Abstract
Despite the importance of hypothalamic neurocircuits in regulating homeostatic and survival-related behaviors, our understanding of the intrinsic molecular identities of neural components involved in these complex multi-synaptic interactions remains limited. In this study, we constructed a Cre recombinase-dependent pseudorabies virus (PRVs) capable of crossing synapses, coupled with transcriptome analysis of single upstream neurons post-infection. By utilizing this retrograde nuclear Connect-seq (nuConnect-seq) approach, we generated a single nuclei RNA-seq (snRNA-seq) dataset of 1,533 cells derived from the hypothalamus of CRH-IRES-Cre (CRH-Cre) mice. To ensure the technical validity of our nuConnect-seq dataset, we employed a label transfer technique against an integrated reference dataset of postnatal mouse hypothalamus comprising 152,524 QC-passed cells. The uniqueness of our approach lies in the integration of diverse datasets for validation, providing a more nuanced diversity of hypothalamic cell types. The presented validated dataset may deepen our understanding of hypothalamic neurocircuits and underscore the essential role of comprehensive integrated transcriptomic data for technical validity.
Similar content being viewed by others
Background & Summary
Understanding the complexity of hypothalamic neurocircuits and identifying their molecular signatures is crucial for comprehending the coordinated physiological responses1,2,3. In past years, the high-throughput single-cell RNA sequencing (scRNA-seq) analytical and experimental approaches have facilitated the understanding of the transcriptional cataloging of cell types in the hypothalamus and many other adjacent tissues of the brain4,5,6. However, very little is known about the molecular identification of neural components involved in complex multi-synaptic circuits. In addition, there is a need for a more comprehensive characterization of cell type populations and their distribution to understand the neural circuit properties.
Recent neuroanatomical tracer studies have provided invaluable insights into visualizing and defining complex presynaptic neural circuits7,8,9. Furthermore, these studies have confirmed the faithful retrograde transmission of pseudorabies viruses (PRVs) between interconnected neurons via direct cell-to-cell contact, rather than extracellular diffusion10,11,12. Upon infecting neurons, PRVs can trigger the activation of adjacent astrocytes and microglia, which then function to sequester heavily infected or dying neurons13,14. It’s worth noting that glial cells lack the necessary machinery to generate and release infectious (enveloped) virions. The glial cells obtained through Connect-seq likely represent neighboring cells in close proximity to neurons, with specific projections to CRH neurons within the PVN of CRH-Cre mice8,15. Here, we focus on a retrograde Connect-seq approach to produce a hypothalamic single nuclei RNA-seq (snRNA-seq) dataset from CRH-IRES-Cre (CRH-Cre) mice. We refer to this approach as “nuclear Connect-seq” (nuConnect-seq), in contrast to the original whole-cell-based Connect-seq method (Fig. 1a–c).
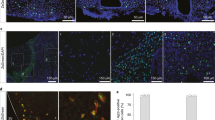
Generation of a nuConnect-seq-derived dataset. (a) A retrograde nuConnect-seq approach was used to generate a single nuclei RNA-seq (snRNA-seq) dataset from CRH-IRES-Cre (CRH-Cre) mice. (b) PRVB180, a pseudorabies virus expressing TK (thymidine kinase)-GFP (green fluorescence protein) in the Cre+ neurons and their upstream. (c) The flow of nuConnect-seq method. (d) UMAP plot showing the manually curated cell types of the presented dataset. (e) Violin plot showing the relative expression of neuronal and non-neuronal gene markers.
In this study, we combine the use of conditional neurotropic viruses that cross synapses with transcriptome analysis of single upstream neurons infected with the virus to identify the molecular identities of upstream CRH neurons in hypothalamic brain regions8 (see Methods, Fig. 1). Thus, we created a nuConnect-seq-derived snRNA-seq dataset of 1,533 cells from the hypothalamus and the bed nucleus of CRH-IRES-Cre (CRH-Cre) mice. To further evaluate the technical robustness of our lab-generated nuConnect-seq dataset, we processed and re-analyzed 10 publicly available scRNA-seq transcriptomic datasets, each compromising unique hypothalamic regions of mice. After merging these datasets, we performed an integrative single-cell transcriptomic analysis through anchor-based mapping, comprising 152,524 QC-passed cells with a total of 36 distinct neuronal and non-neuronal cell types (Fig. 2). Then we conducted cell type label transfer between our nuConnect-seq-derived dataset and the integrated transcriptomic reference dataset.
