Abstract
Odontotermes formosanus (Shiraki) is an important pest in the world. Serratia marcescens have a high lethal effect on O. formosanus, but the specific insecticidal mechanisms of S. marcescens on O. formosanus are unclear, and the immune responses of O. formosanus to S. marcescens have not been clarified. At present, genetic database resources of O. formosanus are extremely scarce. Therefore, using O. formosanus workers infected by S. marcescens and the control as experimental materials, a full-length transcriptome was sequenced using the PacBio Sequel sequencing platform. A total of 10,364 isoforms were obtained as the final transcriptome. The unigenes were further annotated with the Nr, Swiss-Prot, EuKaryotic Orthologous Groups (KOG), Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Ortholog public databases. In a comparison between the control group and a Serratia marcescens-infected group, a total of 259 differentially expressed genes (DEGs) were identified, including 132 upregulated and 127 downregulated genes. Pathway enrichment analysis indicated that the expression of the mitogen-activated protein kinase (MAPK) pathway, oxidative stress genes and the AMP-activated protein kinase (AMPK) pathway in O. formosanus may be associated with S. marcescens treatment. This research intensively studied O. formosanus at the high-throughput full-length transcriptome level, laying a foundation for further development of molecular markers and mining of target genes in this species and thereby promoting the biological control of O. formosanus. Furthermore, these results will be helpful to clarify the action mechanisms of S. marcescens on O. formosanus, and also explore the relationship between O. formosanus and S. marcescens. In addition, this study will identify the immune response of O. formosanus to S. marcescens, which will provide a theoretical foundation for the development of new immunosuppressants for O. formosanus.
Similar content being viewed by others
Introduction
Termites are known for their destructive potential, and they harm buildings, agricultural and forestry crops, the earth dams of reservoirs, cable bridges, traffic facilities, and more, and they often eat noncellulose materials1,2,3. Termites are widely distributed and cause billions of property losses every year. Because of the complex feeding habits of termites, they can cause more or less severe harm to human buildings, vehicles, crops, and so on. More serious effects can include the collapse of houses and result in threats to human life, causing irreparable damage. In 1992, the number of known termite species in China had reached 476, and the termite damage to the south of the Yangtze River was the most serious. At that time, China’s annual damage due to termites had reached more than 2 billion yuan. The economic loss caused by termite damage in the United States in 1998 was approximately $5 billion4.
Odontotermes formosanus (Shiraki) is a derived type of termite that remains underground most of the time. The workers of this species mainly damage bark and roots, and these termites prefer humid but not wet conditions5. Donkeys, mountain frogs, bats, various ants, spiders, pangolins, and other animals are their natural enemies6. O. formosanus is the main pest in reservoir and river dams, often causing dam leakage, collapse and breach7. Li et al. found that the presence of O. formosanus nest has a great impact on the overall and local stability of the embankment. The existence of termite nest reduces the maximum safety factor of the downstream slope by 17%8. Because the nests of O. formosanus are very well concealed, the damage caused by this species is not easily detected. When it is detected, a great deal of damage is already present. In addition, O. formosanus is a generalist and can damage more than 100 kinds of plants, including eucalyptus, cedar, magnolia, and maple9. Kirton et al.10 found the phenomenon of ring-barking and root debarking of saplings in the Malaysian peninsula of Southeast Asia. At the same time, many termite species of termite genera, including Schedorhinotermes, Odontotermes, Procapritermes, Pericapritermes and Nasutitermes, were found on or near the main roots of the saplings, among which the most common was Odontotermes.
Currently, increasing attention is being paid to the prevention and control of termites in China. Large amounts of manpower, material resources and financial resources have been invested in termite prevention and control to minimize termite-related losses to human beings. At present, imidacloprid, fipronil, chlorpyrifos, fenvalerate and bifenthrin are commonly used termite control agents. Although chemical control can achieve certain results, there will be pesticide residues, which will cause serious harm to the ecological environment. At the same time, due to the increase in human environmental awareness, green pesticides are becoming increasingly popular. The research and development of new termite pesticides is currently a hot topic. Notably, in the field of biological control, Serratia marcescens can effectively control some pests and pathogens11,12. However, Serratia marcescens strains from different sources have very different characteristics. Previously, we isolated a strain of S. marcescens from O. formosanus. After separation and purification, a pure strain was obtained13. Then, the bioassay showed that S. marcescens have a high lethal effect on O. formosanus14. At the same time, we found that the insecticidal effect of Metarhizium anisopliae and S. marcescens was better than M. anisopliae or S. marcescens individually14. In addition, we reported that the main components that affected O. formosanus were in a supernatant and that the insecticidal protease in the supernatant resulted in the death of O. formosanus15.These results showed that S. marcescens had good insecticidal effect on O. formosanus, but the action mechanisms of S. marcescens on O. formosanus need to be further clarified. Guo et al. demonstrated that MAPK signaling pathway is a general switch, which can transregulate the differential expression of genes in the midgut of Plutella xylostella, so as to resist the toxic effect of Bacillus thuringiensis (Bt)16. So, we speculate that the signal transduction and energy metabolism of O. formosanus will be affected in varying degrees, and the oxidative stress will be affected when infected by S. marcescens.
In recent years, third-generation high-throughput sequencing technology characterized by single-molecule real-time (SMRT) sequencing has been successfully applied to functional genome research in yeast17, rabbit18, sorghum19, corn20, wheat21 and cotton22. Compared with second-generation sequencing technology, third-generation sequencing technology not only has the advantages of large data sets, long sequence reads and full-length gene transcripts but also does not require sequence splicing and assembly, which greatly improves the accuracy of gene functional annotation23. However, this technology has not previously been used in termites. Therefore, this study used O. formosanus as an experimental material to perform full-length transcriptome sequencing to explore the key metabolic pathways, signaling pathways and genes affected by S. marcescens, and analysis of O. formosanus infected with S. marcescens using the third-generation high-throughput sequencing technology to clarify the action mechanisms of S. marcescens on O. formosanus. Furthermore, these data also explored the relationship between O. formosanus and S. marcescens, which will provide theoretical basis for biological control of O. formosanus by using S. marcescens. In addition, this study will identify the immune response of O. formosanus to S. marcescens, which will provide a theoretical foundation for the development of new immunosuppressants for O. formosanus.
