Abstract
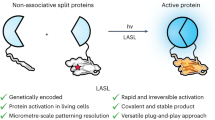
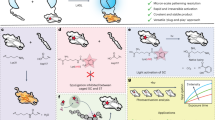
Proteases function as pivotal molecular switches, initiating numerous biological events. Notably, potyviral protease, derived from plant viruses, has emerged as a trusted proteolytic switch in synthetic biological circuits. To harness their capabilities, we have developed a single-component photocleavable switch, termed LAUNCHER (Light-Assisted UNcaging switCH for Endoproteolytic Release), by employing a circularly permutated tobacco etch virus protease and a blue-light-gated substrate, which are connected by fine-tuned intermodular linkers. As a single-component system, LAUNCHER exhibits a superior signal-to-noise ratio compared with multi-component systems, enabling precise and user-controllable release of payloads. This characteristic renders LAUNCHER highly suitable for diverse cellular applications, including transgene expression, tailored subcellular translocation and optochemogenetics. Additionally, the plug-and-play integration of LAUNCHER into existing synthetic circuits facilitates the enhancement of circuit performance. The demonstrated efficacy of LAUNCHER in improving existing circuitry underscores its significant potential for expanding its utilization in various applications.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data in this paper are included in the manuscript or the Supplementary Information. Source data and information for plasmid DNA generated in this study are provided with this paper. Key plasmids and their sequences of LAUNCHER are available in Addgene (210501).
References
Lopez-Otin, C. & Bond, J. S. Proteases: multifunctional enzymes in life and disease. J. Biol. Chem. 283, 30433–30437 (2008).
Lichtenthaler, S. F., Lemberg, M. K. & Fluhrer, R. Proteolytic ectodomain shedding of membrane proteins in mammals—hardware, concepts, and recent developments. EMBO J. 37, e99456 (2018).
Solary, E., Eymin, B., Droin, N. & Haugg, M. Proteases, proteolysis, and apoptosis. Cell Biol. Toxicol. 14, 121–132 (1998).
Neurath, H. Proteolytic processing and physiological regulation. Trends Biochem. Sci. 14, 268–271 (1989).
Pahl, H. L. & Baeuerle, P. A. Control of gene expression by proteolysis. Curr. Opin. Cell Biol. 8, 340–347 (1996).
Chung, H. K. et al. A compact synthetic pathway rewires cancer signaling to therapeutic effector release. Science 364, eaat6982 (2019).
Daringer, N. M., Dudek, R. M., Schwarz, K. A. & Leonard, J. N. Modular Extracellular Sensor Architecture for engineering mammalian cell-based devices. ACS Synth. Biol. 3, 892–902 (2014).
Fink, T. et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells. Nat. Chem. Biol. 15, 115–122 (2019).
Chung, H. K. & Lin, M. Z. On the cutting edge: protease-based methods for sensing and controlling cell biology. Nat. Methods 17, 885–896 (2020).
Gray, D. C., Mahrus, S. & Wells, J. A. Activation of specific apoptotic caspases with an engineered small-molecule-activated protease. Cell 142, 637–646 (2010).
Rachel, B. K. et al. Tobacco etch virus protease mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. Des. Sel. 14, 993–1000 (2001).
Kapust, R. B., Tözsér, J., Copeland, T. D. & Waugh, D. S. The P1′ specificity of tobacco etch virus protease. Biochem. Biophys. Res. Commun. 294, 949–955 (2002).
Wehr, M. C. et al. Monitoring regulated protein-protein interactions using split TEV. Nat. Methods 3, 985–993 (2006).
Baeumler, T. A., Ahmed, A. A. & Fulga, T. A. Engineering synthetic signaling pathways with programmable dCas9-based chimeric receptors. Cell Rep. 20, 2639–2653 (2017).
Dolberg, T. B. et al. Computation-guided optimization of split protein systems. Nat. Chem. Biol. 17, 531–539 (2021).