Schematic representation of the generation of an integrated reference dataset of postnatal mouse hypothalamus for technical validation of nuConnect-seq-derived dataset. Ten independent datasets were integrated and batch-corrected in an anchor-based manner. Cell-level metadata was manually curated and standardized for all the datasets. A uniform informatic pipeline was used for clustering and cell type annotation.
By following the Seurat label transfer workflow as suggested in the “Mapping and annotating query datasets” vignette, we validated our newly generated nuConnect-sec dataset with the reference map of over 100,000 hypothalamic single nuclei (Figs. 3, 4). Through this approach, we effectively demonstrate the merits of utilizing label transfer to address the inherent limitations posed by the relatively small nuConnect-seq datasets. This comparison against a substantially larger reference dataset highlights the advantages of leveraging label transfer. Finally, our validated nuConnect-seq dataset, along with a large-scale single-cell transcriptomic reference dataset with harmonized cell-level metadata, can serve as an asset for exploring genes across the hypothalamus.
Cell type annotation of the integrated reference dataset of young adult mouse hypothalamus. (a) UMAP plots showing neuronal or non-neuronal cells (left) classified by combined expression of pan-neuronal markers (right). (b) Relative expression of pan-neuronal and non-neuronal marker genes is shown. (c) Final UMAP plot showing subtypes of neuronal and non-neuronal cells.
Mapping and transferring labels for technical validation of nuConnect-seq-derived dataset. (a) Schematic representation showing identified anchors, which facilitate the transfer of discrete labels between integrated (reference) and lab-generated nuConnect-seq (query) datasets. (b) UMAP plots show the actual and predicted cell types on the nuConnect-seq dataset by projecting cells of the nuConnect-seq dataset onto UMAP embeddings. (c) Histogram plot showing the distribution of predicted cell type scores across the predicted cell types in the validation dataset. (d) UMAP plot showing neuronal and non-neuronal cells of the final merged (integrated and nuConnect-seq dataset).
Methods
Mice
To produce nuConnect-seq data, CRH-IRES-Cre (CRH-Cre; stock no, 012704) mice were purchased from The Jackson Laboratory. All procedures involving mice were approved by the Ajou University Institutional Animal Care and Use Committee and/or DGIST Institutional Animal Care and Use Committee.
PRV (pseudorabies virus)
PRVB180 was cultivated following established protocols as detailed in prior studies8,16. In brief, PRV180 was propagated by infecting PK15 cells (obtained from the American Type Culture Collection) with the virus, utilizing a multiplicity of infections ranging from 0.1 to 0.01. Following infection, notable cytopathic effects became apparent within approximately two days. The infected cells were subsequently collected through scraping, and the cellular material was swiftly frozen using liquid nitrogen, then rapidly thawed within a water bath at 37 °C. After three freeze-thaw cycles, cellular debris was eliminated through centrifugation, performed twice at 1000 g for 5 minutes each. The resulting supernatant was employed for subsequent experiments. Viral stocks were quantified through standard plaque assays carried out on PK15 cells17, with the titer denoted in terms of plaque-forming units (pfu).
Stereotaxic injections
Viruses were injected into the PVN of CRH-Cre mice as described previously8. Viruses were injected into the brain using a Stereotaxic Alignment System (David Kopf Instruments) with inhalation anesthesia of 2% isoflurane. Briefly, 1 µL of PRVs (PRVB180; 1 to 1.5 × 106 p.f.u.) were loaded into a 1-µL syringe and injected bilaterally into the brain at a rate of 100 nL/min. The needle was inserted into the PVN based on a stereotaxic atlas (anterior-posterior (AP), −0.4 mm; medial-lateral (ML), ±0.3 mm; dorsal-ventral (DV), −5.0 mm)18. After recovery, animals were singly housed with regular 12-h dark/light cycles, and food and water were provided ad libitum.