Materials and methods
Insects and pathogenic bacteria
Odontotermes formosanus was collected from JuRong (Jiangsu Province, China), and the nests were harvested. Then, the nests were placed in an insect incubator under dark conditions (L:D = 0:24) and 75 ± 1% relative humidity at 27 ± 1 °C. The S. marcescens strain SM1 was previously isolated from infected and dead O. formosanus and stored in glycerin at − 80 °C.
Sample processing
The isolated Serratia marcescens was transferred to a bacterial basal medium and cultured in the dark at 27 °C for 24 h. After streaking, a single colony was removed, placed in a 250 mL Erlenmeyer flask containing 50 mL of seed culture medium, and cultured for 12 h at 30 °C in a constant temperature shaker at 200 r/min . Then, 70 mL of the seed solution was added to 250 mL of the fermentation medium and cultured in a shaking incubator at a temperature of 30 °C for 36 h at a rate of 200 r/min. The cultured fermentation medium was used in subsequent experiments. Then, filter paper was placed on the bottom of a 9 cm petri dish and wetted with water. Twenty mature worker termites with good vitality were placed into the petri dish, and ten petri dishes were set up. In the S. marcescens-infected experimental group (SMT), 0.12 μL of S. marcescens solution (culture fermentation medium) was collected and dropped on pronotum of termites. For the control group (CK), 0.12 μL sterile liquid (fermentation medium without S. marcescens) was placed in the same locations on the termites. After 20 h of treatment, the ten termites were collected and stored in liquid nitrogen for subsequent experiments in the CK and the SMT respectively. The CK and the SMT were repeated three times independently.
RNA sample preparation
After the tissue was taken out, it was frozen in liquid nitrogen, then stored in − 80 °C. Total RNA was extracted by Trizol method. Agarose gel electrophoresis was used to analyze the degree of degradation and whether there was contamination in RNA solution. Then, RNA purity was checked using a NanoPhotometer spectrophotometer (IMPLEN, CA, USA). Next, RNA concentration was measured using a Qubit RNA Assay Kit in a Qubit 3.0 Fluorometer (Life Technologies, CA, USA). RNA integrity was assessed using the RNA Nano 6000 Assay Kit of the Agilent Bioanalyzer 2100 system24,25.
Library preparation and SMRT sequencing
A total of 1 μg RNA per sample was used as input material for RNA sample preparation. Sequencing libraries were generated using the NEBNext Ultra RNA Library Prep Kit for Illumina (NEB, USA) following the manufacturer’s recommendations, and index codes were added to attribute the sequences to each sample24,25,26. In short, mRNA was purified from the total RNA using poly-T oligo-attached magnetic beads. Fragmentation was carried out using divalent cations under elevated temperature in NEBNext First Strand Synthesis Reaction Buffer (5×)26. The first strand of cDNA was synthesized by random hexamer primer and M-MuLV Reverse Transcriptase (RNaseH-). The second strand of cDNA was synthesized by DNA polymerase I and RNase H. The rest overhangs were transformed into blunt ends via exonuclease/polymerase activities. After adenylation of the 3′ ends of the DNA fragments, NEBNext Adaptor with hairpin loop structure was connected to prepare for hybridization. In order to select the proper length of cDNA fragment, the library fragment was purified by AMPure XP system (Beckman Coulter, Beverly, USA). Then, 3 μL USER Enzyme (NEB, USA) was incubated with the size-selected, adaptor-ligated cDNA at 37 °C for 15 min followed by 5 min at 95 °C. PCR was then performed with Phusion High-Fidelity DNA polymerase, Universal PCR primers and Index (X) Primer. Finally, the products were purified (AMPure XP system), and library quality was assessed on the Agilent Bioanalyzer 2100 System25,26. The library preparations were sequenced on the PacBio Sequel platform.
Sequel data output and quality control
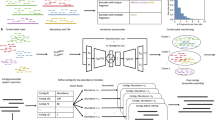
The PacBio sequence sequencing platform is based on single-molecule real-time (SMRT) sequencing technology. We used SMRT Link to preprocess and filter the original sequencing output data. The main parameters were Minimum Subread Length = 50, Maximum Subread Length = 15,000, Minimum Number of Passes = 3, Minimum Predicted Accuracy = 0.8, Minimum Read Score = 0.65, and Minimum Polished Isoform Accuracy 0.99; the remaining parameters were set to the default values. For ISO SEQ sequencing, the data preprocessing and filtering of SMRT Link mainly includes the following steps: the single-molecule polymerase reads are split to obtain subreads, and the subreads obtained from the same polymer reads are self-corrected and combined to form a circular consensus sequence (CCS). The CCSs were classified by detecting the presence of chimeric sequences, sequencing primers and 3′ poly-A sequences. CCSs with sequencing primer sequences, 3′ terminal poly-A sequences and no chimeric sequences were considered full-length nonchimeric (FLNC) sequences. The linker, primer and poly-A sequences corresponding to the FLNC sequences were removed, and all sequences were forward sequences. Then, the iterative clustering and error correction (ICE) tool of the SMRT Link software cluster module was used to cluster and correct multiple FLNC sequences of the same isoform to obtain the redundant isoform sequence. The non-FLNC sequences removed during the generation of the FLNC sequences were used to further correct the isoform sequence to enhance the sequence quality (cluster/polish) to obtain the polished isoform sequence. The polished isoforms were subjected to secondary sequence clustering by cd-hit-est software to further remove sequence redundancy. The sequences obtained in this step were the final non-redundant isoform set and were used for all subsequent analyses.
