Abstract
The epigenetic mechanisms coordinating the maintenance of adult cellular lineages and the inhibition of alternative cell fates remain poorly understood. Here we show that targeted ablation of the histone chaperone HIRA in myogenic cells leads to extensive transcriptional modifications, consistent with a role in maintaining skeletal muscle cellular identity. We demonstrate that conditional ablation of HIRA in muscle stem cells of adult mice compromises their capacity to regenerate and self-renew, leading to tissue repair failure. Chromatin analysis of Hira-deficient cells show a significant reduction of histone variant H3.3 deposition and H3K27ac modification at regulatory regions of muscle genes. Additionally, we find that genes from alternative lineages are ectopically expressed in Hira-mutant cells via MLL1/MLL2-mediated increase of H3K4me3 mark at silent promoter regions. Therefore, we conclude that HIRA sustains the chromatin landscape governing muscle cell lineage identity via incorporation of H3.3 at muscle gene regulatory regions, while preventing the expression of alternative lineage genes.
Similar content being viewed by others
Introduction
Skeletal muscle has a high regeneration potential that relies on tissue-resident muscle stem cells, also known as satellite cells. These adult stem cells were identified nearly 60 years ago as quiescent cells lying on the surface of the myofiber underneath the basal lamina1. Muscle stem cells express the paired-box transcription factor PAX7 which is required for their maintenance and myogenic lineage progression2. In resting muscles, quiescent satellite cells become activated upon injury to repair damaged tissue, while a fraction self-renews to regain quiescence. Activated satellite cells express MYF5 and MYOD and re-enter the cell cycle, proliferating as myoblasts that will either fuse to generate newly formed fibers or return to quiescence to restore the muscle stem cell pool3. Proper regulation of quiescence, activation and differentiation of muscle stem cells ensures the normal homeostasis of the tissue as well as its repair in the case of disease or injury-related damage. Although the transcriptional regulation of these processes is widely studied, the molecular mechanisms regulating PAX7 expression and maintaining muscle stem cell identity have not been identified.
Epigenetic regulation of gene expression is crucial for cell fate commitment and maintenance. The epigenetic memory of a specified cell defines the state of gene expression that is transmitted upon division in order to sustain cell lineage identity4. This mechanism of inheritance of a gene expression pattern relies on the epigenetic signature associated with DNA methylation, chromatin architecture, histone variants deposition, and histone modifications4,5. Nucleosome incorporation of the non-canonical histone H3 variant H3.3 has been associated with epigenetic memory and is required to maintain the transcriptionally active pattern of a tissue6. The H3.3 histone is specifically incorporated into the nucleosomes by two histone chaperones, HIRA and DAXX, depending on the genomic loci7,8. In mouse embryonic stem cells (ESCs), HIRA regulates H3.3 deposition at transcription start sites (TSS), promoters, and gene bodies while DAXX deposits H3.3 at the telomeres and constitutive heterochromatin7,8. H3.3 deposition is not only required for pluripotency maintenance but also for somatic cell gene expression regulation, including the neurogenic and myogenic lineages, playing a central role in cellular differentiation9,10,11,12. While it has been previously shown that HIRA and H3.3 are regulating myogenic cell differentiation9,10,11 their role in regulating PAX7 and sustaining lineage identity has not been addressed.
Here, we show that HIRA is a major upstream regulator of muscle stem cells and myogenic lineage identity. Conditional knockout of HIRA in muscle stem cells leads to defective skeletal muscle regeneration in vivo. We further demonstrate that HIRA-deficient cells lose PAX7 and muscle-specific gene expression, while inducing tissue-specific genes from other lineages. We show that HIRA drives myogenic identity by specifically incorporating H3.3 at regulatory regions of muscle genes. By contrast, induction of non-myogenic genes in HIRA-deficient muscle cells requires MLL1/MLL2-mediated deposition of H3K4me3 mark at silent or bivalent loci. Altogether, our data demonstrate that HIRA is required to maintain muscle stem cell lineage chromatin landscape in order to promote the expression of myogenic-specific genes and prevent the expression of alternative lineages.
Results
HIRA sustains myogenic cell identity
In order to identify upstream epigenetic regulators of the muscle stem cell-specific transcription factor PAX7, we performed targeted deletion of candidate genes in the myoblast C2C12 cell line. Among these, we addressed the role of HIRA as a potential upstream regulator of PAX7 by mutating Exon 10 of Hira using Crispr/Cas9 (Supplementary Fig. 1a–c), to generate a truncated protein lacking the putative H3.3 interaction domain13. The deletion of Hira did not affect the expression of Daxx/DAXX (Supplementary Fig. 1b, c). Upon Hira ablation, expression of PAX7 was drastically reduced and MYOD protein levels decreased (Fig. 1a–c). In addition, Hira mutant myogenic cells displayed impaired differentiation in low-serum-containing medium as seen with the quantification of Myosin-positive cells (Fig. 1d, e), consistent with previous observations11. Hira-deficient cells did not show changes in proliferation or cell death (Supplementary Fig. 1d–g). We analyzed two other independent Hira-KO clones for muscle phenotype by RT-qPCR and the loss of PAX7 and myogenic gene expression was consistent between them (Supplementary Fig. 1h). To characterize the global impact of the loss of Hira we performed RNA-sequencing (RNA-seq) in proliferating Hira-KO and control C2C12 cells cultured in high serum-containing medium and at low density (Fig. 1f, g). Strikingly, we observed major changes in the transcriptome of Hira KO compared to control cells with 30% of expressed genes being dysregulated (10297 out of 33686 detected genes), suggesting a global impact on gene expression in the absence of HIRA (Fig. 1h). This was unexpected given that Hira deletion in mouse ESCs does not result in major transcriptomic changes8,14. Gene ontology and analysis of downregulated genes, included genes linked with muscle stem cell lineage, including muscle stem cell markers, muscle lineage progression, and differentiation (Fig. 1i, k, l). Surprisingly, upregulated transcripts were related to non-myogenic lineages, such as vascular or neuronal lineages (Fig. 1j). Moreover, we found various endothelial, mesenchymal, neuronal, adipogenic, and osteogenic lineage-specific genes to be ectopically upregulated in myogenic cells when Hira was deleted (Fig. 1k, l). In addition, Hoxb cluster genes, previously identified as repressed in C2C12 cells15, showed a strong upregulation in the Hira-KO cell line (Fig. 1k, l). Taken together, these results show that HIRA is required to sustain muscle stem cells and myogenic gene expression and to inhibit the expression of genes from alternative lineages in C2C12 cells, making it an essential epigenetic factor required for the maintenance of myogenic cell identity at a transcriptomic level.
