seeded at a density of 0.5 × 106 cells per well in a 24-well plate and transfected with MYBPHL-specific siRNAs (25 nM) or scrambled negative control siRNA using Lipofectamine RNAiMAX (Invitrogen, Carlsbad, CA, USA). Cells were harvested at 48 h post-transfection, followed by functional assays.
MTS assay
Cellular proliferation was measured by MTS assay (Promega; Madison, WI, USA). In brief, transfected cells of 5 × 104 were resuspended in 100 µL medium and seeded in a 96-well microplate in triplicate. At 48 h post-transfection, MTS reagent of 20 µL was added into each well, followed by incubation for a further 4 h. Absorbance at 490 nm was measured using Epoch Microplate Spectrophotometer (BioTek; Winooski, VT, USA), background subtracted, and normalized to cells transfected with scrambled negative control siRNA. Means of three independent experiments with triplicate in each were plotted.
Trypan blue staining
Cell death was examined by staining with trypan blue (Thermo Fisher Scientific). Briefly, at 48 h post-transfection, number of unstained live cells and stained dead cells were counted in five random microscopic fields under a microscope, followed by normalization to cells transfected with scrambled negative control siRNA. Means of three independent experiments were plotted.
Luciferase reporter assay
Firefly luciferase reporter plasmid constructs (pGL3-Promoter Vector, Promega, Madison, WI, USA) cloned with enhancer DNA of MYBPHL (348 bp) were kind gifts from Dr. Alfred Sze Lok CHENG (School of Biomedical Sciences, CUHK)18. On the enhancer DNA of MYBPHL, each cytosine residue of the three CpG dinucleotides, located at nucleotide position 16, 154, and 292, was individually mutated into adenine residues using QuikChange Lightning Multi Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA) as previously described28. Briefly, SW480 cells (kind gift from Dr. Roberta PANG, Department of Surgery, HKU) were seeded at a density of 2 × 105 cells per well in a 24-well plate overnight, followed by co-transfection with firefly and renilla (pGL4.75[hRluc/CMV] Vector, Promega, Madison, WI, USA) luciferase reporter plasmids at a ratio of 500:0.75 using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Cells were harvested at 24 h post-transfection. Firefly and renilla luciferase activity were sequentially generated using Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA), and their luminescent signals were measured using CLARIOstar microplate reader (BMG LABTECH, Cary, NC, USA). Luciferase activity of firefly was normalized by renilla and relative luciferase activity was obtained as compared with empty pGL3-Promoter Vector. Data represents mean of three independent transfections with triplicate in each.
RNA-seq analysis
We analyzed RNA-seq from newly diagnosed MM (NDMM) samples from the CoMMpass dataset (n = 734). RNA-seq data that were uniformly processed. Salmon21 (v0.7.2) was used to align reads to the transcriptome and quantify expression at the gene and transcript level as previously published37. Expression levels were compared across cytogenetic groups amongst patients that had WES data available (n = 628)38.
Statistical analysis
Methylation and expression of MYBPHL were plotted and analyzed using GraphPad Prism 8 (GraphPad Software; La Jolla, CA, USA). MYBPHL methylation between CD138-sorted normal plasma cells and HMCLs, data between cells transfected with MYBPHL-specific siRNAs and scrambled negative control siRNA were compared by Student’s t-test. All P values were two-sided, and P < 0.05 was defined as a significant difference.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Kumar, S. K. et al. Multiple myeloma. Nat. Rev. Dis. Primers 3, 17046. https://doi.org/10.1038/nrdp.2017.46 (2017).
Rajkumar, S. V. et al. International myeloma working group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol. 15, e538–e548 (2014).
Corre, J., Munshi, N. & Avet-Loiseau, H. Genetics of multiple myeloma: another heterogeneity level?. Blood 125, 1870–1876 (2015).
Morgan, G. J., Walker, B. A. & Davies, F. E. The genetic architecture of multiple myeloma. Nat. Rev. Cancer 12, 335–348 (2012).
Jones, P. A. & Baylin, S. B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 3, 415–428 (2002).
Dor, Y. & Cedar, H. Principles of DNA methylation and their implications for biology and medicine. TLancet 392, 777–786 (2018).
Plass, C. Cancer epigenomics. Hum. Mol. Genet. 11, 2479–2488 (2002).
Llinas-Arias, P. & Esteller, M. Epigenetic inactivation of tumour suppressor coding and non-coding genes in human cancer: An update. Open Biol. 7, 170152 (2017).
Chim, C. S., Fung, T. K., Cheung, W. C., Liang, R. & Kwong, Y. L. SOCS1 and SHP1 hypermethylation in multiple myeloma: Implications for epigenetic activation of the Jak/STAT pathway. Blood 103, 4630–4635. https://doi.org/10.1182/blood-2003-06-2007 (2004).
Wong, K. Y. et al. Epigenetic inactivation of the MIR34B/C in multiple myeloma. Blood 118, 5901–5904. https://doi.org/10.1182/blood-2011-06-361022 (2011).
Cao, Q. & Yip, K. Y. A survey on computational methods for enhancer and enhancer target predictions. Comput. Biol. Bioinform. 1, 3–27 (2016).