Functional annotation of transcripts
The obtained non-redundant transcript sequences were mapped to five databases to obtain transcript annotation information.
The NCBI nonredundant protein sequences (Nr) include the protein-coding sequence of GenBank gene, Protein Data Bank (PDB) protein database, Swiss-Prot protein sequence and Protein Information Resource (PIR) and the protein sequence of a database such as Protein Research Foundation (PRF). Gene Ontology (GO) is a set of internationally standardized classification systems for gene function descriptions. GO is divided into three ontologies: biological process (BP), molecular function (MF) and cellular component (CC). The basic unit of GO is the term27. EuKaryotic Orthologous Groups (KOG) is a gene orthology database for eukaryotes. KOG combines evolutionary relationships to classify homologous genes from different species into different ortholog clusters. Orthologous genes have the same function, so functional annotations can be directly transferred to other members of the same KOG cluster28. Kyoto Encyclopedia of Genes and Genomes (KEGG) includes a systematic analysis of the metabolic pathways of gene products and compounds in cells and a database of the functions of these gene products. It integrates data about genomes, chemical molecules, and biochemical systems. The KO (KEGG Ortholog) system links the various KEGG annotation systems, and KEGG has established a complete KO annotation system to perform functional annotation of the genomes or transcriptomes of newly sequenced species29. The Swiss-Prot Protein Sequence Database (Swiss-Prot) contains carefully examined and accurately annotated protein sequences from the EMBL nucleic acid sequence database27.
Digital gene expression library preparation and analysis
Using RSEM, the comparison results of Bowtie2 were counted, the number of reads from each sample compared to each transcript was obtained, and the fragments per kilobase per million bases (FPKM) was calculated. Paired-end reads from the same fragment were counted as one fragment, and then, the gene and transcript expression levels were obtained. A difference analysis was performed using the R language package DESeq2. To identify the differentially expressed genes (DEGs) between the SMT and CK libraries, the false discovery rate (FDR) method was used to determine the threshold of the P-value in multiple tests. The significance of the difference in gene expression was judged using a threshold of FDR < 0.05 and an absolute value of log2 fold change (FC, (condition 2/condition 1) for a gene) > 1 or log2FC < − 1. Then, GO enrichment analysis and KEGG pathway enrichment analysis were used to further annotate the genes expressed at different levels among the samples.
Results
SMRT sequencing data output
Using the PacBio SMRT sequencing technology, 283,232 polymerase reads were generated (Table 1). With this technology, circular sequences are obtained from single molecules, and the effective inserts produced during the sequencing process are the polymerase reads. The sequence fragments remaining from the polymerase reads after removing the adaptor sequences are called subreads. After preprocessing, 20.06 Gb of subreads were obtained (Table 2). Circular Consensus Sequences (CCSs) are sequences with low error rates obtained by mutual correction of multiple sequencing results. A total of 218,147 CCSs were obtained (Table 3). A total of 172,632 FLNC with a mean length of 1418 bp were assembled from CCSs with an N50 length of 1478 bp.
Transcript clustering analysis
Posttranscriptional modification of eukaryotic genes can produce a variety of different isoforms, and multiple FLNC sequences can be obtained from multiple sequencing of the same isoform. Using the ICE tools of the PacBio SMRT Link software to cluster the FLNC sequences and remove redundancy, a set of nonredundant isoform sequences can be obtained. Furthermore, the Quiver tool of the SMRT Link software was used to correct the isoform sequences obtained from the non-FLNC short sequences, and the obtained sequences were considered the polished isoforms. Then, a total of 10,364 polished isoform sequences with average length of 1493 bp were assembled from FLNC with an N50 length of 1648 bp. After the ICE clustering, 10,364 consensus isoforms were obtained, including 10,359 high-quality isoforms and five low-quality isoforms. The second sequence clustering was done by the software cd-hit-est to further remove the sequence redundancy. The sequence obtained in this step was the final non redundant isoform set and used for all subsequent analysis. Finally, a total of 9888 isoforms with average length of 1499 bp were assembled from polished isoform with an N50 length of 1659 bp, for further study.
Alternative splicing analysis
A total of 24 alternative splicing events were identified, as shown in Supplementary Table S1. However, because no reference genome was available for SMRT transcriptome sequencing in O. formosanus, we were unable to determine the types of the alternative splicing events.
Functional annotation of transcripts
In total, 9888 transcripts were annotated in the database: 3062 were annotated in GO, 4299 in KEGG (KO), 3536 in KOG, 5856 in Swiss-Prot and 7156 in NR. Moreover, 2719 transcripts were not annotated in any of the eight databases. The highest unigene hit rates were clearly in insect genomes; Zootermopsis nevadensis (3930, 54.92%), other (1582, 22.11%), Coptotermes formosanus (879, 12.28%), Odontotermes formosanus (159, 2.22%), Clastoptera arizonana (125, 1.75%), Cuerna arida (91, 1.27%), Reticulitermes flavipes (90, 1.26%), Blattella germanica (84, 1.17%), Periplaneta americana (82, 1.15%), Daphnia magna (67, 0.94%) and Locusta migratoria (67, 0.94%) accounted for unigenes with Nr annotations (Fig. 1).
The GO database is maintained by the Gene Ontology Consortium and includes a clearly structured, clearly defined, universal and controlled vocabulary to describe the roles of genes and gene products in any organism. The GO annotation system is a directed acyclic graph that includes three categories: BP, MF, and CC.
Using Blast2GO software, the transcriptome of O. formosanus was successfully mapped to the three main functional processes, which comprised 51 GO terms.