a Co-immunostaining with PAX7 (red) and MYOD (green) antibodies and the nuclear marker DAPI (blue) in C2C12 and Hira-KO cell lines. b Quantification of the number of PAX7-positive cells among DAPI-positive cells in a (n = 4 control and n = 4 cKO independent culture experiments). Error bars, mean ± SD, two-tailed unpaired t-test. c Western blot for MYOD and H3 (control) in C2C12 (n = 4 independent protein samples) and Hira KO (n = 4 independent protein samples). Uncropped blots in Source Data. Quantification of the signal intensity performed in four independent blots, comparing samples within each blot (bottom, arb. units: arbitrary units). Error bars, mean ± SD, two-tailed unpaired t-test. d Immunostaining with MF20 antibody (green) to visualize myosins and the nuclear marker DAPI (blue) in C2C12 and Hira-KO cell lines. e Quantification of the number of MF20-positive nuclei among DAPI-positive nuclei in d (C2C12 n = 6, Hira KO n = 5, independent differentiation assays). Error bars, mean ± SD, two-tailed unpaired t-test. f Differential expression analysis of the C2C12 (n = 3 independent RNA samples) and Hira KO (n = 3 independent RNA samples) RNA-seq samples. g MA-plot of C2C12 over Hira-KO RNA-seq data. Significantly dysregulated genes are highlighted in red (FDR < 0.05). h Number of upregulated (4920 genes, blue) and downregulated (5377 genes, red) genes in Hira KO compared to C2C12. i, j Gene ontology analysis for biological processes of the downregulated (i) and upregulated (j) genes. Selected enriched terms are presented according to the fold enrichment. k Heatmap with the normalized reads per gene of C2C12 and Hira-KO RNA-seq data in individual triplicates. l RT-qPCR analyses of the mRNA levels of selected genes from the C2C12 RNA-seq performed in control and Hira-KO cells (n = 6 for Fgfr4, Cdh5, Pdgfra, Efna5, Hoxb13; n = 5 Pax7, Myf5, Myh3, Hoxb5, Hoxb6, Hoxb7, Hoxb8, Hoxb9; n = 4 Myod1, Eya4, Angpt2; n = 3 Nefm, Nefl; independent RNA samples). For each gene, the mRNA levels of the control cells were normalized to 1. Muscle lineage (blue), alternative lineages (green), and Hoxb cluster (red). Error bars, mean ± SD, two-sided unpaired t-test. Scale bars, 40 μm.
H3.3 deposition by HIRA maintains H3K27ac mark in myogenic gene loci
We next evaluated whether the maintenance of skeletal muscle lineage identity is linked with the histone chaperone activity of HIRA and H3.3 incorporation. In order to determine if the deletion of Hira had an impact on the H3.3 incorporation in the genome, we performed H3.3 chromatin immunoprecipitation (ChIP) followed by sequencing (ChIP-seq) in proliferating Hira-deficient and control cells (cultured in high serum-containing medium and at low density). Genome-wide analysis showed a tenfold reduction in the H3.3 genomic deposition in the absence of Hira compared to control cells (38439 vs. 3744 detected peaks) (Fig. 2a). In addition, we observed that in myoblasts lacking Hira the number of H3.3 peaks is evenly decreased across genomic loci (Fig. 2b), suggesting a global pattern for H3.3 incorporation by HIRA in the genome of C2C12 cells, in contrast to findings in ESCs8,14,16. To investigate the H3.3 deposition pattern in muscle cells, we plotted the H3.3 average intensity signal along the promoter (±3 kb around the TSS), TSS and gene bodies in control and Hira-KO cells. We observed that H3.3 signal is diminished in Hira-KO cells when analyzing all protein-coding genes (Fig. 2c). To evaluate if decreased H3.3 deposition is linked with gene expression, we analyzed H3.3 average signal in upregulated or downregulated genes of the Hira-KO cell line compared with control cells. This analysis demonstrated that reduced H3.3 incorporation in promoters and TSS is specifically associated with genes that are downregulated in Hira KO but not with genes that are upregulated, the latter displaying H3.3 levels similar to control (Fig. 2c). In addition, we confirmed that skeletal muscle genes (GO:0007519) in Hira KO follow the same pattern of H3.3 levels as the downregulated genes (Fig. 2c). Given that incorporation of H3.3 is required for acetylation and promoter-distal enhancer (±3 kb around the TSS) activity in ESCs14 we further analyzed the distribution of the acetylated histone H3 Lys27 (H3K27ac) that is associated with active enhancers and promoters17, in Hira-deficient and control myoblasts using ChIP-seq. While H3K27ac modification was not altered in average when examined across all protein-coding genes, specific enrichment was observed in upregulated genes and conversely, downregulated genes showed decreased H3K27ac levels (Fig. 2d). This result suggests that loss of H3.3 incorporation in promoters and TSS of downregulated genes correlates with the loss of transcriptionally permissive H3K27ac mark (Fig. 2d). As actively expressed genes have been shown to reside within accessible chromatin, we performed an assay for transposase-accessible chromatin sequencing (ATAC-seq) to determine if the chromatin accessibility is modified in muscle genes when Hira is deleted in myogenic cells compared with controls. The chromatin accessibility was greatly decreased in Hira KO compared to control cells as shown by the reduced ATAC-seq peaks (Supplementary Fig. 2a, b). This decreased accessibility is likely due to the replacement of H3.3 by H3.1 upon loss of HIRA (Supplementary Fig. 2e, f). In addition, we observed that both H3.3 incorporation and H3K27ac histone modification were strongly reduced at sites where ATAC-seq peaks were lost in Hira-KO cells, demonstrating that H3.3 and H3K27ac modification are associated with open chromatin genomic regions (Fig. 2e, f). Since we detected a specific loss in the expression of myogenic-related genes in Hira-deficient cells, we further analyzed H3.3 enrichment at muscle-specific loci. The regulatory regions of Pax7 have not been characterized, but putative enhancer regions have been identified by H3K27ac ChIP-seq in quiescent satellite cells18. We first scanned Pax7 locus for H3.3 deposition and H3K27ac at opened chromatin sites in control cells. Strikingly, all detected regions presented a common pattern for H3.3, H3K27ac, and ATAC-seq peaks in myogenic cells and lost all of these marks in the absence of HIRA (Fig. 2g). We next analyzed the genomic regions of muscle stem cells and myogenic markers downregulated in Hira-deficient cells (Fig. 1k). These include the satellite cell markers Pax7 (Fig. 2g), Heyl (Fig. 2h), Col5a1 (Fig. 2i), Sdc3 (Supplementary Fig. 2c), the previously identified regulatory regions of Dmrt2 (−18 kb) (Fig. 2j), Myod1 (−20 kb) (Fig. 2k), and Fgfr4 (+19.2 kb) (Fig. 2l)19,20,21, as well as Eya4 (Fig. 2m) and Fgf2 (Supplementary Fig. 2d). We observed that all these muscle stem cell and muscle-specific regulatory regions presented H3.3 incorporation associated with H3K27ac histone mark at open chromatin sites in C2C12 myogenic cells (Fig. 2g–m, Supplementary Fig. 2c, d). Consistently, H3.3 incorporation, H3K27ac modification, and ATAC-seq peaks are abrogated in Hira-deleted cells (Fig. 2g–m, Supplementary Fig. 2c, d). Combining the analysis of the RNA-seq, ChIP-seq, and ATAC-seq in control and Hira-KO cells we conclude that the incorporation of H3.3 by HIRA is required for chromatin accessibility and H3K27 histone acetylation at regulatory regions of PAX7 and myogenic-specific genes in order to maintain their expression in the muscle lineage.
a Total number of H3.3 called peaks (tag number) (q- value = 5e−2) in the ChIP-seq of C2C12 (gray, 38439 peaks) (n = 1) and Hira-KO cells (black, 3744 peaks) (n = 1). b Percentage of the total number of H3.3 peaks distributed in distinct genomic regions as indicated in C2C12 or Hira-KO cells. c, d ChIP-seq average signal profiles (ratio to input) in the promoter region (±3 kb around the TSS), TSS, gene body, and TES for H3.3 (c) and H3K27ac (d) in C2C12 (orange) and Hira KO (blue) shown for all genes, upregulated genes in Hira KO, downregulated in Hira-KO and skeletal muscle genes (GO:0007519). e, f ChIP-seq average signal profiles (ratio to input) of H3.3 (e) and H3K27ac (f) plotted on the control ATAC-seq peaks in C2C12 (orange) and Hira KO (blue). g–m ATAC-seq (green), ChIP-seq profiles for H3.3 (orange), and H3K27ac (gray) in the genomic loci of Pax7 (g), HeyL (h), Col5a1(i), Dmrt2 (j), Myod1 (k), Fgfr4 (l), and Eya4 (m).