Isolation of single nuclei
nuConnect-seq experiments were performed as previously described8 with modifications for nuclei isolations, Adult 8-week-old CRH-Cre male mice were injected with PRVB180. After 3 days, mice were euthanized by cervical dislocation, and the brain was quickly removed. The hypothalamus was carefully micro-dissected under a microscope and immediately placed into a nuclei isolation medium (sucrose 0.25 M, KCl 25 mM, MgCl2 5 mM, Tris-Cl 10 mM, dithiothreitol, 0.1% Triton). Tissue underwent Dounce homogenization, facilitating the mechanical isolation of nuclei from cells19. Hoechst 33342 nucleic acid stain (5 μM, Life Technologies) was introduced into the media to enhance nuclei visualization. Subsequently, the samples were washed, reconstituted in a nuclei storage buffer (composed of sucrose, 5 mM MgCl2, and 10 mM TrisCl), and then filtered. It’s important to note that solutions and samples were maintained cold throughout the procedural steps. Then, PRV-infected single nuclei were isolated based on the fluorescence emitted by TK-GFP using flow cytometry (FACSAria II; BD Biosciences) according to methods described previously8, with some modifications for isolating single nuclei20. Hoechst-positive nuclei were gated first, followed by the exclusion of debris using forward and side scatter pulse area parameters (FSC-A and SSC-A), and the exclusion of aggregates using pulse width (FSC-W and SSC-W). Next, single nuclei with high GFP were placed in ice-cold methanol and then stored at −80 °C immediately. Single-nuclei suspensions obtained from the cortex of CRH-Cre mice injected with PRVB180 were analyzed first in the initial experiments to determine the threshold for fluorescence, and then single-nuclei suspensions from CRH-Cre mice were analyzed to sort GFP+ fluorescent nuclei.
Single nuclei capture
FACS-sorted single nuclei were loaded onto an independent single channel of a Chromium Controller (10 × Genomics) single-cell platform. In summary, we gathered a cumulative total of 10,000 single nuclei from five CRH-Cre mice, which were subsequently pooled and loaded onto the capture setup utilizing the Chromium Single Cell 3′ Reagent kit with, v2 Chemistry (10 × Genomics). Following the capture process and cellular lysis, complementary DNA was synthesized and amplified through a 14-cycle process following the manufacturer’s instructions (10 × Genomics). The amplified cDNA was then used to create an Illumina sequencing library, which was subsequently sequenced on a single lane of a NovaSeq. 6000 (Illumina). To facilitate FASTQ generation and subsequent alignments, the initial Illumina basecall files (*.bcl) were converted to FASTQs using Cell Ranger v.1.3 (10 × Genomics), which uses bcl2fastq v.2.17.1.14. FASTQ files were then aligned to the mm10 genome and transcriptome using the Cell Ranger v.1.3 pipeline. This pipeline results in the generation of a ‘gene vs cell’ expression matrix.
nuConnect-seq-derived data
The Connect-seq dataset was processed using the Seurat package v.4.3.021 in R v.4.2.1 (as for integrated reference dataset) (Supplementary Fig. S1). After loading the dataset into R, the Seurat object was created using the “CreateSeuratObject” function. Before main analyses, metadata for each cell was meticulously added, encapsulating various parameters including age (8 weeks), sex (male), region (whole hypothalamus + BNST), technological method (FACS), and mouse strain (CRH-Cre). This cell-level metadata standardization ensures consistency and clarity in our dataset. To mitigate the potential biases arising from disparate sequencing depths across cells, the dataset was normalized using the “LogNormalize” method in Seurat with default parameters. Then the top 2000 variable genes were identified using the FindvariableFeatures() function.
Using these top 2000 variable genes nCount_RNA was regressed out in the scaling step followed by PCA (principal component analysis). Then we employed the UMAP (uniform manifold approximation and projection) on the top 20 principal components in a two-dimensional space to visualize a clear depiction of cell distribution. Further graph-based clustering was performed by using the FindCluster() function in Seurat. For cell clustering, resolution was set as 0.25 to ensure clear granularity and avoid overlap of comparable cells, and 8 cell clusters were identified using specific marker genes, including IPCs (Mki67+), OPC (Pdgfra+), Astro (Agt+), Endo1 (Pecam+), Endo2 (Itm2a+), Mac (C1qa+), MO (Mog+) and one neuronal cluster expressing Syp (Fig. 1).