Lee, D. et al. Temporally precise labeling and control of neuromodulatory circuits in the mammalian brain. Nat. Methods 14, 495–503 (2017).
Lee, D., Hyun, J. H., Jung, K., Hannan, P. & Kwon, H. B. A calcium- and light-gated switch to induce gene expression in activated neurons. Nat. Biotechnol. 35, 858–863 (2017).
Kim, M. W. et al. Time-gated detection of protein-protein interactions with transcriptional readout. eLife 6, e30233 (2017).
Ross, B., Mehta, S. & Zhang, J. Molecular tools for acute spatiotemporal manipulation of signal transduction. Curr. Opin. Chem. Biol. 34, 135–142 (2016).
Shekhawat, S. S. & Ghosh, I. Split-protein systems: beyond binary protein-protein interactions. Curr. Opin. Chem. Biol. 15, 789–797 (2011).
Voss, S., Klewer, L. & Wu, Y. W. Chemically induced dimerization: reversible and spatiotemporal control of protein function in cells. Curr. Opin. Chem. Biol. 28, 194–201 (2015).
Stanton, B. Z., Chory, E. J. & Crabtree, G. R. Chemically induced proximity in biology and medicine. Science 359, eaao5902 (2018).
Kramer, M. M., Lataster, L., Weber, W. & Radziwill, G. Optogenetic approaches for the spatiotemporal control of signal transduction pathways. Int. J. Mol. Sci. 22, 5300 (2021).
Sanchez, M. I., Nguyen, Q. A., Wang, W., Soltesz, I. & Ting, A. Y. Transcriptional readout of neuronal activity via an engineered Ca2+-activated protease. Proc. Natl Acad. Sci. USA 117, 33186–33196 (2020).
Xu, X. et al. A single-component optogenetic system allows stringent switch of gene expression in yeast cells. ACS Synth. Biol. 7, 2045–2053 (2018).
Kaberniuk, A. A., Baloban, M., Monakhov, M. V., Shcherbakova, D. M. & Verkhusha, V. V. Single-component near-infrared optogenetic systems for gene transcription regulation. Nat. Commun. 12, 3859 (2021).
Chung, H. K. et al. Tunable and reversible drug control of protein production via a self-excising degron. Nat. Chem. Biol. 11, 713–720 (2015).
Jacobs, C. L., Badiee, R. K. & Lin, M. Z. StaPLs: versatile genetically encoded modules for engineering drug-inducible proteins. Nat. Methods 15, 523–526 (2018).
Zhang, W. et al. Optogenetic control with a photocleavable protein, PhoCl. Nat. Methods 14, 391–394 (2017).
Lu, X. et al. Photocleavable proteins that undergo fast and efficient dissociation. Chem. Sci. 12, 9658–9672 (2021).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl Acad. Sci. USA 112, 112–117 (2015).
Berger, J., Hauber, J., Hauber, R., Geiger, R. & Cullen, B. R. Secreted placental alkaline phosphatase: a powerful new quantitative indicator of gene expression in eukaryotic cells. Gene 66, 1–10 (1988).
Seo, J. K., Choi, H. S. & Kim, K. H. Engineering of soybean mosaic virus as a versatile tool for studying protein-protein interactions in soybean. Sci. Rep. 6, 22436 (2016).
Nallamsetty, S. et al. Efficient site-specific processing of fusion proteins by tobacco vein mottling virus protease in vivo and in vitro. Protein Expr. Purif. 38, 108–115 (2004).
Cella, F., Wroblewska, L., Weiss, R. & Siciliano, V. Engineering protein-protein devices for multilayered regulation of mRNA translation using orthogonal proteases in mammalian cells. Nat. Commun. 9, 4392 (2018).
Fernandez-Rodriguez, J. & Voigt, C. A. Post-translational control of genetic circuits using Potyvirus proteases. Nucleic Acids Res. 44, 6493–6502 (2016).
Gao, X. J., Chong, L. S., Kim, M. S. & Elowitz, M. B. Programmable protein circuits in living cells. Science 361, 1252–1258 (2018).