Hira-deficient satellite cells lack regenerative capacity
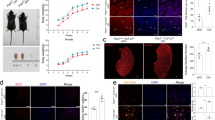
To demonstrate if HIRA is also required to preserve myogenic stem cell identity in vivo, we analyzed the consequences of ablating Hira in activated and proliferating satellite cells. To specifically delete Hira in adult muscle stem cells we inter-crossed Pax7CreErt2 and Hirafl/fl mouse lines22,23. Skeletal muscle satellite cells from adult Pax7CreErt2;Hirafl/fl mice treated with tamoxifen diet for 2 weeks displayed a sharp decrease in Hira expression (80%) and HIRA protein was not detected compared with control Pax7CreErt2;Hirafl/+ mice, without affecting DAXX (Supplementary Fig. 3a, b). Because muscle stem cells reside in a quiescence state in homeostasis, we performed injury experiments in the Tibialis anterior (TA) muscles using BaCl2 injection (Fig. 3a) that efficiently triggers satellite cell activation and muscle regeneration24. We analyzed the impact on regeneration 7 days post injury (dpi), to allow amplification of the proliferating satellite cell pool after activation, and at 28 dpi when the regeneration is completed3. At 7 dpi the number of PAX7-positive cells was significantly reduced in the conditional knockout (cKO) mice when normalized to the fiber number (Fig. 3b, c). Since HIRA regulates PAX7 expression (Fig. 1k, l), we further used M-Cadherin as a satellite cell marker and confirmed that the number of M-Cadherin-associated nuclei is decreased in the absence of HIRA in satellite cells, 7 dpi (Fig. 3d, e). At 28 dpi, the number of PAX7-positive cells in the Pax7CreErt2;Hirafl/fl muscles was further reduced compared to 7 dpi (Fig. 3f, g), showing that satellite cells lacking Hira are unable to replenish the muscle stem cell pool after muscle injury. Similar results were obtained 28 dpi in Pax7CreErt2;Hirafl/fl muscles when satellite cell number was normalized per area (Supplementary Fig. 3c, d), confirming the lack of capacity for the regenerating muscle to maintain the satellite cell pool. Consistently, the number of M-Cadherin-positive associated nuclei at 7 dpi, normalized to the unit area, was significantly decreased when Hira is deleted in satellite cells (Supplementary Fig. 3e). Consistent with the in vitro myogenic cell analysis, proliferation and cell death rates were not significantly affected in Pax7CreErt2;Hirafl/fl satellite cells (Supplementary Fig. 3f–h). Loss of satellite cells in Pax7CreErt2;Hirafl/fl injured muscles was associated with severely impaired regeneration capacity as shown by the high number of newly formed fibers 7 dpi labeled with embryonic myosin heavy chain (MYH3) antibody (Fig. 3h, i) and the failure to generate fibers of large cross-sectional area 28 dpi (Fig. 3j, k). These data demonstrate that muscle satellite cells lacking Hira fail to preserve the muscle stem cell pool and to efficiently regenerate damaged skeletal muscle.
a Schematic representation of experimental timeline. b Co-immunostaining using PAX7 (green), Laminin (LAM, red) and the nuclear marker DAPI (blue) in Pax7CreErt2;Hirafl/+ and Pax7CreErt2;Hirafl/fl TA muscles 7 dpi. c Quantification of the PAX7-positive cell number per 100 fibers of b (control n = 3 mice, cKO n = 4 mice). Error bars, mean ± SD, two-sided unpaired t-test. d Co-immunostaining using PAX7 (red), M-Cadherin (M-CADH, red: first panel; gray: last panel), Laminin (LAM, green) and the nuclear marker DAPI (blue) in Pax7CreErt2;Hirafl/+ and Pax7CreErt2;Hirafl/fl TA muscles 7 dpi. Arrows indicate M-CADH-positive cells. e Quantification of the M-CADH-positive cell number per 100 fibers in d (control n = 3 mice, cKO n = 3 mice). Error bars, mean ± SD, two-sided unpaired t-test. f Co-immunostaining using PAX7 (red), Laminin (LAM, green), and the nuclear marker DAPI (blue) in Pax7CreErt2;Hirafl/+ and Pax7CreErt2;Hirafl/fl TA muscles 28 dpi. Arrows indicate PAX7-positive cells. g Quantification of the PAX7-positive cell number per 100 fibers in f (control n = 3 mice, cKO n = 4 mice). Error bars, mean ± SD, two-tailed unpaired t-test. h Co-immunostaining using MYH3 (green), Laminin (LAM, red), and the nuclear marker DAPI (blue) in Pax7CreErt2;Hirafl/+ and Pax7CreErt2;Hirafl/fl TA muscles 7 dpi. i Quantification of the MYH3-positive fibers per area in h (control n = 3 mice, cKO n = 3 mice). Error bars, mean ± SD, two-tailed unpaired t-test. j Immunostaining using Laminin (LAM, red) and the nuclear marker DAPI (blue) in Pax7CreErt2;Hirafl/+ and Pax7CreErt2;Hirafl/fl TA muscles 28 dpi. k Quantification of the CSA in j as percentage of total fibers (control n = 3 mice, cKO n = 3 mice). Error bars, mean ± SD. Scale bars, 40 μm.
Hira-deficient satellite cells lose myogenic cell identity
Satellite cells lacking Hira are unable to sustain muscle regeneration. In order to analyze if Hira-deficient muscle stem cells display an alteration of lineage-specific gene expression, as observed in C2C12 cells, we performed RNA-seq on 5 dpi FACS-isolated satellite cells from control (Pax7CreErt2;Hirafl/+) and cKO (Pax7CreErt2;Hirafl/fl) mice. Sorted satellite cells were further cultured for 48 h to homogenize the stem cell population and ensure activation of all cells. We observed expression changes in 2925 genes when comparing control and cKO samples (1297 downregulated and 1628 upregulated) (Fig. 4a, b). Gene ontology analysis of downregulated genes in satellite cells lacking Hira included terms related to skeletal muscle satellite cell differentiation, fiber development, and regeneration (Fig. 4c). In contrast, gene ontology of upregulated genes was linked to angiogenic, cardiac, and endothelial processes, which suggests an increase in the expression of alternative lineage genes in the absence of HIRA in satellite cells (Fig. 4d). These results are consistent with the C2C12 gene ontology analysis (Fig. 1i, j). In the Hira-deficient satellite cells, muscle stem cell markers and myogenic markers are consistently downregulated (Fig. 4e, f), while genes from cardiac, endothelial, and other alternative lineages are upregulated (Fig. 4e, f). Moreover, satellite cells lacking Hira display positive VE-Cadherin (Cdh5 gene) immunostaining (Supplementary Fig. 3i, j). These results suggest that a similar mechanism is operating in C2C12 and in satellite cells to sustain the myogenic identify of these cells. In order to test this, we performed H3.3 ChIP-RT-qPCR in satellite cells and analyzed the known regulatory regions of Myod1 (−20 kb) and Fgfr4 (+19.2 kb)19,20, the genomic regions of Pax7 (+62.4 kb) and Eya4 (−380 bp) that display H3.3 enrichment in C2C12 cells (Fig. 2g, m) and the negative control for H3.3 binding (Myod1, −15 kb)11. All of these myogenic genes are downregulated in satellite cells (Fig. 4e, f). We observed that similar to C2C12 cells, H3.3 recruitment to the regulatory regions of these genes is lost in Hira-deficient satellite cells (Fig. 4g). Taken together, we conclude that H3.3 deposition by HIRA is required to maintain muscle stem cell gene expression and sustain a myogenic identity in satellite cells.