The GO gene functional classification system classified 3062 unigenes into three main functional ontologies (BP, MF, CC) (Fig. 2). Based on the GO analysis, 2047 genes were classified as BP, 1831 genes were classified as associated with CC, and 2427 were associated with MF. With regard to the BP category, the main subcategories were metabolic process (1517) and cellular process (1339), followed by single-organism process (994). In terms of MF, catalytic activity (1460) and binding (1429) were the most highly represented. In the CC category, cell part (1162), cell (1162) and membrane (931) were highly enriched. In total, 3536 sequences had KOG classifications. They were divided into 25 KOG groups using Diamond with the following comparison parameters: − f 6 − e 1e − 2—more-sensitive, and the other parameters were set to the default values (Fig. 3). Among the 25 KOG classifications, the largest group was posttranslational modification, protein turnover, chaperones (447), followed by general function prediction only (445) and cytoskeleton (411). These results may be related to the fact that there is still little data on O. formosanus in the KOG database at this stage. A total of 6352 unigenes with unknown function were obtained by sequencing and are presumed to represent new genes specific to O. formosanus.
DEGs in O. formosanus in response to S. marcescens infection
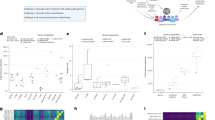
To explore the mechanism of S. marcescens infection, we tried to identify DEGs that were up- or downregulated in O. formosanus infected with S. marcescens by using PacBio Sequel sequencing. To maximize the accuracy of expression measurement, data from three biological replicates were combined, FPKM values were computed using the combined data, and the results were compared between the replicate SMT and CK groups (Fig. 4). DEGs between the SMT and CK groups were considered significant when FDR < 0.5 and log2FC > 1 or log2FC < − 1. Among the 9888 unigenes, a total of 259 DEGs were identified (Fig. 5), of which 132 were upregulated and 127 were downregulated.
Overview of DEGs. Heatmaps illustrating differences in normalized log signal intensity for the identified O. formosanus genes. SMT1, SMT2 and SMT3 are S. marcescens treatment groups, and CK1, CK2 and CK3 are control groups. Red and blue indicate genes expressed at high and low levels, respectively. The colors from blue to red indicate gradually increasing expression.
DEG analysis
GO was used to classify the functions of the O. formosanus DEGs. A total of 69 unigenes were divided into 26 functional groups. For the BP category, the dominant subcategories were metabolic process (35) and cellular process (29), followed by single-organism process (26). In the CC category, cell part (28), cell (28) and organelle (21) were highly enriched. In MF, catalytic activity (37) and binding (37) were the most highly represented terms (Fig. 6).
We mapped the DEGs to the typical KEGG pathways and then determined the biological pathways involved in response to S. marcescens treatment. In total, 94 DEGs were assigned to 115 KEGG pathways (Supplementary Table S2). Pathways with p-values < 0.05 were considered highly enriched (Table 4). The DEG enrichment analyses showed that the top three pathways enriched in up- or downregulated genes in response to S. marcescens treatment were “Adrenergic signaling in cardiomyocytes” (12 unigenes), “Cardiac muscle contraction” (12 unigenes) and “AMPK signaling pathway” (6 unigenes). These annotations provide valuable resources for further study of the specific processes, functions and pathways of O. formosanus under S. marcescens treatment.
Discussion
Termites are among the most famous pests in the world, causing great damage, and their infestations are difficult to control. There have been some studies on termites using the molecular technology. Harrison et al. revealed molecular basis of termite eusociality using hemimetabolous genomes30; Korb et al. compared the genomes of zootermopsis nevadensis and Macrotermes natalensis, and focused on genes involved in communication, immune defenses, mating biology and symbiosis that were likely important in termite social evolution31; Poulsen et al. reported the complementary symbiont contributioned to plant composition in a fungus-farming Macrotermes natalensis worker using the high-quality annotated draft genomes32; Terrapon et al. studied the molecular traces of alternative social organization in a termite genome by sequencing the genome and stage specific transcripts of Zootermopsis nevadensis (Blattodea) and comparing them with similar data of eusocial Hymenoptera33; Wu et al. clarified that soldier-biased gene expression in Reticulitermes flavipes implied functional specialization of the defensive caste using RNA sequencing34. However, few studies on the molecular biology of termites by ‘third-generation’ sequencing technology, such as the excavation, cloning and functional identification of genes, are available. Although, we have proved the insecticidal activity of S. marcescens through some bioassay experiments, but the specific insecticidal mechanisms of S. marcescens on O. formosanus are unclear, and the immune responses of O. formosanus to S. marcescens have not been clarified. Therefore, we use the third-generation high-throughput sequencing technology to explore these issues. In this study, third-generation SMRT sequencing technology was used to sequence for the first time the full-length transcriptome of termites under S. marcescens stress, and 10,364 full-length unigenes were obtained. In addition, 9888 unigenes were successfully annotated without a reference genome. These results indicate that the sequencing method used in this study yields rich data with high quality. These results will be helpful to clarify the action mechanisms of S. marcescens on O. formosanus, and explore the relationship between O. formosanus and S. marcescens. In addition, this study will identify the immune response of O. formosanus to S. marcescens, which will provide a theoretical foundation for the development of new immunosuppressants for O. formosanus. Furthermore, these high-quality unigene data will provide an important reference for subsequent functional genomics research, including termite-related gene discovery, cloning and functional identification.