Data Records
Raw counts data and processed sequencing data generated in this study have been submitted to the NCBI Gene Expression Omnibus (GEO) under the accession number GSE24076522. Our nuConnect-seq dataset is mainly composed of three primary files, (1) “features.tsv.gz”, a tab-separated values (TSV) file that is compressed using gzip, provides a comprehensive catalog of the genomic features considered in the nuConnect-seq dataset. (2) The “barcodes.tsv.gz” file, compressed using gzip, contains the information of unique molecule identifiers that are very important for cell identification. (3) The “matrix.mtx.gz” file represents the sparse matrix where each row represents the genes, and each column represents the individual cell. All these files with the final annotated nuConnect-seq dataset named as “Validation. rds” file containing the information about cell-level metadata, are also deposited in figshare, (https://doi.org/10.6084/m9.figshare.21981251.v1)23. Furthermore, the matrix data derived from the 10X CellRanger metrics has been provided in Supplementary Table 1.
Our integrated transcriptomic reference dataset named the “final_hypo_ann.rds” is accessible in the figshare repository (https://doi.org/10.6084/m9.figshare.21981251.v1)23. The data can be used to construct quality control (QC) metrics and obtain standardized cell-level metadata including information on age, brain area, sex, single-cell technology, mouse strain, and manually annotated neuronal and non-neuronal cell types.
Technical Validation
Generation of the integrated reference scRNA-seq dataset
To assess the technical validity of our lab-generated nuConnect-seq derived single nuclei RNA-seq (snRNA-seq) dataset, we collected ten publicly available scRNA-seq datasets from the NCBI GEO under the accession numbers GSE8754424, GSE13235525, GSE9337426, GSE1135765, GSE12683627, GSE7467228,29,30, GSE1250656, GSE14669231, GSE1399238 and https://doi.org/10.17632/ypx3sw2f7c.1 (Mendeley Data)32. These datasets of mice originated from various hypothalamic brain regions including arcuate hypothalamus and median eminence (ARC-ME), lateral hypothalamus (LH), medial preoptic area (MPOA), midline hypothalamus, posterior hypothalamus, ventromedial hypothalamus (VMHvl), and the bed nucleus of the stria terminalis (BNST), with ages ranging from P14 to 14 weeks. The datasets were obtained from mice of various strains including C57BL/6, C57 × D2F1, Agrp-Cre × Ai6, Nr5a1-Cre × Ai14, and CRH-Cre5,6,8,24,25,26,27,31,33, which were further processed using the Seurat package34 in R (v. 4.2.1).
To facilitate the re-use of the integrated data, we manually curated cell-level attributes, such as study, age, brain region, sex, and strain of mouse of each dataset, and used standardized texts for metadata (Fig. 2). After merging all these datasets, a threshold of 15% mitochondrial content was chosen based on the observed distribution of mitochondrial percentages across the cells followed by feature counts over 2,800 or less than 100 were filtered out (Supplementary Fig. 2a,b). Cells with very low gene counts also tend to have an abnormally high percentage of mitochondrial genes. But we also were specific regarding the inherent nature of brain cells that specific specialized and quiescent neurons in the brain may have a reduced gene expression profile, leading to inherently lower detection rates. Our threshold ensures that these critical cells are not inadvertently filtered out. Our filtering approach, therefore, not only retains potentially quiescent or low transcript count cells but also effectively removes cells likely to be poor quality or damaged. A total of 152,524-QC passed cells were subsequently included for further analyses, and variable features of these cells were identified after normalization. The FindIntegrateionAnchors() function of the Seurat was applied to merged cells to find the shared cell population across datasets. Then we identified the cross-dataset pairs of cells with similar biological states (“anchors”) to correct for technical differences between datasets. The number of principal components (PCs) of the dataset was determined using the Elbow plot before the clustering of cells (Supplementary Fig. 2c). To comprehensively identify distinct cell populations, cells of the integrated dataset were clustered into distinct cell populations using the FindClusters() function by implementing multiple resolutions (Supplementary Fig. 2d).
Supplementary Fig S3 shows QC-passed cells colored by different cell-level attributes before and after batch removal, further divided by cell type proportion across all datasets. We quantitatively evaluated the quality of clustering results for each resolution to examine whether subpopulations of cells could be split or merged. To ensure clear granularity and avoid overlap of comparable cells in different clusters, we used a resolution of 0.25 for downstream analyses. The resulting Louvain clusters were visualized in a two-dimensional UMAP representation and were manually annotated with known cell types using canonical neural and non-neural marker genes, as well as signatures chosen in the original publications. We first distinguished neuronal cells from non-neuronal cells using the known pan-neural genes (Snap25, Syp, Elval2, Slc17a6, and Slc32a1) and canonical non-neural genes (Olig1, Cldn5, C1qa, Sox9, and Mustn1) (Fig. 3a,b).