Pennacchio, L. A., Bickmore, W., Dean, A., Nobrega, M. A. & Bejerano, G. Enhancers: Five essential questions. Nat. Rev. Genet. 14, 288–295 (2013).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: From properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Vernimmen, D. & Bickmore, W. A. The hierarchy of transcriptional activation: From enhancer to promoter. Trends Genet. 31, 696–708 (2015).
Bhagwat, A. S., Lu, B. & Vakoc, C. R. Enhancer dysfunction in leukemia. Blood 131, 1795–1804 (2018).
Sur, I. & Taipale, J. The role of enhancers in cancer. Nat. Rev. Cancer 16, 483 (2016).
Kimura, H. Histone modifications for human epigenome analysis. J. Hum. Genet. 58, 439–445 (2013).
Cao, Q. et al. Reconstruction of enhancer-target networks in 935 samples of human primary cells, tissues and cell lines. Nat. Genet. 49, 1428. https://doi.org/10.1038/ng.3950 (2017).
Ordoñez, R., Martínez-Calle, N., Agirre, X. & Prosper, F. DNA methylation of enhancer elements in myeloid neoplasms: Think outside the promoters?. Cancers 11, 1424 (2019).
Cheng, S. et al. Identification of key genes in invasive clinically non-functioning pituitary adenoma by integrating analysis of DNA methylation and mRNA expression profiles. J. Transl. Med. 17, 407 (2019).
Li, F. et al. Identification of characteristic and prognostic values of chromosome 1p abnormality by multi-gene fluorescence in situ hybridization in multiple myeloma. Leukemia 30, 1197–1201 (2016).
Barefield, D. Y. et al. Experimental modeling supports a role for MyBP-HL as a novel myofilament component in arrhythmia and dilated cardiomyopathy. Circulation 136, 1477–1491 (2017).
Lahm, H. et al. Myosin binding protein H-like (MYBPHL): A promising biomarker to predict atrial damage. Sci. Rep. 9, 1–8 (2019).
Walker, B. A. et al. Aberrant global methylation patterns affect the molecular pathogenesis and prognosis of multiple myeloma. Blood 117, 553–562. https://doi.org/10.1182/blood-2010-04-279539 (2011).
Chim, C. S., Liang, R. & Kwong, Y. L. Hypermethylation of gene promoters in hematological neoplasia. Hematol. Oncol. 20, 167–176 (2002).
Costello, J. F. & Plass, C. Methylation matters. J. Med. Genet. 38, 285–303. https://doi.org/10.1136/jmg.38.5.285 (2001).
Agirre, X. et al. Whole-epigenome analysis in multiple myeloma reveals DNA hypermethylation of B cell-specific enhancers. Genome Res. 25, 478–487 (2015).
Zou, B. et al. Correlation between the single-site CpG methylation and expression silencing of the XAF1 gene in human gastric and colon cancers. Gastroenterology 131, 1835–1843. https://doi.org/10.1053/j.gastro.2006.09.050 (2006).
Pogribny, I. P., Pogribna, M., Christman, J. K. & James, S. J. Single-site methylation within the p53 promoter region reduces gene expression in a reporter gene construct: Possible in vivo relevance during tumorigenesis. Can. Res. 60, 588–594 (2000).
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Fulciniti, M. et al. Non-overlapping control of transcriptome by promoter-and super-enhancer-associated dependencies in multiple myeloma. Cell Rep. 25, 3693–3705 (2018).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Chipumuro, E. et al. CDK7 inhibition suppresses super-enhancer-linked oncogenic transcription in MYCN-driven cancer. Cell 159, 1126–1139 (2014).
Koch, A. et al. Analysis of DNA methylation in cancer: Location revisited. Nat. Rev. Clin. Oncol. 15, 459–466 (2018).
Ledergor, G. et al. Single cell dissection of plasma cell heterogeneity in symptomatic and asymptomatic myeloma. Nat. Med. 24, 1867–1876 (2018).
Wong, K. Y. et al. Frequent functional activation of RAS signalling not explained by RAS/RAF mutations in relapsed/refractory multiple myeloma. Sci. Rep. 8, 13522 (2018).
Bauer, M. A. et al. Differential RNA splicing as a potentially important driver mechanism in multiple myeloma. Haematologica (2020).
Walker, B. A. et al. Identification of novel mutational drivers reveals oncogene dependencies in multiple myeloma. Blood 132, 587–597 (2018).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Fishilevich, S. et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database 2017 (2017).
Rohde, C., Zhang, Y., Reinhardt, R. & Jeltsch, A. BISMA-Fast and accurate bisulfite sequencing data analysis of individual clones from unique and repetitive sequences. BMC Bioinform. 11, 230 (2010).
Acknowledgements
The authors would like to thank MMRF CoMMpass Network for their contributions to the CoMMpass patient and data collection and the Multiple Myeloma Research Foundation.
Author information
Authors and Affiliations
Contributions
C.S.C. conceived and designed the study; K.Y.W. designed and conducted the experiments; E.M.B. performed analysis of data from the CoMMpass study; A.S.L.C. contributed luciferase reporter plasmids; all authors drafted, critically revised, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wong, K.Y., Morgan, G.J., Boyle, E.M. et al. A proof-of-concept study for the pathogenetic role of enhancer hypomethylation of MYBPHL in multiple myeloma. Sci Rep 11, 7009 (2021). https://doi.org/10.1038/s41598-021-86473-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-86473-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.