a Differential expression analysis of the RNA-seq analysis performed in 5 dpi satellite cells (n = 3 independent RNA samples for control and cKO). b MA-plot of control over Hira cKO satellite cells RNA-seq data. Significantly dysregulated genes are highlighted in red (FDR < 0.05). c, d Gene ontology analysis for biological processes of the downregulated (c) and upregulated (d) genes in cKO. Selected enriched terms are presented according to the fold enrichment. e Heatmap with the normalized reads per gene of control and Hira cKO RNA-seq data in individual triplicates. f RT-qPCR analyses in satellite cells (5 dpi) from Pax7CreErt2;Hirafl/+ and Pax7CreErt2;Hirafl/fl (n = 5 mice for Pax7, Myod1, Eya4, Angtp2; n = 4 mice for Myf5, Cdh5, Pdgfra; n = 3 mice for Fgfr4). For each gene, the mRNA levels of the control cells were normalized to 1. Error bars, mean ± SD, two-tailed paired t-test. g ChIP-RT-qPCR for H3.3 in Myod1 (−20 kb), Fgfr4 (+19.2 kb), Pax7 (+62.4 kb), Eya4 (−380 bp) and the negative control (−15 kb Myod1) of control (blue, n = 4) and Hira-depleted (red, n = 3) satellite cells (5 dpi). Error bars, mean ± SD, two-tailed unpaired t-test.
Alternative lineage genes are expressed in Hira-KO myoblasts
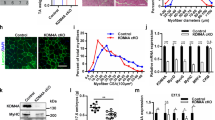
We identified a significant number of non-myogenic genes that were upregulated in Hira-deficient myogenic cells and that were previously shown to be silent in muscle cells, displaying either no histone marks or a bivalent state in C2C1215. Bivalent promoters are associated with silent genes characterized by both active H3K4me3 and repressive H3K27me3 histone marks, remaining in a poised state that can be resolved into an active or repressed state upon differentiation25. To define the epigenetic status of the genes upregulated in Hira-deficient cells, we performed ChIP-seq for H3K4me3 and H3K27me3 in control and Hira-mutant C2C12 myogenic cells. We observed that H3K4me3 signal intensity was strongly increased in promoters, TSS, and gene bodies of all protein-coding genes in Hira-deficient myogenic cells, while H3K27me3 signal was unchanged (Supplementary Fig. 4a, b). In addition, we analyzed the bivalency-associated marks in the promoter regions of upregulated genes, identifying a disruption of the H3K4me3 and H3K27me3 signal balance consistent with their induced expression (Fig. 5a). In contrast, the promoters of the downregulated genes do not show a bivalent profile in C2C12 or in Hira-KO cells (Fig. 5a). The Hoxb cluster, which was strongly upregulated in Hira-deficient myogenic cells (Fig. 1k, l), was previously described as presenting only repressive marks in C2C12 cells;15 on the contrary, we observed bivalent features at these loci with H3K27me3 and H3K4me3 marks (Fig. 5b), similarly to quiescent satellite cells26. Upon Hira deletion, the Hoxb cluster showed a concomitant increase in H3K4me3 modification and loss of H3K27me3 mark (Fig. 5b). Other bivalent genes, associated with adipogenic fate (Sox6, Cebpa) had increased H3K4me3 signal in their promoters but did not display modifications in H3K27me3 upon Hira deletion, indicating, together with the general absence of H3K27me3 changes in all genes, that the major histone mark associated with activation and expression of the bivalent genes is H3K4me3 (Supplementary Fig. 5c, d). Upregulated genes in Hira-deficient cells belonging to neuronal (Nefl, Nefm), endothelial (Cdh5, Angpt2), mesenchymal (Pdgfra), or osteogenic lineages (Runx2) did not display histone marks in myogenic cells (Fig. 5c–f and Supplementary Fig. 4e, f), associated with an absence of RNA expression of these genes. However, upon Hira deletion, these genes acquired H3K4me3 marks and became expressed (Fig. 5c–f and Supplementary Fig. 4e, f). Since MLL1/MLL2 methyltransferase complex is responsible for H3K4 trimethylation27, we investigated if this complex mediated the activation of alternative lineage genes in myogenic cells depleted of Hira. We analyzed the expression of Mll1 (Kmt2a) and Mll2 (Kmt2b) genes in Hira-deficient cells and found that they were upregulated 1.9- and 1.5-fold, respectively, compared with controls (Fig. 5g, h). To investigate if the upregulation of Mll1/Mll2 genes is promoting the expression of alternative lineage genes when Hira is deleted in myogenic cells, we knocked out either Mll1 or Mll2 in Hira-deficient cells (Supplementary Fig. 4g, h). Deleting Mll1 or Mll2 significantly decreased their RNA levels and both Hira/Mll1 dKO and Hira/Mll2 dKO lacked MLL1 protein (Supplementary Fig. 4g, h). Strikingly, abolishing Mll1 or Mll2 expression in Hira mutant cells prevented the expression of non-myogenic lineage genes (Cdh5, Nefl, Nefm) and Hoxb cluster gene Hoxb13 (Fig. 5i). The expression levels of Pax7, shown to be regulated by MLL1 in activated satellite cells28 were partially rescued in Hira/Mll1 and Hira/Mll2 dKO cell lines compared to Hira KO alone (Supplementary Fig. 4i). To evaluate if the expression of non-myogenic genes was linked to the H3K4 trimethyltransferase activity of MLL1/MLL2, we performed H3K4me3 ChIP-RT-qPCR and analyzed the promoter regions of Cdh5, Nefl, and Hoxb13. We confirmed that at these loci the enrichment of the H3K4me3 histone mark was reduced in Hira/Mll1 dKO cells compared to Hira mutant cells (Fig. 5j), confirming that increased H3K4me3 is linked to MLL1/MLL2 expression. Since H3K4me3 histone modification is increased across the genome in Hira-KO cells both in genes that are upregulated and downregulated (Supplementary Fig. 4a), we asked whether other histone modifications could be associated with the expression of alternative lineage genes. Since loss of H3.3 and decreased H3K27ac modification in myogenic gene loci is associated to decreased expression in Hira-KO cells, we asked whether acetylation in promoter regions of alternative lineage genes could be associated with their activation. Thus, we performed H3-acetyl ChIP-RT-qPCR (that recognizes acetylation in all N-terminal residues of H3 histones) at the promoter regions of Cdh5, Nefl, and Hoxb13. We observed that in Hira mutant cells, promoters of alternative lineage genes and Hox genes display an increased acetylation compared with control cells (Fig. 5k). Taken together these data demonstrate that, HIRA is required to prevent the expression of silent alternative non-myogenic lineage genes by (1) inhibiting Mll1/Mll2 expression and consequently preventing the H3K4 methyltransferase activity of these proteins and by (2) preventing acetylation of H3 histones at promoter regions of those genes.