Oxidative stress is associated with S. marcescens in O. formosanus
In vertebrates, ferritin is mainly present in the cytoplasm. Ferritin includes both heavy (H) chain and light (L) chain subunits35. When the content of free iron in cells is insufficient, ferritin can release stored iron ions and maintain normal life activities; when too many free iron ions are present in cells, large amounts of ferritin can be synthesized to store the excess iron ions. Thus, ferritin can prevent the damage to lipids and proteins caused by Fenton reactions and participate in the process of eliminating antioxidant stress36. As in vertebrates, ferritin in insects can also reduce antioxidant stress, acting as a protective agent against cytotoxicity. Insect ferritin is also composed of two subunits, heavy chain homolog (HCH) and light chain homolog (LCH) subunits37. These two subunits of ferritin are assembled depending on the ingestion, storage and release of iron ions in the body to form heteropolymers in different proportions to achieve the purpose of regulating iron ions. Overall, ferritin plays an important role in the various stages of organism proliferation, growth and development.
The most basic functions of ferritin in the body are to regulate iron metabolism and maintain iron homeostasis. It has been reported that ferritin in Drosophila played an important role in combating oxidative stress. When iron ions were added to the diet of Drosophila, the expression of the LCH and HCH subunits was significantly upregulated38,39. Similarly, ferritin played a role in regulating oxidative stress in mosquitoes40. In all animals, the expression of the LCH and HCH genes can be induced by dried blood40. As an iron chelator, ferritin can eliminate the Fenton reactions caused by excessive iron ions in insect food, thus protecting the body from reactive oxygen species (ROS). During the attack by invading pathogenic microorganisms, ferritin can store excess iron ions in the body, thereby inhibiting the growth of the pathogens. A previous study showed that the content of ferritin in the hemolymph increased significantly when beetle larvae were stimulated by fungi41, and another study showed that the content of ferritin in bumblebees increased significantly after iron stimulation42, and it has been reported that after LPS or bacterial stimulation, the expression of ferritin mRNA in horseshoe crabs was upregulated43. At present, the cause of the change in ferritin in O. formosanus under oxidative stress and how ferritin in O. formosanus helps to resist oxidative stress have not been reported. In our study, we found that the expression of ferritin was downregulated, which may be related to bacterial infection. This result might be due to a large quantity of iron ions obtaining from the host after bacterial infection, resulting in a decrease in the expression of ferritin.
Insect transferrin (TF) can be used as iron transporter, antibiotic or larval hormone regulatory protein to perform physiological functions. TF plays an important role in insect resistance to oxidative stress. It has been found that heat treatment, fungal invasion and H2O2 stimulation can lead to upregulated TF gene expression in the fat body of beetles and subsequently to the upregulation of TF gene expression in the hemolymph43. To prevent the harmful effects of iron, extracellular iron binds to TF and protects tissues from iron-induced oxidative stress damage44. In addition, insect TF plays an important role in fighting against bacterial infection45,46. Toe et al. stimulated Macrobrachium rosenbergii with Aeromonas hydrophila and confirmed that the content of TF in M. rosenbergii changed significantly47. In the present study, our results showed that the expression of TF in termites changed significantly after bacterial infection, and five insect TF genes were found to be downregulated (1.9–12.7-fold), while one gene was upregulated (23.31-fold), in O. formosanus following S. marcescens infection. The expression of the most TF genes was downregulated, which would lead to the reduction of transferrin synthesis, so that the excessive iron could not be eliminated in time, and further lead to the decline of antioxidant stress capacity of termites. This might be one of the reasons for the high insecticidal activity of S. marcescens on O. formosanus, which needed further experimental verification.
The MAPK pathway is associated with bacterial stress in O. formosanus
The mitogen-activated protein kinase (MAPK) pathway is an important protein-based signaling system in which eukaryotic cells transduce extracellular signals into cells to cause cellular responses48. An extensive and in-depth study of apoptosis-related genes and apoptosis signaling pathways found that the MAPK pathway plays an important role in regulating bacteria-induced apoptosis.
In vitro experiments have identified a large number of bacteria as mediators of apoptosis. These bacteria include Gram-positive bacteria, Gram-negative bacteria, and mycobacteria. Tseng et al.49 showed that the MAPK pathway was the main pathway in AGS apoptosis induced by Helicobacter pylori-derived Smase (Sphingomyelinase). Watanabe et al.50 used Porphyromonas gingivalis to infect human gingival epithelial cells (GECs), and the results showed that P. gingivalis acted selectively on the MAPK pathway. Dahan et al.51 used enterohemorrhagic Escherichia coli (EHEC) to infect intestinal epithelial cells (T84) for 3 h and showed the phosphorylation of three protein kinases (ERK1/2, P38 and JNK) in the MAPK family, demonstrating the role of the MAPK pathway in EHEC infection. These studies showed that the MAPK pathway played an important role in signal transduction related to bacterial-induced apoptosis. In our reseach, 14-3-3 protein bet/alpha (YWHAB) in the MAPK pathway was strongly downregulated (tenfold), meanwhile, YWHAB also existed in a immune pathway, JAN/STAT signaling pathway which was the branch of P13K-Akt signaling pathway in O. formosanus. This suggested that certain immune function of O. formosanus was inhibited to some extent by S. marcescens infection. This might be one of the reasons for the high insecticidal activity of S. marcescens.
MAPKs also play a role in phagocytosis and the survival of some microbial pathogens in macrophages. The process by which bacteria invade cells is often associated with activation of the MAPK pathway. For example, MAPKs in macrophages can be activated by bacterial lipopolysaccharide (LPS). Infection of epithelial cells with pathogenic Staphylococcus aureus can induce the activation of genes involved in the MAPK pathway (ERK1/2, P38 and JNK)52. In contrast, Yersinia pestis can reduce the activity of genes involved in the MAPK pathway (ERK1/2, P38 and JNK) in both macrophages and epithelial cells. In addition, curcumin has been found to inhibit the expression of macrophages by inhibiting MAPK/NF-κB signaling pathway53. This indirectly indicates that MAPK pathway is associated with phagocytosis. In our study, following S. marcescens infection, three genes involved in phagocytosis were upregulated (1–23-fold), and one gene was downregulated (22-fold). This phenomenon indicated that phagocytosis in termites was active but limited in the process of resistance to S. marcescens infection. Therefore, the MAPK pathway in O. formosanus may mediate S. marcescens-induced stress responses. However, further studies are needed to establish the possible regulatory role of the O. formosanus MAPK pathway during infection by S. marcescens. Exploring the relationship between O. formosanus and S. marcescens will help to elucidate the mechanisms of S. marcescens at the molecular level and provide an important theoretical basis for the development of new immunosuppressants for O. formosanus.