Characterization of cell subpopulations
To identify the various populations of excitatory and inhibitory neurons, we sub-setted the neuronal cell cluster of the integrated dataset and performed a second round of dimensional reduction and clustering using the FindVariableFeatures(), ScaleData(), RunPCA(), and RunUMAP() functions of the Seurat package. To group the cells with similar transcriptional profiles, we applied the resolution of the FindClusters() function as 0.3. We next examined the expression patterns of markers for excitatory and inhibitory neurons, which are Slc17a6 for glutamatergic neurons and Slc32a1, Gad1, and Gad2 for GABAergic neurons, respectively35. Of the 13 identified neuronal clusters, there are 4 glutamatergic clusters, 8 GABAergic clusters, and one cluster neither glutamatergic nor GABAergic, but Cck + (Supplementary Fig. S4a,b). Overall, within the identified neuronal cell populations, clusters can be identified by the unique combination of genes that are expressed at different levels. Most neuronal clusters contain potential sub-type markers and others are defined by combinatorial expression of marker genes. For example, GABA7 is specified with the high expression of arginine vasopressin (Avp), a neuropeptide controlling osmoregulation, and low expression of pro-opiomelanocortin (Pomc) neurons. Similarly, a neuropeptide such as Agrp that controls feeding, is highly expressed in the GABA6 cluster. Similarly, Slc1a3, Nr4a2, Lhx1, Adcyap1, Lhx8, Crabp1, and Npy2r are each expressed in only one neuron subtype, while Htr2c, Six3, Meis2, Tubb3, and Tmem163 are each expressed in two or four neurons’ subtypes with distinct patterns (Supplementary Fig. S4c).
Next, we identified a total of 17 distinct clusters of non-neural cells. Oligodendrocytes were found using markers, such as Olig1/2, Pdgfra, Fyn, and Mobp, all of which show a gradient of gene expression in postnatal development24,31,36,37 (Supplementary Fig. S4d,e). These cells also include oligodendrocytes progenitor cells (OPC), newly formed oligodendrocytes (NFO), and myelinating oligodendrocytes (MO) that indicate axon myelination is still taking in the adult hypothalamus. We also identified one cluster as intermediate progenitor cells (IPCs) by the expression of Hmgb2, which encodes a chromatin-related transcription activator38. OPCs are another type of neural progenitor cells that express Pdgfra, which regulates processes of cell proliferation, differentiation, and migration through the Wnt and Notch signaling pathways. Subsequently, we also identified three distinct subtypes of astrocytes (Astro1, 2, and 3) based on overall levels of relative gene expression from none or low to high (Supplementary Fig. S4e,f). Each of these astrocyte subtypes expresses Agt, which regulates synaptic communication and cerebral blood flow at varying levels. The Astro1 cell cluster exhibits an elevated expression of proteins like Gfap, pivotal for cytoskeletal function, while the Astro2 cell cluster has a high expression of C1qa and a low expression of Unc13c39,40. The cells of Astro3 highly express Unc13c and Rgcc, which encode for membrane fusion and cell cycle regulation41. Tanycytes (Tany) and tanycyte-like ependymal cells (Tany-like ependy) were also found using the expression levels of Rax and Ccdc153, respectively, that have role in cell development and response to injury24. In addition, we found other distinct clusters readily identifiable as macrophages (Mac: Mrc1+), microglia (Micro: Cx3cr1+), mural cells (Mustn1+), two populations of endothelial cells (Endo1/2: Pecam1+), vascular leptomeningeal cells or VLMCs (Dcn+) and one erythrocyte representing cluster (Eryth: Alas2+). These findings of major non-neural cells obtained from the hypothalamus at varying postnatal stages closely align with previous scRNA-seq analyses from mouse brains8,31,42,43. Similarly, myelinating oligodendrocytes and tanycytes were subsetted to run a second round of dimensional reduction and clustering and thus to characterize cellular heterogeneity, (Supplementary Fig. S4g,h)44,45,46. Subtyping of myelinating oligodendrocytes (MO) revealed 3 clusters: (1) early mature oligodendrocytes (EMOL) expressing Egr2 (Early Growth Response 2; also known as Krox20), during oligodendrocytes maturation, (2) mature oligodendrocytes (MOL) expressing late oligodendrocyte differentiation genes, such as Klk6 and Anxa5, and (3) myelin-forming oligodendrocytes (Myl-OL) expressing genes involved in myelin formation (Supplementary Fig. S4g). Similarly, tanycytes were subdivided into four subtypes – α1, α2, β1, and β2 – based on their gene expression. Interestingly, we found one cell cluster expressing S100b, which is known to be involved in the regulation and migration of astrocytes47 (Supplementary Fig. S4h). This rigorous analysis yielded six additional sub-clusters, bringing the total number of non-neuronal subtypes to 23, further emphasizing the robust cellular differentiation of our integrated dataset. Finally, our large-scale integrated dataset surpasses the scope of individual studies by encompassing a broader array of a total of 36 distinct cell types that encompass all major and sub-cell types found in the hypothalamic regions, including glial-like cells such as ependymal cells, tanycytes, astrocytes, oligodendrocytes, previously undefined IPCs and neuronal cells with their sub-populations, thereby providing unprecedented perspective and insight into the hypothalamic molecular landscape. Furthermore, insight information on each cluster’s characteristics is detailed in Supplementary Table 2.
