Abstract
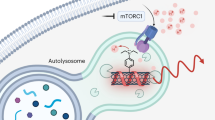
Autophagy—the lysosomal degradation of cytoplasmic components via their sequestration into double-membraned autophagosomes—has not been detected non-invasively. Here we show that the flux of autophagosomes can be measured via magnetic resonance imaging or serial near-infrared fluorescence imaging of intravenously injected iron oxide nanoparticles decorated with cathepsin-cleavable arginine-rich peptides functionalized with the near-infrared fluorochrome Cy5.5 (the peptides facilitate the uptake of the nanoparticles by early autophagosomes, and are then cleaved by cathepsins in lysosomes). In the heart tissue of live mice, the nanoparticles enabled quantitative measurements of changes in autophagic flux, upregulated genetically, by ischaemia–reperfusion injury or via starvation, or inhibited via the administration of a chemotherapeutic or the antibiotic bafilomycin. In mice receiving doxorubicin, pre-starvation improved cardiac function and overall survival, suggesting that bursts of increased autophagic flux may have cardioprotective effects during chemotherapy. Autophagy-detecting nanoparticle probes may facilitate the further understanding of the roles of autophagy in disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The raw and analyzed datasets generated during the study are too large to be publicly shared but are available for research purposes from the corresponding authors on reasonable request. Source data are provided with this paper.
References
Levine, B. & Klionsky, D. J. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev. Cell 6, 463–477 (2004).
Levine, B. Eating oneself and uninvited guests: autophagy-related pathways in cellular defense. Cell 120, 159–162 (2005).
Choi, A. M., Ryter, S. W. & Levine, B. Autophagy in human health and disease. N. Engl. J. Med. 368, 651–662 (2013).
Nishino, I. et al. Primary LAMP-2 deficiency causes X-linked vacuolar cardiomyopathy and myopathy (Danon disease). Nature 406, 906–910 (2000).
Mizushima, N. & Levine, B. Autophagy in human diseases. N. Engl. J. Med. 383, 1564–1576 (2020).
Nah, J., Fernandez, A. F., Kitsis, R. N., Levine, B. & Sadoshima, J. Does autophagy mediate cardiac myocyte death during stress? Circ. Res. 119, 893–895 (2016).
Doherty, J. & Baehrecke, E. H. Life, death and autophagy. Nat. Cell Biol. 20, 1110–1117 (2018).
Savini, M. & Wang, M. C. Does autophagy promote longevity? It depends. Cell 177, 221–222 (2019).
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000).
Kimura, S., Noda, T. & Yoshimori, T. Dissection of the autophagosome maturation process by a novel reporter protein, tandem fluorescent-tagged LC3. Autophagy 3, 452–460 (2007).
Gottlieb, R. A., Andres, A. M., Sin, J. & Taylor, D. P. Untangling autophagy measurements: all fluxed up. Circ. Res. 116, 504–514 (2015).
Kobayashi, S. et al. Artificial induction of autophagy around polystyrene beads in nonphagocytic cells. Autophagy 6, 36–45 (2010).
Remaut, K., Oorschot, V., Braeckmans, K., Klumperman, J. & De Smedt, S. C. Lysosomal capturing of cytoplasmic injected nanoparticles by autophagy: an additional barrier to non viral gene delivery. J. Control. Release 195, 29–36 (2014).
Lewin, M. et al. Tat peptide-derivatized magnetic nanoparticles allow in vivo tracking and recovery of progenitor cells. Nat. Biotechnol. 18, 410–414 (2000).
Schmidt, N., Mishra, A., Lai, G. H. & Wong, G. C. Arginine-rich cell-penetrating peptides. FEBS Lett. 584, 1806–1813 (2010).
Nakase, I. et al. Arginine-rich cell-penetrating peptide-modified extracellular vesicles for active macropinocytosis induction and efficient intracellular delivery. Sci. Rep. 7, 1991 (2017).
Dowaidar, M. et al. Role of autophagy in cell-penetrating peptide transfection model. Sci. Rep. 7, 12635 (2017).
Robison, A. D. et al. Polyarginine interacts more strongly and cooperatively than polylysine with phospholipid bilayers. J. Phys. Chem. B 120, 9287–9296 (2016).
Najjar, K. et al. Unlocking endosomal entrapment with supercharged arginine-rich peptides. Bioconjug. Chem. 28, 2932–2941 (2017).
Appelbaum, J. S. et al. Arginine topology controls escape of minimally cationic proteins from early endosomes to the cytoplasm. Chem. Biol. 19, 819–830 (2012).
Qian, Z. et al. Early endosomal escape of a cyclic cell-penetrating peptide allows effective cytosolic cargo delivery. Biochemistry 53, 4034–4046 (2014).
Uchiyama, Y. Autophagic cell death and its execution by lysosomal cathepsins. Arch. Histol. Cytol. 64, 233–246 (2001).
Chen, H. H. et al. Fluorescence tomography of rapamycin-induced autophagy and cardioprotection in vivo. Circ. Cardiovasc. Imaging 6, 441–447 (2013).
Wu, P. et al. Myocardial upregulation of cathepsin D by ischemic heart disease promotes autophagic flux and protects against cardiac remodeling and heart failure. Circ. Heart Fail. 10, e004044 (2017).
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004).
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004).
Zhou, J. et al. Activation of lysosomal function in the course of autophagy via mTORC1 suppression and autophagosome–lysosome fusion. Cell Res. 23, 508–523 (2013).
McConnell, H. L. et al. Ferumoxytol nanoparticle uptake in brain during acute neuroinflammation is cell-specific. Nanomedicine 12, 1535–1542 (2016).