The AMPK pathway is associated with bacterial stress in O. formosanus
AMP-activated protein kinase (AMPK) is a serine/threonine kinase. It is a key metabolic and stress sensor/effector that is activated under conditions of nutrient deprivation, strenuous exercise or heat shock54, as well as an important regulator of cellular energy balance55.
The energy state of an organism is closely related to its rate of aging, and a low energy state delays aging in multiple species56. AMPK regulates senescence-associated pathways, including TOR/S6K, FOXO, sirtuins, cAMP response element binding protein (CREB)-regulated transcriptional coactivator (CRTCs), NF-κB signaling and other mediators57. Studies have shown that activation of AMPK is associated with prolonged life and improved health55. Funakoshi et al.58 showed that in fruit flies, the AMPK pathway could be activated by dietary restriction, which could extend the lifespan of fruit flies. Su et al.59 found that feeding alpha-ketoglutarate could activate AMPK signaling and prolong the lifespan of fruit flies. Piegholdt et al.60, by feeding prunetin, found that the prolongation of the lifespan of male fruit flies was dependent on the changes in energy homeostasis regulated by AMPK. Yang et al.61 found that dietary administration of β-guanidinopropionic acid at a concentration higher than 900 mmol led to a significant extension of the lifespan of Drosophila melanogaster in repeated experiments. Ulgherait et al.62 found that when the AMPK pathway was activated, cells cleared their aging and damaged components through autophagy. Furthermore, the lifespan of fruit flies was prolonged when the amount of AMPK in the intestine was increased. Burkewitz et al.57 showed that AMPK could prolong fruit fly lifespan by inhibiting the activity of the CRTC-1 protein. When AMPK content increased, CRTC-1 could not bind to CREB, thus ensuring the normal function of mitochondria throughout the body and prolonging life. Funakoshi et al.58 found that overexpression of lkb1 increased the level of phosphorylated AMPK, leading to prolong the fruit fly life. At present, studies of the AMPK pathway and related genes have not been reported in termites. In our study, following S. marcescens infection, three genes in the AMPK pathway were upregulated (1.15–23.28-fold), and three genes were downregulated (1.27–22.76-fold). These results indicate that the AMPK pathway in O. formosanus may mediate S. marcescens-induced stress responses. Other studies have found that the activation of AMPK pathway was closely related to energy metabolism. When insects were stimulated (pesticides, starvation, etc.), AMPK pathway would be activated to reduce the anabolism of cells, reduce the consumption of materials and energy, so as to maintain the survival of cells. It could also activate autophagy to resist the invasion of some intracellular pathogens. In our results, we found that there were significant changes in the expression of a large number of genes in this pathway, which may be related to the resistance of O. formosanus to S. marcescens invasion. Termites control energy metabolism by activating AMPK, reduce the loss, and use a lot of energy to resist S. marcescens invasion. However, further studies are needed to establish the possible regulatory role of the AMPK pathway in O. formosanus infection by S. marcescens.
References
Zhang, X. W. et al. Research progress inanatomic structures of digestive system and symbiotes in termites. J. Nanjing For. Univ. Nat. Sci. Ed. 39, 155–161 (2015).
Wang, Y. Z. et al. Induction feeding activity of fungus garden of Odontotermes formosanus (Isoptera: Termitidae) to foragers. Sci. Silv. Sin. 54, 117–123 (2018).
Xu, L. J. et al. Studies on the anatomical structures of labial gland in Odontotermes formosanus. J. Environ. Entomol. 41, 914–921 (2019).
Zhang, F. Y., Li, C. & Fan, D. F. Studies sunialpha 5FL against termites-laboratory tests of soil treatment. Pesticides 34, 18–20 (1995).
Huang, Q. Y., Lei, C. L. & Xue, D. Field evaluation of a fipronil bait against subterranean termite odontotermes formosanus (isoptera: termitidae). J. Econ. Entomol. 99, 455–461 (2006).
Zhang, Z. H. & Li, P. R. Investigation on the natural enemies of blackwing termites (Odontotermes formosanus shiraki) outside the termitarium. Acta Entomolo. Sin. 25, 227–230 (1982).
Yang, T. C., Mo, J. R., Pan, C. Y. & Cheng, J. Ultrastructure of the salivary gland in the worker of odontotermes formosanus (isoptera: termitidae). Sociobiology 48, 5–14 (2006).
Li, Y., Dong, Z. Y., Pan, D. Z. & Pan, C. H. Effects of subterranean termite nest architectures on earth embankment seepage and stability. Paddy Water Environ. 18, 367–384 (2020).
Huang, Y. Q., Lei, C. L. & Xue, D. Food choice of the underground termite Odontotermes formosanus. Sci. Silv. Sin. 41, 91–95 (2005).
Kirton, L. G. & Cheng, S. Ring-barking and root debarking of dipterocarp saplings by termites in an enrichment planting site in Malaysia. J. Trop. For. Sci. 19, 67–72 (2007).
Babashpour, S. et al. Characterization of a Chitinase (Chit62) from Serratia marcescens B4A and its efficacy as a bioshield against plant fungal pathogens. Biochem. Genet. 50, 722–735 (2012).