Validation of transcriptional profiles
In our study, we utilized the integrated transcriptomic dataset as an established benchmarked reference to ascertain the accuracy of our nuConnect-seq dataset. The manually annotated cell types of the nuConnect-seq dataset were compared with the cell types of the integrated dataset using the ‘label transfer’ method from Seurat via the FindTransferAnchors() and TransferData() functions (Fig. 4a). These comparative approaches demonstrate that the projected cell types in our nuConnect-seq dataset closely match the cell types of the integrated reference dataset, validating the technical validity of the generated nuConnect-seq dataset (Fig. 4b). The histogram, merged UMAP and feature plots displaying the strength of the cell type predictions are shown in Fig. 4c,d, Supplementary Fig. S5 confirms the similarity of transcriptional profiles between neuronal and non-neuronal cell types in the two datasets. This transcriptional profiling comparison with an integrated transcriptomic reference dataset highlights the reliability and accuracy of our nuConnect-seq dataset.
Code availability
The R script used to generate the integrated dataset, conduct validation dataset analysis, and produce relevant plots can be found at: https://github.com/Junaid13913/PostnatalMouseHypothalamus_SingleCellTranscriptome. Our final datasets with UMAP embeddings are available at the figshare repository: (https://doi.org/10.6084/m9.figshare.21981251.v1)23.
References
Cooper-Kuhn, C. M. et al. Impaired adult neurogenesis in mice lacking the transcription factor E2F1. Mol Cell Neurosci 21, 312–323, https://doi.org/10.1006/mcne.2002.1176 (2002).
Kleinridders, A., Konner, A. C. & Bruning, J. C. CNS-targets in control of energy and glucose homeostasis. Curr Opin Pharmacol 9, 794–804, https://doi.org/10.1016/j.coph.2009.10.006 (2009).
Jovanovic, P. et al. A sex-specific thermogenic neurocircuit induced by predator smell recruiting cholecystokinin neurons in the dorsomedial hypothalamus. Nat Commun 14, 4937, https://doi.org/10.1038/s41467-023-40484-7 (2023).
Mu, Q., Chen, Y. & Wang, J. Deciphering Brain Complexity Using Single-cell Sequencing. Genomics Proteomics Bioinformatics 17, 344–366, https://doi.org/10.1016/j.gpb.2018.07.007 (2019).
Moffitt, J. R. et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 362, https://doi.org/10.1126/science.aau5324 (2018).
Mickelsen, L. E. et al. Single-cell transcriptomic analysis of the lateral hypothalamic area reveals molecularly distinct populations of inhibitory and excitatory neurons. Nat Neurosci 22, 642–656, https://doi.org/10.1038/s41593-019-0349-8 (2019).
Pomeranz, L. E. et al. Gene Expression Profiling with Cre-Conditional Pseudorabies Virus Reveals a Subset of Midbrain Neurons That Participate in Reward Circuitry. J Neurosci 37, 4128–4144, https://doi.org/10.1523/JNEUROSCI.3193-16.2017 (2017).
Hanchate, N. K. et al. Connect-seq to superimpose molecular on anatomical neural circuit maps. Proc Natl Acad Sci USA 117, 4375–4384, https://doi.org/10.1073/pnas.1912176117 (2020).