Icha, J., Weber, M., Waters, J. C. & Norden, C. Phototoxicity in live fluorescence microscopy, and how to avoid it. Bioessays 39, 1700003 (2017).
Cong, F., Schweizer, L. & Varmus, H. Wnt signals across the plasma membrane to activate the β-catenin pathway by forming oligomers containing its receptors, Frizzled and LRP. Development 131, 5103–5115 (2004).
Metcalfe, C., Mendoza-Topaz, C., Mieszczanek, J. & Bienz, M. Stability elements in the LRP6 cytoplasmic tail confer efficient signalling upon DIX-dependent polymerization. J. Cell Sci. 123, 1588–1599 (2010).
Lee, S. et al. A rationally designed optochemogenetic switch for activating canonical Wnt signaling. iScience 26, 106233 (2023).
Shahi, P. et al. Activation of Wnt signaling by chemically induced dimerization of LRP5 disrupts cellular homeostasis. PLoS ONE 7, e30814 (2012).
Zha, J. P., Weiler, S., Oh, K. J., Wei, M. C. & Korsmeyer, S. J. Posttranslational N-myristoylation of BID as a molecular switch for targeting mitochondria and apoptosis. Science 290, 1761–1765 (2000).
Lee, D. & Kwon, H. B. Current and future techniques for detecting oxytocin: focusing on genetically-encoded GPCR sensors. J. Neurosci. Methods 366, 109407 (2022).
Mignocchi, N., Kruessel, S., Jung, K., Lee, D. & Kwon, H. B. Development of a genetically-encoded oxytocin sensor. Preprint at bioRxiv, https://doi.org/10.1101/2020.07.14.202598 (2020).
Kanaji, S., Iwahashi, J., Kida, Y., Sakaguchi, M. & Mihara, K. Characterization of the signal that directs Tom20 to the mitochondrial outer membrane. J. Cell Biol. 151, 277–288 (2000).
Gossen, M. & Bujard, H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl Acad. Sci. USA 89, 5547–5551 (1992).
Rivera, V. M. et al. A humanized system for pharmacologic control of gene expression. Nat. Med. 2, 1028–1032 (1996).
Hyun, J. H. et al. Tagging active neurons by soma-targeted Cal-Light. Nat. Commun. 13, 7692 (2022).
Lu, X., Shen, Y. & Campbell, R. E. Engineering photosensory modules of non-opsin-based optogenetic actuators. Int. J. Mol. Sci. 21, 6522 (2020).
Klewer, L. & Wu, Y. W. Light-induced dimerization approaches to control cellular processes. Chemistry 25, 12452–12463 (2019).
Zhang, K. & Cui, B. Optogenetic control of intracellular signaling pathways. Trends Biotechnol. 33, 92–100 (2015).
Krueger, D. et al. Principles and applications of optogenetics in developmental biology. Development 146, dev175067 (2019).
Rogers, K. W. & Muller, P. Optogenetic approaches to investigate spatiotemporal signaling during development. Curr. Top. Dev. Biol. 137, 37–77 (2020).
H, M. et al. Photo-induced peptide cleavage in the green-to-red conversion of a fluorescent protein. Mol. Cell. 12, 1051–1058 (2003).
McEvoy, A. L. et al. mMaple: a photoconvertible fluorescent protein for use in multiple imaging modalities. PLoS ONE 7, e51314 (2012).
Miyamae, Y., Chen, L. C., Utsugi, Y., Farrants, H. & Wandless, T. J. A method for conditional regulation of protein stability in native or near-native form. Cell Chem. Biol. 27, 1573–1581 e3 (2020).
Goh, C. J. & Hahn, Y. Analysis of proteolytic processing sites in potyvirus polyproteins revealed differential amino acid preferences of NIa-Pro protease in each of seven cleavage sites. PLoS ONE 16, e0245853 (2021).
Chen, S. et al. Identification of highly selective covalent inhibitors by phage display. Nat. Biotechnol. 39, 490–498 (2021).