Wang, K. et al. Potential of chitinolytic Serratia marcescens strain JPP1 for biological control of Aspergillus parasiticus and Aflatoxin. Biomed. Res. Int. 2013, 397142. https://doi.org/10.1155/2013/397142 (2013).
Fu, R. J., Qi, X. L., Feng, K. & Xia, X. R. Identification and characters of a strain of Serratia marcescens isolated from the Odontotermes formosanus. J. Nanjing For. Univ. Nat. Sci. Ed. 43, 76–82 (2019).
Fu, R. J., Luo, J., Feng, K., Lu, X. Y. & Tang, F. Effects of Serratia marcescens (SM1) and its interaction with common biocontrol agents on the termite, Odontotermes formosanus (Shiraki). J. For. Res. https://doi.org/10.1007/s11676-020-01122-w (2020).
Fu, R. J. et al. Termite-killing components in Serratia marcescens (SM1). J. For. Res. https://doi.org/10.1007/s11676-020-01172-0 (2020).
Guo, Z. J. et al. MAPK-dependent hormonal signaling plasticity contributes to overcoming Bacillus thuringiensis toxin action in an insect host. Nat. Commun. 11, 3003. https://doi.org/10.1038/s41467-020-16608-8 (2020).
Kuang, Z., Boeke, J. D. & Canzar, S. The dynamic landscape of fission yeast meiosis alternative-splice isoforms. Genome Res. 27, 145–156 (2017).
Chen, S. Y., Deng, F. L., Jia, X. B., Li, C. & Lai, S. J. A transcriptome atlas of rabbit revealed by PacBio single-molecule long-read sequencing. Sci. Rep. 7, 7648. https://doi.org/10.1038/s41598-017-08138-2 (2017).
Abdel-Ghany, S. E. et al. A survey of the sorghum transcriptome using single-molecule long reads. Nat. Commun. 7, 11706. https://doi.org/10.1038/ncomms11706 (2016).
Wang, B. et al. Unveiling the complexity of the maize transcriptome by single-molecule long-read sequencing. Nat. Commun. 7, 11708. https://doi.org/10.1038/ncomms11708 (2016).
Dong, L. L. et al. Single-molecule real-time transcript sequencing facilitates common wheat genome annotation and grain transcriptome research. BMC Genomics 16, 1039. https://doi.org/10.1186/s12864-015-2257-y (2015).
Wang, M. J. et al. A global survey of alternative splicing in allopolyploid cotton: Landscape, complexity and regulation. New Phytol. 217, 163–178 (2018).
Liu, Y. H., Wang, L. & Yu, L. The principle and application of the single-molecule real-time sequencing technology. Yichuan (Hereditas) 37, 259–268 (2015).
Jin, Q., Pan, L. Q., Liu, T. & Hu, F. X. RNA-seq based on transcriptome reveals differ genetic expressing in Chlamys farreri exposed to carcinogen PAHs. Environ. Toxicol. Pharmacol. 39, 313–320 (2015).
Fei, Q. H. et al. Transcriptome profiling and phytohormone responses of Arabidopsis roots to different ambient temperatures. J. Plant Interact. 14, 314–323 (2019).
Shi, K. P. et al. RNA-seq reveals temporal differences in the transcriptome response to acute heat stress in the Atlantic salmon (Salmo salar). Comp. Biochem. Physiol. Part D Genomics Proteomics 30, 169–178 (2019).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 129–140 (2010).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280 (2004).
Harrison, M. C. et al. Hemimetabolous genomes reveal molecular basis of termite eusociality. Nat. Ecol. Evol. 2, 557–566 (2018).
Korb, J. et al. A genomic comparison of two termites with different social complexity. Front. Genet. 6, 9. https://doi.org/10.3389/fgene.2015.00009 (2015).
Poulsen, M. et al. Complementary symbiont contributions to plant decomposition in a fungus-farming termite. Proc. Natl. Acad. Sci. USA 111, 14500–14505 (2014).
Terrapon, N. et al. Molecular traces of alternative social organization in a termite genome. Nat. Commun. 5, 3636. https://doi.org/10.1038/ncomms4636 (2014).
Wu, T., Dhami, G. K. & Thompson, G. J. Soldier-biased gene expression in a subterranean termite implies functional specialization of the defensive caste. Evol. Dev. 20, 3–16 (2018).
Zhou, Q. Evolution of insect immunity. Fujian J. Agric. Sci. 22, 448–452 (2007).
Orino, K. & Watanabe, K. Molecular, physiological and clinical aspects of the iron storage protein ferritin. Vet. J. 178, 191–201 (2008).
Hamburger, A. E., West, A. P., Hamburger, Z. A., Hamburger, P. & Bjorkman, P. J. Crystal structure of a secreted insect ferritin reveals a symmetrical arrangement of heavy and light chains. J. Mol. Biol. 349, 558–569 (2005).
Missirlis, F. et al. Homeostatic mechanisms for iron storage revealed by genetic manipulations and live imaging of Drosophila ferritin. Genetics 177, 89–100 (2007).
Rosas-Arellano, A. et al. Ferritin assembly in enterocytes of Drosophila melanogaster. Int. J. Mol. Sci. 17, 27. https://doi.org/10.3390/ijms17020027 (2016).
Geiser, D. L., Chavez, C. A., Flores-Munguia, R., Winzerling, J. J. & Pham, D. Q. Aedes aegypti ferritin. Eur. J. Biochem. 270, 3667–3674 (2003).
Kim, B. Y. et al. Insect transferrin functions as an antioxidant protein in a beetle larva. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 150, 161–169 (2008).
Kozma, M. M., Chowrimootoo, G., Debnam, E. S., Epstein, O. & Srai, S. K. Developmental changes in mucosal iron binding proteins in the guinea pig. Expression of transferrin, H and L ferritin and binding of iron to a low molecular weight protein. Biochim. Biophys. Acta 1201, 229–234 (1994).