Morishita, H., Kaizuka, T., Hama, Y. & Mizushima, N. A new probe to measure autophagic flux in vitro and in vivo. Autophagy 13, 757–758 (2017).
Nahrendorf, M. et al. The healing myocardium sequentially mobilizes two monocyte subsets with divergent and complementary functions. J. Exp. Med. 204, 3037–3047 (2007).
Simonson, B. et al. DDiT4L promotes autophagy and inhibits pathological cardiac hypertrophy in response to stress. Sci. Signal. 10, eaaf5967 (2017).
Dhingra, R. et al. Bnip3 mediates doxorubicin-induced cardiac myocyte necrosis and mortality through changes in mitochondrial signaling. Proc. Natl Acad. Sci. USA 111, E5537–E5544 (2014).
Li, D. L. et al. Doxorubicin blocks cardiomyocyte autophagic flux by inhibiting lysosome acidification. Circulation 133, 1668–1687 (2016).
Abdullah, C. S. et al. Doxorubicin-induced cardiomyopathy associated with inhibition of autophagic degradation process and defects in mitochondrial respiration. Sci. Rep. 9, 2002 (2019).
Whitehead, N. P., Kim, M. J., Bible, K. L., Adams, M. E. & Froehner, S. C. A new therapeutic effect of simvastatin revealed by functional improvement in muscular dystrophy. Proc. Natl Acad. Sci. USA 112, 12864–12869 (2015).
Liu, Y. et al. Visnagin protects against doxorubicin-induced cardiomyopathy through modulation of mitochondrial malate dehydrogenase. Sci. Transl. Med. 6, 266ra170 (2014).
Asnani, A. et al. Highly potent visnagin derivatives inhibit Cyp1 and prevent doxorubicin cardiotoxicity. JCI Insight 3, e96753 (2018).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (4th edition). Autophagy 17, 1–382 (2021).
Perez, J. M., Josephson, L., O’Loughlin, T., Hogemann, D. & Weissleder, R. Magnetic relaxation switches capable of sensing molecular interactions. Nat. Biotechnol. 20, 816–820 (2002).
Taktak, S., Sosnovik, D., Cima, M. J., Weissleder, R. & Josephson, L. Multiparameter magnetic relaxation switch assays. Anal. Chem. 79, 8863–8869 (2007).
Nahrendorf, M. et al. Dual channel optical tomographic imaging of leukocyte recruitment and protease activity in the healing myocardial infarct. Circ. Res. 100, 1218–1225 (2007).
Chen, H. H. et al. Theranostic nucleic acid binding nanoprobe exerts anti-inflammatory and cytoprotective effects in ischemic injury. Theranostics 7, 814–825 (2017).
Yuan, H. et al. Heat-induced-radiolabeling and click chemistry: a powerful combination for generating multifunctional nanomaterials. PLoS ONE 12, e0172722 (2017).
Messroghli, D. R. et al. Clinical recommendations for cardiovascular magnetic resonance mapping of T1, T2, T2* and extracellular volume: a consensus statement by the Society for Cardiovascular Magnetic Resonance (SCMR) endorsed by the European Association for Cardiovascular Imaging (EACVI). J. Cardiovasc. Magn. Reson. 19, 75 (2017).
Acknowledgements
This study was supported in part by the following grants from the National Institutes of Health: K99/R00HL121152 (to H.H.C.), R01HL112831 and R01HL141563 (to D.E.S.), R01HL122547 and R01HL102368-06A1 (to S.D.), R01HL131635 (to C.M.) and R01HL131831 (to R.M.B.). Electron microscopy was performed in the Microscopy Core of the Center for Systems Biology/Program in Membrane Biology, which is partially supported by an Inflammatory Bowel Disease Grant (DK043351) and a Boston Area Diabetes and Endocrinology Research Center (BADERC) award (DK057521). The funders had no role in the design of the study, in data collection and analysis, in the decision to publish the manuscript or in its preparation.
Author information
Authors and Affiliations
Contributions
H.H.C. and D.E.S. conceived the study, designed the experiments and performed data acquisition, data analysis, data interpretation, figure preparation and manuscript writing. Z.K. and L.W. contributed to data acquisition and data analysis. C.M. and S.J.W.K. contributed to data analysis and figure preparation. D.P., A.B., E.E., L.M., Y.I.C., L.K., G.L., D.E.C., P.S. and A.T.N.K. contributed to data acquisition and data analysis. R.M.B. contributed to data acquisition, data analysis and data interpretation. H.Y. contributed to experimental design, data acquisition and data analysis. S.D. contributed to experimental design, data analysis and data interpretation. L.J. contributed to the conception of the study and to experimental design and data interpretation. All authors contributed to the editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Richard Kitsis, Ben Loos and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary information
Supplementary results, methods and figures.
Source data
Source data for Fig. 4
Uncropped gels for a.
Source data for Fig. 6
Uncropped gels for n.
Rights and permissions
About this article
Cite this article
Chen, H.H., Khatun, Z., Wei, L. et al. A nanoparticle probe for the imaging of autophagic flux in live mice via magnetic resonance and near-infrared fluorescence. Nat. Biomed. Eng 6, 1045–1056 (2022). https://doi.org/10.1038/s41551-022-00904-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-022-00904-3
This article is cited by
-
Ultra-photostable small-molecule dyes facilitate near-infrared biophotonics
Nature Communications (2024)
-
Recent advances in molecular and nanoparticle probes for fluorescent bioanalysis
Nano Research (2024)
-
Non-invasive monitoring of autophagy
Nature Biomedical Engineering (2022)