a ChIP-seq average signal profiles (ratio to input) in the promoter region (±3 kb around the TSS) and TSS for H3K4me3 and H3K27me3 in C2C12 and Hira KO, plotted with the upregulated or the downregulated genes (n = 1). b–f ATAC-seq (green) and ChIP-seq profiles for H3K4me3 (blue) and H3K27me3 (dark red) in the Hoxb cluster (b) and in the genomic loci of Nefl (c), Cdh5 (d), Pdgfra (e), and Runx2 (f) genes. Gene lineages are indicated. g Heatmap with the normalized reads per gene of C2C12 and Hira-KO RNA-seq data for Mll1 and Mll2. h RT-qPCR analyses of the mRNA expression levels of Mll1 (light gray) and Mll2 (dark gray) in C2C12 (n = 4 independent RNA samples) and Hira KO (n = 4 independent RNA samples) cells. For each gene, the mRNA levels of the control cells were normalized to 1. Error bars, mean ± SD. two-tailed paired t-test. i RT-qPCR analyses of the mRNA expression levels of Cdh5, Nefl, Nefm, and Hoxb13, in C2C12, Hira KO, Hira/Mll1 dKO, and Hira/Mll2 dKO cells (n = 3 independent RNA samples per condition). Error bars, mean ± SD, two-tailed unpaired t-test. j ChIP-RT-qPCR for the H3K4me3 histone mark on the promoter region of Cdh5, Nefl, and Hoxb13 genes and in the negative control (intronic region of Cdh5) of Hira KO (blue) and Hira/Mll1 dKO (gray) cell lines (n = 3 independent biological samples per condition). Error bars, mean ± SD, two-tailed paired t-test. k ChIP-RT-qPCR for the H3ac (pan-acetyl) histone mark on the promoter region of Cdh5, Nefl, and Hoxb13 genes and in the negative control (intronic region of Cdh5) of control C2C12 (blue) and Hira KO (green) (n = 3 independent biological samples per condition). Error bars, mean ± SD, two-tailed unpaired t-test.
Discussion
In the murine myogenic cell line C2C12, it was previously shown that the histone chaperone activity of HIRA is required to deposit H3.3 in the core enhancer region of Myod1, to sustain its expression during differentiation11. In this myoblast cell line, forced expression of the canonical H3 variant H3.1 disrupts the balance of the H3.3/H3.1 in the nucleosomes, which inhibits myogenic gene expression and suppresses the differentiation potential of the cells10. Here we demonstrate that HIRA plays a considerably broader function by providing an appropriate chromatin landscape that promotes muscle gene expression via H3.3 incorporation, which is required to maintain the active transcription-associated mark H3K37ac at myogenic loci.
Whether H3.3 deposition-associated regulation of myogenic gene expression is an intrinsic function of the histone chaperone HIRA or if other factors are involved remains to be determined. The ubiquitous chromodomain helicases CHD1 and CHD2, which are recruited to active chromatin regions associated with TSS and enhancers in a transcription-coupled manner participate in the deposition of H3.39,29,30. Moreover, CHD2 deletion leads to reduced H3.3 occupancy at TSS and enhancers in a human lymphoblast cell line29. It was previously shown that MYOD physically interacts with CHD2 and that this interaction is required for H3.3 incorporation in the regulatory regions of myogenic terminal differentiation genes, such as MYOG, regulating their expression9. Although CHD1 physically associates with HIRA to mediate H3.3 deposition into chromatin in human lymphoblast cell lines, CHD1 is not involved in H3.3 deposition in Myod1 promoter or upstream regulatory regions in myogenic cells11,30. The H3.3 histone chaperone DAXX regulates the deposition of this histone variant in constitutive heterochromatin but it was also shown to deposit H3.3 in regulatory regions of neuronal genes during development, regulating their expression7,8,12. However, the role of DAXX as a histone chaperone of H3.3 during myogenesis remains to be evaluated. Altogether, HIRA appears to be the main histone chaperone required for H3.3 deposition in the myogenic lineage, while CHD2 could act either as a co-factor or incorporate H3.3 at the locus of MYOD downstream target genes.
The epigenetic determinants of lineage-committed adult stem cells are poorly understood compared to those of ESCs. In this study, we identify two major epigenetic differences between the adult myogenic stem cell lineage and what has been identified for ESCs: (1) the drastic transcriptomic changes when HIRA is deleted vs. no changes in ESCs8,14 and (2) the general loss of H3.3 deposition in the absence of HIRA in myoblasts vs. loss of H3.3 deposition in specific genomic loci in ESCs8. Although the pluripotent transcriptomic state of ESCs can be maintained in the absence of HIRA and depletion of H3.3 in TSS and gene bodies8, the deposition of H3.3 in regulatory regions of developmental genes is required for ESCs differentiation into the different germ layers14,31. In addition, while the deposition of H3.3 in promoter-distal enhancers (±3 kb around the TSS) of ESCs is required for H3K27 acetylation and enhancer activation, an effect in gene expression is only observed when the cells are triggered to differentiate14. These results suggest that H3.3 plays a major role in lineage commitment and maintenance, consistent with our observation that H3.3 is required to preserve myogenic gene expression and lineage potential of skeletal muscle stem cells. Although, it remains to be investigated if this mechanism is specific to the myogenic lineage or also operates in other tissues.
In this study, we combined the analysis of skeletal muscle stem cells with a myogenic cell line (C2C12) to address the requirement of HIRA-H3.3 in the acquisition of myogenic commitment and in its maintenance, respectively. We showed that the deposition of H3.3 by HIRA in myogenic genomic loci is required for gene expression in activated satellite cells upon muscle injury, as well as for the maintenance of muscle identity in already committed cells (C2C12). In vivo deletion of Hira reduces satellite cell numbers and severely compromises muscle regeneration. We showed that this is associated with loss of myogenic gene expression and muscle stem cell markers in the absence of HIRA. The percentage of proliferating PAX7-positive cells is not altered in Hira-deficient satellite cells, yet we cannot exclude that loss of muscle stem cells could be linked to an unbalanced asymmetric cell division, which is known to be required for replenishing the quiescent pool of satellite cells32,33. Given the consistency between the in vivo and in vitro data, we believe that the regeneration impairment phenotype is associated with the transcriptomic changes observed in satellite cells-lacking Hira and therefore to the role of HIRA in the maintenance of myogenic identity. Although, an alternative hypothesis in the context of muscle regeneration could be linked to the production of impaired myofibers. Future studies will evaluate the impact of myofiber specific conditional ablation of Hira to clarify this point.
We observed a striking and specific loss in muscle stem cell lineage gene expression linked to the decreased H3.3 deposition in Hira-deficient cells. Yet, the activation of gene expression from non-muscle lineages in cells depleted for HIRA did not occur through changes in H3.3 deposition at their loci and was associated with the increased and global activity of the H3K4 methyltransferases MLL1 and MLL2. Indeed, the MLL1/MLL2 complex has been shown to play a major role in lineage commitment during development, associated with regulation of Hoxb cluster expression, demonstrating that this methyltransferase complex is associated with the specification of new lineages34,35,36.