Jia, F. et al. Optimization of the Fluorescent Protein Expression Level Based on Pseudorabies Virus Bartha Strain for Neural Circuit Tracing. Front Neuroanat 13, 63, https://doi.org/10.3389/fnana.2019.00063 (2019).
Ekstrand, M. I., Enquist, L. W. & Pomeranz, L. E. The alpha-herpesviruses: molecular pathfinders in nervous system circuits. Trends Mol Med 14, 134–140, https://doi.org/10.1016/j.molmed.2007.12.008 (2008).
Ugolini, G. Advances in viral transneuronal tracing. J Neurosci Methods 194, 2–20, https://doi.org/10.1016/j.jneumeth.2009.12.001 (2010).
Nguyen, T. D. et al. Targeted single-neuron infection with rabies virus for transneuronal multisynaptic tracing. J Neurosci Methods 209, 367–370, https://doi.org/10.1016/j.jneumeth.2012.06.019 (2012).
Card, J. P. et al. Pseudorabies virus infection of the rat central nervous system: ultrastructural characterization of viral replication, transport, and pathogenesis. J Neurosci 13, 2515–2539, https://doi.org/10.1523/JNEUROSCI.13-06-02515.1993 (1993).
Rinaman, L., Card, J. P. & Enquist, L. W. Spatiotemporal responses of astrocytes, ramified microglia, and brain macrophages to central neuronal infection with pseudorabies virus. J Neurosci 13, 685–702, https://doi.org/10.1523/JNEUROSCI.13-02-00685.1993 (1993).
Briscoe, J. & Marin, O. Looking at neurodevelopment through a big data lens. Science 369, https://doi.org/10.1126/science.aaz8627 (2020).
Lee, E. J. et al. A psychological stressor conveyed by appetite-linked neurons. Sci Adv 6, eaay5366, https://doi.org/10.1126/sciadv.aay5366 (2020).
Card, J. P. & Enquist, L. W. Transneuronal circuit analysis with pseudorabies viruses. Curr Protoc Neurosci 68, 1 5 1–1 5 39, https://doi.org/10.1002/0471142301.ns0105s68 (2014).
Keith, B. J., Franklin, G. P. The Mouse Brain in Stereotaxic Coordinates. (Elsevier Science, 2008).
Grindberg, R. V. et al. RNA-sequencing from single nuclei. Proc Natl Acad Sci USA 110, 19802–19807, https://doi.org/10.1073/pnas.1319700110 (2013).
Lacar, B. et al. Nuclear RNA-seq of single neurons reveals molecular signatures of activation. Nat Commun 7, 11022, https://doi.org/10.1038/ncomms11022 (2016).
Stuart, T. et al. Comprehensive Integration of Single-Cell Data. Cell 177, 1888–1902 e1821, https://doi.org/10.1016/j.cell.2019.05.031 (2019).
Junaid, M., Lim, S. B. & Lee, E. J. GEO. https://identifiers.org/geo/GSE240765 (2023).
Junaid, M., Choe, H. K., Lee, E. J. & Lim, S. B. Large-scale single-cell RNA-seq characterizes neural stem cells and progenitor cells in postnatal and young adult mouse hypothalamus. figshare https://doi.org/10.6084/m9.figshare.21981251.v1 (2023).
Chen, R., Wu, X., Jiang, L. & Zhang, Y. Single-Cell RNA-Seq Reveals Hypothalamic Cell Diversity. Cell Rep 18, 3227–3241, https://doi.org/10.1016/j.celrep.2017.03.004 (2017).
Kim, D. W. et al. The cellular and molecular landscape of hypothalamic patterning and differentiation from embryonic to late postnatal development. Nat Commun 11, 4360, https://doi.org/10.1038/s41467-020-18231-z (2020).
Campbell, J. N. et al. A molecular census of arcuate hypothalamus and median eminence cell types. Nat Neurosci 20, 484–496, https://doi.org/10.1038/nn.4495 (2017).
Welch, J. D. et al. Single-Cell Multi-omic Integration Compares and Contrasts Features of Brain Cell Identity. Cell 177, 1873–1887 e1817, https://doi.org/10.1016/j.cell.2019.05.006 (2019).
Romanov, R. A. et al. Molecular interrogation of hypothalamic organization reveals distinct dopamine neuronal subtypes. Nat Neurosci 20, 176–188, https://doi.org/10.1038/nn.4462 (2017).