Ong, D. S. T., Wang, L., Zhu, Y., Ho, B. & Ding, J. L. The response of ferritin to lps and acute phase of pseudomonas infection. J. Endotoxin Res. 11, 267–280 (2005).
Du, X. L. et al. Apotransferrin is internalized and distributed in the same way as holotransferrin in K562 cells. J. Cell. Physiol. 201, 45–54 (2004).
Yun, E. Y. et al. Molecular cloning and characterization of a c DNA encoding a transferrin homolog from Bombyx mori. Biol. Chem. 380, 1455–1459 (1999).
Yoshiga, T. et al. Drosophila melanogaster transferrin. Cloning, deduced protein sequence, expression during the life cycle, gene localization and up-regulation on bacterial infection. Eur. J. Biochem. 260, 414–420 (2010).
Toe, A., Areechon, N. & Srisapoome, P. Molecular characterization and immunological response analysis of a novel transferrin-like, pacifastin heavy chain protein in giant freshwater prawn, Macrobrachium rosenbergii (de man, 1879). Fish Shellfish Immunol. 33, 801–812 (2012).
Ma, X. L. et al. Inhibition of p38mitogen -activated protein kinase decreases cardiomyocyte aoptosis and improves cardiac function after myocardial ischemia and reperfusion. Circulation 99, 1685–1691 (1999).
Tseng, H. J., Chan, C. C. & Chan, E. C. Sphingomyelinase of helicobacter pylori-induced cytotoxi city in AGS gastric epithelial cells via activation of JNK kinase. Bio. Biophys. Res. Commun. 314, 513–518 (2004).
Watababe, K., Yilmaz, O., Nakhjiri, S. F., Belton, C. M. & Lamont, R. J. Association of mitogen-activated protein kinase pathways with gingival epithelial cell responses to Porphyromonas gingivalis infection. Infect. Immun. 69, 6731–6737 (2001).
Dahan, S. et al. Enterohemorrhagic Escherichia coli infection induces interleukin-8 production via activation of mitogen-activated protein kinases and the transcription factors NF-κB and AP-1 in T84 cells. Infec. Immun. 70, 2304–2310 (2002).
Guan, L. Z. et al. The effect of Selenium on the transcriptional level of key factors in Nod2/MAPK/mTORs signaling pathways in S. aureus infected bovine mammary epithelial cells. Chinese J. Prev. Vet. Med. 41, 879–884 (2019).
Tang, B. & Zhou, Y. Y. Effect of curcumin on macrophage polarization and inflammatory response via TLR4-MAPK/NF-κB pathway. J. Electrocardiol. Circ. 38, 389–394 (2019).
Turnley, A. M. et al. Cellular distribution and developmental expression of AMP-activated protein kinase isoforms in mouse central nervous system. J. Neurochem. 72, 1707–1716 (1999).
Bitto, A., Wang, A. M., Bennett, C. F. & Kaeberlein, M. Biochemical genetic pathways that modulate aging in multiple species. Cold Spring Harbor Perspect. Med. 5, a025114. https://doi.org/10.1101/cshperspect.a025114 (2015).
Burkewitz, K., Zhang, Y. & Mair, W. B. AMPK at the nexus of energetics and aging. Cell Metab. 20, 10–25 (2014).
Burkewitz, K. et al. Neuronal CRTC-1 governs systemic mitochondrial metabolism and lifespan via a catecholamine signal. Cell 160, 842–855 (2015).
Funakoshi, M. et al. A gain-of-function screen identifies wdb and lkb1 as lifespan-extending genes in Drosophila. Biochem. Biophys. Res. Commun. 405, 667–672 (2011).
Su, Y. et al. Alpha-ketoglutarate extends Drosophila lifespan by inhibiting mTOR and activating AMPK. Aging 11, 4183–4197 (2019).
Piegholdt, S., Rimbach, G. & Wagner, A. E. The phytoestrogen prunetin affects body composition and improves fitness and lifespan in male Drosophila melanogaster. Faseb J. 30, 948–958 (2015).
Yang, S. et al. β-Guanidinopropionic acid extends the lifespan of Drosophila melanogaster via an AMP-activated protein kinase-dependent increase in autophagy. Aging Cell 14, 1024–1033 (2015).
Ulgherait, M., Rana, A., Rera, M., Graniel, J. & Walker, D. W. AMPK modulates tissue and organismal aging in a non -cell-autonomous manner. Cell Rep. 8, 1767–1780 (2014).
Acknowledgements
We gratefully acknowledge Dr. Meng Xu of the Nanjing Forestry University for their assistance. This research was supported by Postgraduate Research & Practice Innovation Program of Jiangsu Province [Grant numbers KYCX20_0871] and the Priority Academic Program Development Fund of Jiangsu Higher Education Institutions. There was no additional external funding received for this study. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
K.F. performed the experiments, analyzed the data and drafted the manuscript. X.Y.L. collected samples and analyzed the data. J.L. collected samples. F.T. conceived and designed the experiments, reviewed manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Feng, K., Lu, X., Luo, J. et al. SMRT sequencing of the full-length transcriptome of Odontotermes formosanus (Shiraki) under Serratia marcescens treatment. Sci Rep 10, 15909 (2020). https://doi.org/10.1038/s41598-020-73075-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73075-3
This article is cited by
-
Unraveling metagenomics through long-read sequencing: a comprehensive review
Journal of Translational Medicine (2024)
-
Effect of Glycosylation on the Physicochemical Properties, Structure and Iron Bioavailability of Ferritin Extracted from Tegillarca granosa
Journal of Ocean University of China (2023)
-
Full-length SMRT transcriptome sequencing and microsatellite characterization in Paulownia catalpifolia
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.