In this study, we identified an epigenetic mechanism by which HIRA, via H3.3 incorporation, stabilizes the myogenic identity of muscle stem cells and committed myoblasts by maintaining their gene expression active and by silencing alternative lineage genes through MLL1/MLL2 inhibition. Our study therefore supports a crucial role for HIRA-mediated H3.3 incorporation that subsequently stabilizes the active transcription-associated histone mark H3K27ac in lineage-specific cells to maintain the balanced expression of tissue-specific genes.
Methods
Mouse lines
All mice were kept under pathogen-free conditions with a 12 h/12 h light/dark cycle and a constant ambient temperature (22 °C) and humidity (34%). Adult male mice aged between 8 and 12 weeks were used as heterozygous Pax7CreErt2;Hirafl/+ (controls) or homozygous Pax7CreErt2;Hirafl/fl for the Hira floxed allele22,23. Animals were handled according to the European Community guidelines, implementing the 3Rs rule. Protocols were validated by the ethic committee of the French Ministry, under the reference number APAFIS#13695-2018021408521124.v2.
Tamoxifen administration and muscle injury
Mice were put on a solid tamoxifen diet (400 mg/kg of food) for 2 weeks and then kept 1 week with regular food. Mice were anesthetized intra-peritoneally with 3.5 μL/g of Ketamine/Xylazine at 20 and 10 mg/mL, respectively. The TA muscles were injected with 50 μl of 0.6% BaCl2 (Sigma, 202738).
Immunohistochemistry
Cryosections of 10 µm were performed on TA muscles previously dissected and frozen in liquid-nitrogen-cooled isopentane 7 or 28 dpi. The muscle sections or the cultured cells were fixed in 4% PFA for 15 min at room temperature, permeabilized in cold methanol for 6 min, boiled in citrate buffer (Dako, S1699) for epitope retrieval and blocked with 5% immunoglobin G (IgG)-free BSA (Jackson, 001-000-162) for 1 h at room temperature. To reduce background, the slides were incubated 30 min at room temperature with anti-mouse IgG Fab fragment (Jackson, 115-007-003). The sections were incubated with primary antibodies (Supplementary Table 1) diluted in 5% IgG-free BSA overnight at 4 °C. Secondary antibody incubation (Invitrogen, Alexa-Fluor) was performed 1 h at room temperature followed by DAPI (Sigma, D9542) staining (1:5000) 10 min at room temperature. EdU staining reaction was performed according to manufacturer’s guidelines (Invitrogen, EdU Click-iT PLUS Kit, C10640).
Western blot
Cells were lysed in RIPA buffer (50 mM Tris HCl (Sigma, T5941), 150 mM NaCl (Sigma, 9625), 1% Igepal (Sigma, I8896), 0.5% Sodium deoxycholate (Sigma, D6750), 0.1% SDS (Sigma, L3771), 1 mM EDTA (Sigma, E5134), 1x Protease Inhibitor Cocktail (Roche, 04-693-116-001); pH8) for protein extraction. Overall, 10 μg of protein were used per sample. The blocking of the membrane was performed in 0.5% of gelatin from cold water fish (Sigma, G7765) and 5% Tween20 (Sigma, P9416) in PBS (Invitrogen, 003002) 1 h at room temperature. Primary antibodies were diluted in blocking solution and incubated 2 h at room temperature (Supplementary Table 1). Secondary antibodies were diluted in blocking solution and incubated 1 h at room temperature (Supplementary Table 1). Revelation reaction was performed 5 min at room temperature using SuperSignal West Pico (ThermoScientific, 35060). Western blot quantification was performed using Image J37. Uncropped and unprocessed blots are displayed in the Source Data file.
Muscle dissection and satellite cell isolation
FACS-isolation of satellite cells was performed as following18. Muscles were dissected, minced, and incubated in digestion buffer: HBSS (Gibco, 14025092), 0.2% BSA (Sigma, A7906), 2 μg/ml Collagenase A (Roche, 11088793001), and 3.25 μg/ml Dispase II (Roche, 04942078001) for 2 h at 37 °C, and purified by filtration using 100 µm and 40 µm cell strainers (BD Falcon, 352360 and 352340). Cell suspensions were incubated 30 min on ice with the following antibodies: Alexa700-anti-ITGA7, BV421-anti-CD34, PE-Cy7-anti-TER119, PE-Cy7-anti-CD45, and PE-anti-Ly-6/E (SCA1) (Supplementary Table 1). Cells were sorted by gating CD34 + ITGA7+ double positive lineage as shown in Supplementary Fig. 5.
Cell culture of C2C12 cells
C2C12 cell line (ATCC: CRL1772), obtained from DSMZ Germany, were grown in DMEM (Gibco, 41966029) supplemented with 10% fetal calf serum (FCS) (Eurobio, CVFSVF00-01) and 1% Pen/Strep (Gibco, 15070063). For differentiation assays, C2C12 were seeded at a confluence of 80% with 2% FCS for 3 days. For EdU proliferation analysis, cells were incubated with 10 mM EdU (Invitrogen, A10044) for 24 h (Invitrogen, EdU Click-iT PLUS Kit, C10640).
Cell culture of satellite cells
FACS-isolated satellite cells were plated on Matrigel (Corning, 354248)-coated dishes and cultured in DMEM (Gibco, 41966029) with 20% FCS (Eurobio, CVFSVF00-01), 1% Pen/Step (Gibco, 15070063), and 4 ng/mL basic FGF (bFGF) (Peprotech, 450-33).
Generation of KO C2C12 line using CRISPR-Cas9
Hira, Mll1, and Mll2 knockouts were generated by CRISPR/Cas9 using the pU6-(BbsI)_CBh-Cas9-T2A-mCherry plasmid, a gift from Ralf Kuehn (Addgene plasmid # 64324; http://n2t.net/addgene:64324; RRID:Addgene_64324)38. The single guide RNAs (sgRNAs) were obtained from the sgRNA optimized library39. The primers were designed with added BbsI (ThermoScientific, FD1014) restriction sites and an extra G/C for increased hU6 promoter efficiency (Supplementary Table 2), annealed and ligated to BbsI-linearized pU6-CBh-Cas9-T2A-mCherry plasmid using the T4-ligase (ThermoScientific, EL0011). A total of 1.5 × 105 C2C12 cells were transfected with 7 μg of plasmid with Lipofectamine LTX PLUS reagent (Invitrogen, 15338030) and 3 days post transfection the cells were FACS-isolated for mCherry-positive. The sorted cells were seeded at a low confluence (500 cells per 10 cm dish) in dishes previously coated with 0.1% gelatin from pork skin (Sigma, G1890). Individual clones were isolated, expanded, genotyped by PCR and sequenced to confirm the mutation.
RNA extraction, cDNA synthesis and RT-qPCR
A minimum of 2 × 105 C2C12 cells, cultured in high serum-containing medium and at low density, were collected per sample for RNA extraction (Macherey Nagel, 740955) following the manufacturer protocol. A minimum of 105 FACS-isolated satellite cells were collected per sample for RNA extraction. Reverse transcription was performed using the SuperScript III Reverse Transcriptase (Invitrogen, 18080-093) following the manufacturer’s guidelines. RT-qPCR was performed using the PowerUp SYBR Green Master Mix (Applied Biosystems, A25742). The relative mRNA levels were calculated using the 2^−ΔΔCt method40. The ΔCt were obtained from Ct normalized to the housekeeping gene Tbp levels in each sample. The RT-qPCR primers used are listed in the Supplementary Table 2.