Acknowledgements
This study was supported by grants from the National Research Foundation of Korea, the Korea Government Ministry of Science and Information and Communications Technology (grant nos. NRF‐2018M3C7A1024597 and NRF-2020R1C1C1014793 to D.L.), the Korea University Grant (grant no. K2320381 to D.L.) and the National Institutes of Health (grant no. DP1MH119428 to H.-B.K.). We thank all members of the laboratory for their critical discussions and comments. We also thank Yunho Lee and Yuni Lee for the encouragement.
Author information
Authors and Affiliations
Contributions
M.C., H.-B.K. and Dongmin Lee conceptualized the research. The design of the plasmid vector was spearheaded by M.C., S.L., S.H.B. and Dongmin Lee. The SEAP assay in HEK293T cells was executed and the associated data collected by M.C., S.L., S.H.B., M.S., J.Y.K., S.K.C. and J.H. J.R.R. was responsible for the isolation of hippocampal neurons from rat embryos. M.C. captured all fluorescence images, with quantitative fluorescence analysis undertaken by M.C., M.S. and S.H.Y. Comprehensive data analysis was carried out by M.C., S.L., S.H.B., S.H.Y., Y.K., K.H., Donghun Lee, W.S., H.-B.K. and Dongmin Lee. All figures and schematics were crafted by M.C. and Dongmin Lee. The manuscript’s initial drafts were penned by M.C., S.L., S.H.B., J.R.R., H.-B.K. and Dongmin Lee, under the overarching supervision of Dongmin Lee. All authors engaged in discussions and provided critical feedback on the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The Korea University has filed a patent application that includes portions of the research described in this manuscript.
Peer review
Peer review information
Nature Chemical Biology thanks Carl Denard and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Screening and validation of size-tunable light-controlled linkers.
a, Structural map illustrating the size-tunable light-controlled linker. The intermodular locations of the first linker (1st LK) and the second linker (2nd LK) are depicted within the structural map. b, Summary graph displaying SEAP expressions of screened variants with different combinations of intermodular linkers. c, Heatmap of fold changes in SEAP expression levels for variable linker-size variants. d, graph shows the SEAP expression level of four constructs selected in a previous screening (c), with varying amounts of transfected DNA. e, Heatmap of fold changes in SEAP expression levels of four constructs with different amount of transfection. c, e Fold-change (blue-light vs. dark) values of individual candidates are indicated by their colors on a three-color gradient scale. Data (b and d) were collected from six biologically independent samples and data points represent individual measurements. In box-and-whisker plots, the upper and lower whiskers indicate the maximum and minimum, respectively, whereas the boxes bracket the 25th, 50th, and 75th percentiles. The significance of differences between dark vs. blue-light conditions was tested using an unpaired t-test with Welch correction. *p < 0.05, **p < 0.005, ***p < 0.001, ****p < 0.0001. Error bars, SD.
Extended Data Fig. 2 Screening of flexible linker with variable length and tandem NESs.
a, Single-component photocleavable variants with flexible linkers of different lengths following TevN. b, Light-dependent SEAP expression and fold-changes according to intermodular linker size. c, SEAP activities of all candidates in the third round screening with combinations of variable length and tandem NESs. Data (b, c) were collected from at least four biologically independent samples, and the data points represent individual measurements. In box-and-whisker plots, the upper and lower whiskers indicate the maximum and minimum, respectively, whereas the boxes bracket the 25th, 50th, and 75th percentiles. The significance of differences between dark vs. blue-light conditions was tested using an unpaired t-test with Welch correction. *p < 0.05, **p < 0.005, ***p < 0.001, ****p < 0.0001. Error bars, SD.