Alpar, A. et al. Hypothalamic CNTF volume transmission shapes cortical noradrenergic excitability upon acute stress. EMBO J 37, https://doi.org/10.15252/embj.2018100087 (2018).
Korchynska, S. et al. A hypothalamic dopamine locus for psychostimulant-induced hyperlocomotion in mice. Nat Commun 13, 5944, https://doi.org/10.1038/s41467-022-33584-3 (2022).
Mickelsen, L. E. et al. Cellular taxonomy and spatial organization of the murine ventral posterior hypothalamus. Elife 9, https://doi.org/10.7554/eLife.58901 (2020).
Kim, D. W. et al. Multimodal Analysis of Cell Types in a Hypothalamic Node Controlling Social Behavior. Cell 179, 713–728 e717, https://doi.org/10.1016/j.cell.2019.09.020 (2019).
Saucisse, N. et al. Functional heterogeneity of POMC neurons relies on mTORC1 signaling. Cell Rep 37, 109800, https://doi.org/10.1016/j.celrep.2021.109800 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 e3529, https://doi.org/10.1016/j.cell.2021.04.048 (2021).
Hernandez, V. S. et al. Hypothalamic Vasopressinergic Projections Innervate Central Amygdala GABAergic Neurons: Implications for Anxiety and Stress Coping. Front Neural Circuits 10, 92, https://doi.org/10.3389/fncir.2016.00092 (2016).
Raj, B. et al. Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat Biotechnol 36, 442–450, https://doi.org/10.1038/nbt.4103 (2018).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142, https://doi.org/10.1126/science.aaa1934 (2015).
Hao, Z. Z. et al. Single-cell transcriptomics of adult macaque hippocampus reveals neural precursor cell populations. Nat Neurosci 25, 805–817, https://doi.org/10.1038/s41593-022-01073-x (2022).
Morel, L. et al. Intracortical astrocyte subpopulations defined by astrocyte reporter Mice in the adult brain. Glia 67, 171–181, https://doi.org/10.1002/glia.23545 (2019).
Batiuk, M. Y. et al. Identification of region-specific astrocyte subtypes at single cell resolution. Nat Commun 11, 1220, https://doi.org/10.1038/s41467-019-14198-8 (2020).
Sadick, J. S. et al. Astrocytes and oligodendrocytes undergo subtype-specific transcriptional changes in Alzheimer’s disease. Neuron 110, 1788–1805 e1710, https://doi.org/10.1016/j.neuron.2022.03.008 (2022).
Jackson, A. C. A common framework for mouse hypothalamic cell atlases. Nat Metab 4, 1227–1228, https://doi.org/10.1038/s42255-022-00660-3 (2022).
Furube, E. et al. Neural stem cell phenotype of tanycyte-like ependymal cells in the circumventricular organs and central canal of adult mouse brain. Sci Rep 10, 2826, https://doi.org/10.1038/s41598-020-59629-5 (2020).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol 32, 381–386, https://doi.org/10.1038/nbt.2859 (2014).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat Methods 14, 979–982, https://doi.org/10.1038/nmeth.4402 (2017).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502, https://doi.org/10.1038/s41586-019-0969-x (2019).
Brozzi, F., Arcuri, C., Giambanco, I. & Donato, R. S100B Protein Regulates Astrocyte Shape and Migration via Interaction with Src Kinase. J Biol Chem 284, 8797–8811, https://doi.org/10.1074/jbc.M805897200 (2009).
Acknowledgements
S.B.L is supported by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (HR22C1734) and the National Research Foundation (NRF) of Korea (2020R1A6A1A03043539, 2020M3A9D8037604, 2022R1C1C1004756). E.J.L is supported by the NRF of Korea (2022R1C1C1005741 and RS-2023-00217595) and the new faculty research fund of Ajou University School of Medicine.
Author information
Authors and Affiliations
Contributions
S.B.L. and E.J.L. conceptualized and designed the study. E.J.L. and H.K.C. generated the validation dataset. K.K. provided PRV. M.J., S.B.L. and E.J.L. analyzed and interpreted the data. All authors reviewed and contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Junaid, M., Choe, H.K., Kondoh, K. et al. Unveiling Hypothalamic Molecular Signatures via Retrograde Viral Tracing and Single-Cell Transcriptomics. Sci Data 10, 861 (2023). https://doi.org/10.1038/s41597-023-02789-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02789-6