RNA-sequencing of C2C12 cells
RNA was prepared as described for RNA extraction and sent to Integragen. Libraries were prepared with TruSeq Stranded Total RNA Sample preparation kit according to supplier recommendations. Briefly, the key stages of this protocol are successively, the removal of ribosomal RNA fraction from 1 µg of total RNA using the Ribo-Zero Gold Kit; fragmentation using divalent cations under elevated temperature to obtain `300 bp pieces; double strand cDNA synthesis using reverse transcriptase and random primers, and finally Illumina adapters ligation and cDNA library amplification by PCR for sequencing. Sequencing was carried out on paired-end 75 bp of Illumina HiSeq4000.
RNA-sequencing of satellite cells
RNA was prepared as described for RNA extraction and sent to the IMRB (Institut Mondor de Recherche Biomédicale) genomic platform. Libraries were prepared with TruSeq Stranded Total Library preparation kit according to supplier recommendations. Briefly, the key stages of this protocol are successively, the removal of ribosomal RNA fraction from 400 ng of total RNA using the Ribo-Zero Gold Kit; fragmentation using divalent cations under elevated temperature to obtain ~300 bp pieces; double strand cDNA synthesis using reverse transcriptase and random primers, and finally Illumina adapters ligation and cDNA library amplification by PCR for sequencing. Sequencing was carried out on single-end 75 bp of Illumina NextSeq500.
RNA-sequencing analysis
The RNA-seq analysis was performed using the Galaxy web platform41, public server https://usegalaxy.org. The FASTQ files were uploaded in Galaxy and formatted as Sanger using the FASTQ groomer tool42. Quality of the data was analyzed with FastQC tool v0.72. FASTQ Sanger files were aligned to the mm10 mouse genome using the built-in index of BOWTIE2 v2.3.4.243. Genes were counted using featureCounts v1.6.3 + galaxy244 and differently expressed genes were determined by DESeq2 v2.11.40.6 + galaxy145, obtaining also the MA-Plot and the sample-to-sample distances plot. Gene ontology analyses were performed on http://geneontology.org/ using the biological process option. The gene expression heatmap was created with displayR software with normalized reads for each triplicate.
Chromatin immunoprecipitation (ChIP)
Native ChIP was performed as following46. Briefly, C2C12 cells, cultured in high serum-containing medium and at low density, or FACS-isolated satellite cells (as described above) were trypsinized, washed, and subjected to nuclei isolation, chromatin fragmentation using Micrococcal Nuclease (MNase) (Sigma, N5386) and nucleosome purification by Hydroxyapatite (BioRad, 158-2000) chromatography. Immunoprecipitation was performed overnight at 4 °C with 6 μg of chromatin and 5 μg of antibody (Supplementary Table 1), previously incubated with Protein-A- (for H3K27ac, H3K4me3, H3K27me3, and H3ac antibodies) or Protein-G-coated (for H3.1 and H3.3 antibodies) magnetic beads (Diagenode, C03010020, and C03010021, respectively) and analyzed by RT-qPCR as percentage of the input or by sequencing. For sequencing, samples were sent to the IRCM (Institut de recherches cliniques de Montréal) molecular biology platform. Library was prepared using KAPA Hyper Prep Kits with PCR Library Amplification/Illumina series (Roche, 07962363001) with IDT for Illumina TruSeq DNA-RNA UD 96 Indexes (UDI) (Illumina, 2023784) and quantified by RT-qPCR using NEBNext Library Quant Kit for Illumina (NEB, E7640AA). Sequencing was carried out on paired-end 50 bp of Illumina HiSeq4000.
ChIP-sequencing analysis
The ChIP-seq analysis was performed using the Galaxy web platform41 public server https://usegalaxy.org. The FASTQ files were uploaded in Galaxy and formatted as Sanger using the FASTQ groomer tool42. Quality of ChIP-seq data was analyzed with FastQC tool v0.72. FASTQ Sanger files were aligned to the mm10 mouse genome using the built-in index of BOWTIE2 v2.3.4.243. The peaks were called with MACS2 v2.1.1.247 using paired-end BAM files with the cutoff q value 5−e2 and the broad region calling off, the input was used as control. MACS2 resulting bedgraph file was converted to bigwig format using the wig/bedgraph-to-bigwig converter v1.1.1 for visualization in the Integrated Genome Browser v9.1.448. The called peaks were annotated using the ChIPseeker v1.18.0 tool49. Average signal graphs were performed using computeMatrix Galaxy version 3.3.2.0.050 to prepare data for plotting using plotProfile Galaxy version 3.3.2.0.050.
ATAC-sequencing
C2C12 cells, cultured in high serum-containing medium and at low density, were harvested and frozen in culture media containing 10% FCS (Eurobio, CVFSVF00-01) and 10% DMSO (ThermoScientific, 20688). Cryopreserved cells were sent to Active Motif to perform the ATAC-seq assay. The cells were then thawed in a 37 °C water bath, pelleted, washed with cold PBS, and tagmented as previously described51, with some modifications based on52. Briefly, cell pellets were resuspended in lysis buffer, pelleted and tagmented using the enzyme and buffer provided in the Nextera Library Prep Kit (Illumina). Tagmented DNA was then purified using the MinElute PCR purification kit (Qiagen), amplified with ten cycles of PCR and purified using Agencourt AMPure SPRI beads (Beckman Coulter). Resulting material was quantified using the KAPA Library Quantification Kit for Illumina platforms (KAPA Biosystems) and sequenced with PE42 sequencing on the NextSeq 500 sequencer (Illumina).
ATAC-sequencing analysis
Reads were aligned using the BWA algorithm (mem mode; default settings). Duplicate reads were removed, only reads mapping as matched pairs and only uniquely mapped reads (mapping quality ≥ 1) were used for further analysis. Alignments were extended in silico at their 3′-ends to a length of 200 bp and assigned to 32-nt bins along the genome. The resulting histograms (genomic “signal maps”) were stored in bigWig files. Peaks were identified using the MACS 2.1.0 algorithm at a cutoff of p value 1e−7, without control file and with the –nomodel option. Signal maps and peak locations were used as input data to Active Motif’s proprietary analysis program, which creates Excel tables containing detailed information on sample comparison, peak metrics, peak locations, and gene annotations.
Image capturing, quantification, and statistical analysis
Image capturing was performed using a Zeiss LSM 800 confocal microscope with the associated Zeiss Zen Lite v2.3software. Quantifications were performed in at least five pictures taken randomly within each sample, per experiment, using Image J37 and statistics were done with GraphPad Prism version 8 and Excel. The statistical test performed in each analysis is described in the associated figure legend.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The RNA-seq, ChIP-seq, and ATAC-seq sequencing data that support the findings of this study have been deposited in GEO NCBI with the accession codes “GSE161056” and “GSE167911”. All other relevant data supporting the key findings of this study are available within the article and its Supplementary Information files or from the corresponding author upon reasonable request. Source data are provided with this paper. A reporting summary for this article is available as a Supplementary Information file. Source data are provided with this paper.
References
Mauro, A. Satellite cell of skeletal muscle fibers. J. Biophys. Biochem. Cytol. 9, 493–495 (1961).
Buckingham, M. & Relaix, F. The role of pax genes in the development of tissues and organs: Pax3 and Pax7 regulate muscle progenitor cell functions. Annu. Rev. Cell Dev. Biol. 23, 645–673 (2007).
Baghdadi, M. B. & Tajbakhsh, S. Regulation and phylogeny of skeletal muscle regeneration. Dev. Biol. 433, 200–209 (2018).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Ng, R. K. & Gurdon, J. B. Epigenetic inheritance of cell differentiation status. Cell Cycle 7, 1173–1177 (2008).