Extended Data Fig. 3 Screening of TEVp substrate with P1′ substitution.
a, Schematics presents a single-component photocleavable variant with P1′ substitution (denoted X). b, Light-dependent SEAP expression levels of single-component photocleavable variants with different 20 aa in the P1′ site of the TEV cleavage sequence. Data (b) were collected from at least four biologically independent samples, and the data points represent individual measurements. In box-and-whisker plots, the upper and lower whiskers indicate the maximum and minimum, respectively, whereas the boxes bracket the 25th, 50th, and 75th percentiles. The significance of differences between dark vs. blue-light conditions was tested using an unpaired t-test with Welch correction. *p < 0.05, **p < 0.005. Error bars, SD.
Extended Data Fig. 4 Spectral characterization of LAUNCHER.
a, Summary graph showing the SEAP expression levels of LAUNCHER with increasing light intensity (0%–100%). b, Spectral specificity of the LAUNCHER system. HEK293T cells were irradiated with light of three different wavelengths (blue: 470 nm, green: 520 nm, and red: 630 nm). c, Schematic diagram of the lighting duty cycle with various frequencies (4/60 Hz, 2/60 Hz, and 1/60 Hz). d, Summary graph of SEAP expression levels according to duty cycles. Data (a, b, d) were collected from at least four biologically independent samples and data points represent individual measurements. In box-and-whisker plots, the upper and lower whiskers indicate the maximum and minimum, respectively, whereas boxes bracket the 25th, 50th, and 75th percentiles. One-way ANOVA with Dunnett’s multiple comparisons test was used to assess the significance of differences between the dark group and other groups in (a, d) and between the blue-light group and other groups in (b). ****p < 0.0001. Error bars, SD.
Extended Data Fig. 5 Photomask-guided spatial expression of LAUNCHER.
a, Configuration of LAUNCHER construct including TRE-tdTomato as a reporter and EGFP as a transfection marker to visualize the spatial resolution of LAUNCHER. b, The cartoon illustrates the sequential procedures for achieving photomask-guided spatial expression of LAUNCHER. A three mm width laser-cutting acryl is used as a photomask to define the spatial expression of tdTomato. Light-induced tdTomato expression was acquired by a fluorescent imaging scanner. c, Scanned images of EGFP (left), tdTomato (middle), and merge image (right) of LAUNCHER. Scale bar, 1 cm.
Extended Data Fig. 6 Light-dependent fluorescent reporter expression of LAUNCHER orthologs.
a, Schematic representations of LAUNCHERTEVp (top), LAUNCHERSbMVp (middle), and LAUNCHERTVMVp (bottom) with corresponding three cleavage sequences (TEVp: ENLYFQG, SbMVp: ESVSLQG, TVMVp: ETVRFQG). b, Representative confocal images of three LAUNCHER orthologs with their matched cleavage sequences in the condition of dark and blue. EGFP (green colored) as transfection marker, TRE-tdTomato (red colored) as reporter, and LAUNCHER were transfected into HEK293T cells. Scale bar, 50 μm. c, Quantitative measurement of LAUNCHER orthologs in the condition of dark and blue. An automated cell counting device (Invitrogen Countess 3) was used to count tdTomato-positive cells within the population of EGFP-positive cells. The results of each measurement are indicated as percentage. Data (c) were collected from at least four biologically independent samples. In the box plots, the upper and lower whiskers indicate the maximum and minimum, respectively, and the boxes bracket the 25th, 50th, and 75th percentiles. The significance of differences between dark vs. blue-light illumination was tested using an unpaired t-test with Welch correction. ****p < 0.0001. Error bars, SD.
Extended Data Fig. 7 Impact of LAUNCHERized iTango2 activity by proteolytic cleavage-dependent βArr2-TevC release.
a, A schematic illustration of LAUNCHERized DRD2-iTango2 (DRD2-TevN-iLID-tTA and NTOM20-LAUNCHERTEVp-βArr2-TevC) with photocleavable (glycine, left) or non-photocleavable (proline, right) TEVseq within LAUNCHER. NonDRD2-iTango2 insert active SbMVseq (left) or inactive SbMVseq (right) into LAUNCHERTEVp-βArr2-TevC. b, SEAP assay of LAUNCHERized iTango2 activity depending on proteolytic release of βArr2-TevC. Photocleavable TEVseq with glycine (left panel) and non-photocleavable TEVseq with proline (right panel) were presented in the summary graph. Data (b) were collected from four biologically independent samples. In the box-and-whisker plots, the upper and lower whiskers indicate the maximum and minimum, respectively, and the boxes bracket the 25th, 50th, and 75th percentiles. One-way ANOVA with Dunnett’s multiple comparisons test was used to assess the significance of differences between the blue-light/quinpirole group and other groups. ****p < 0.0001. Error bars, SD.
Extended Data Fig. 8 Light- and ligand-dependent gene expression of LAUNCHER-DRD2-iTango2.
a, Configuration of the LAUNCHERSbMVp-DRD2-iTango2 construct, including TRE-tdTomato as a fluorescent reporter. b, Confocal images of LAUNCHERSbMVp-DRD2-iTango2. Light- and ligand-dependent gene expression patterns in HEK293T cells. Green signals of EGFP (transfection marker) indicate mitochondrial localization, red signals of tdTomato indicate LAUNCHER activity, and blue-signals of DAPI indicate nuclear localization. Scale bar, 50 μm. c, Quantitative analysis of LAUNCHERized iTango2 in HEK293T cells. Percentages of tdTomato-positive cells within the EGFP-positive cell population are measured using a fluorescence-based cell counter. Data (c) were collected from four biologically independent samples. In box-and-whisker plots, the upper and lower whiskers indicate the maximum and minimum respectively, the boxes bracket the 25th, 50th, and 75th percentiles. One-way ANOVA with Dunnett’s multiple comparisons test was used to assess the significance of differences between the blue-light/quinpirole group and other groups. ****p < 0.0001.
Extended Data Fig. 9 Optochemogenetic Tandem AND-Gate Neural Activity Recorder Using LAUNCHER in Neurons.
a, Tandem AND-logic gate for the neural activity readout system. The three modulators are blue-light, neural activity, and rapamycin. b, Schematics of the LAUNCHER-based neural activity readout system and the three constructs used in the configuration are shown. (1) N-terminal Myc tag and KA2 motif fused to LAUNCHERTEVP-TetR-FKBP. (2) FRB-VP16-P2A-EGFP expressed by c-Fos promoter, with EGFP as a neural activity marker. (3) TRE-tdTomato as a system output reporter. Blue-light or neural activity induces release of TetR-FKBP or expression of FRB-VP16, respectively. Rapamycin acts as an output switch to turn on reporter expression (tdTomato) by coupling of TetR and VP16. c, Confocal images of primary cultured rat hippocampal neurons. Myc tag immunostaining (Cy5) marks transfection while c-Fos-based EGFP indicates activity. Light- and rapamycin- gated neural activity can be monitored by the expression of tdTomato (Red). Neural activity is controlled by the treatment of 2 μM TTX or 30 μM bicuculline (BIC) Scale bars 100 μm. d, Population percentages of tdTomato positive neurons over Cy5 positive neurons in each group. Data (d) were collected from 10 biologically independent samples. In box plots, the upper and lower whiskers indicate the maximum and minimum, respectively, and boxes bracket the 25th, 50th, and 75th percentiles. One-way ANOVA with Dunnett’s multiple comparisons test was used to assess the significance of differences between the blue-light/BIC/rapamycin group and others. ****p < 0.0001. Error bars, SD.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9 and Table 1 (Sequence of plasmid DNA used in this study).
Supplementary Video 1
LANUNCHER and PhoCl2 live video.
Source data
Source Data Figs. 1, 2, 3, 4, 5, 6 and Extended Data Figs. 1, 2, 3, 4, 6, 7, 8, 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cui, M., Lee, S., Ban, S.H. et al. A single-component, light-assisted uncaging switch for endoproteolytic release. Nat Chem Biol 20, 353–364 (2024). https://doi.org/10.1038/s41589-023-01480-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01480-6