Ng, R. K. & Gurdon, J. B. Epigenetic memory of an active gene state depends on histone H3.3 incorporation into chromatin in the absence of transcription. Nat. Cell Biol. 10, 102–109 (2008).
Drane, P., Ouararhni, K., Depaux, A., Shuaib, M. & Hamiche, A. The death-associated protein DAXX is a novel histone chaperone involved in the replication-independent deposition of H3.3. Genes Dev. 24, 1253–1265 (2010).
Goldberg, A. D. et al. Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell 140, 678–691 (2010).
Harada, A. et al. Chd2 interacts with H3.3 to determine myogenic cell fate. EMBO J. 31, 2994–3007 (2012).
Harada, A. et al. Incorporation of histone H3.1 suppresses the lineage potential of skeletal muscle. Nucleic Acids Res. 43, 775–786 (2015).
Yang, J.-H. et al. Myogenic transcriptional activation of MyoD mediated by replication-independent histone deposition. Proc. Natl Acad. Sci. 108, 85–90 (2011).
Michod, D. et al. Calcium-dependent dephosphorylation of the histone chaperone DAXX regulates H3.3 loading and transcription upon neuronal activation. Neuron 74, 122–135 (2012).
Ricketts, M. D. & Marmorstein, R. A molecular prospective for HIRA complex assembly and H3.3-specific histone chaperone function. J. Mol. Biol. 429, 1924–1933 (2017).
Martire, S. et al. Phosphorylation of histone H3.3 at serine 31 promotes p300 activity and enhancer acetylation. Nat. Genet. 51, 941–946 (2019).
Asp, P. et al. Genome-wide remodeling of the epigenetic landscape during myogenic differentiation. Proc. Natl Acad. Sci. USA. 108, E149–E158 (2011).
Campos, E. I. & Reinberg, D. New chaps in the histone chaperone arena. Genes Dev. 24, 1334–1338 (2010).
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009).
Machado, L. et al. In situ fixation redefines quiescence and early activation of skeletal muscle stem cells. Cell Rep. 21, 1982–1993 (2017).
Goldhamer, D. J. et al. Embryonic activation of the myoD gene is regulated by a highly conserved distal control element. Development 121, 637–649 (1995).
Lagha, M. et al. Pax3 regulation of FGF signaling affects the progression of embryonic progenitor cells into the myogenic program. Genes Dev. 22, 1828–1837 (2008).
Sato, T., Rocancourt, D., Marques, L., Thorsteinsdóttir, S. & Buckingham, M. A Pax3/Dmrt2/Myf5 regulatory cascade functions at the onset of myogenesis. PLoS Genet. 6, e1000897 (2010).
Murphy, M. M., Lawson, J. A., Mathew, S. J., Hutcheson, D. A. & Kardon, G. Satellite cells, connective tissue fibroblasts and their interactions are crucial for muscle regeneration. Dev. Camb. Engl. 138, 3625–3637 (2011).
White, J. K. et al. Genome-wide generation and systematic phenotyping of knockout mice reveals new roles for many genes. Cell 154, 452–464 (2013).
Hardy, D. et al. Comparative study of injury models for studying muscle regeneration in mice. PloS One 11, e0147198 (2016).
Blanco, E., González-Ramírez, M., Alcaine-Colet, A., Aranda, S. & Di Croce, L. The bivalent genome: characterization, structure, and regulation. Trends Genet. 36, 118–131 (2020).
Liu, L. et al. Chromatin modifications as determinants of muscle stem cell quiescence and chronological aging. Cell Rep. 4, 189–204 (2013).
Shilatifard, A. MoleculaR Implementation and Physiological Roles for Histone H3 lysine 4 (H3K4) methylation. Curr. Opin. Cell Biol. 20, 341–348 (2008).
Addicks, G. C. et al. MLL1 is required for PAX7 expression and satellite cell self-renewal in mice. Nat. Commun. 10, 4256 (2019).
Siggens, L., Cordeddu, L., Rönnerblad, M., Lennartsson, A. & Ekwall, K. Transcription-coupled recruitment of human CHD1 and CHD2 influences chromatin accessibility and histone H3 and H3.3 occupancy at active chromatin regions. Epigenetics Chromatin 8, 4 (2015).
Konev, A. Y. et al. CHD1 motor protein is required for deposition of histone variant H3.3 into chromatin in vivo. Science 317, 1087–1090 (2007).
Semba, Y. et al. Chd2 regulates chromatin for proper gene expression toward differentiation in mouse embryonic stem cells. Nucleic Acids Res. 45, 8758–8772 (2017).
Kuang, S., Kuroda, K., Le Grand, F. & Rudnicki, M. A. Asymmetric self-renewal and commitment of satellite stem cells in muscle. Cell 129, 999–1010 (2007).
Le Grand, F., Jones, A. E., Seale, V., Scimè, A. & Rudnicki, M. A. Wnt7a activates the planar cell polarity pathway to drive the symmetric expansion of satellite stem cells. Cell Stem Cell 4, 535–547 (2009).
Deng, C. et al. HoxBlinc RNA recruits Set1/MLL complexes to activate hox gene expression patterns and mesoderm lineage development. Cell Rep. 14, 103–114 (2016).
Iimura, T. & Pourquié, O. Collinear activation of Hoxb genes during gastrulation is linked to mesoderm cell ingression. Nature 442, 568–571 (2006).
Liu, H.-C. et al. Expression of HOXB genes is significantly different in acute myeloid leukemia with a partial tandem duplication of MLL vs. a MLL translocation: a cross-laboratory study. Cancer Genet. https://doi.org/10.1016/j.cancergen.2011.02.003 (2011).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Chu, V. T. et al. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 33, 543–548 (2015).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods San. Diego Calif. 25, 402–408 (2001).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 44, W3–W10 (2016).
Blankenberg, D. et al. Manipulation of FASTQ data with Galaxy. Bioinformatics 26, 1783–1785 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Brand, M., Rampalli, S., Chaturvedi, C.-P. & Dilworth, F. J. Analysis of epigenetic modifications of chromatin at specific gene loci by native chromatin immunoprecipitation of nucleosomes isolated using hydroxyapatite chromatography. Nat. Protoc. 3, 398–409 (2008).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Freese, N. H., Norris, D. C. & Loraine, A. E. Integrated genome browser: visual analytics platform for genomics. Bioinforma. Oxf. Engl. 32, 2089–2095 (2016).
Yu, G., Wang, L.-G. & He, Q.-Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Acknowledgements
We thank Matthew Borok, Despoina Mademtzoglou, Valentina Taglietti, Philippos Mourikis, and Delphine Duprez for reading and commenting on the manuscript. We thank Carole Conejero and Stéphane Kerbrat from the IMRB genomic platform. We thank Adeline Henry, Aurélie Guguin, and Odile Ruckebusch from the IMRB cytometry platform. We thank Odile Neyret from the IRCM molecular biology platform. We thank the animal facilities EP3 from IMRB and TAAM from CDTA. This work was supported by funding to FR from Association Française contre les Myopathies (AFM) via TRANSLAMUSCLE (PROJECT 19507 and 22946), Agence Nationale pour la Recherche (ANR) grant Epimuscle (ANR 11 BSV2 017 02) and RHU CARMMA (ANR-15-RHUS-0003). This work was also supported by Labex REVIVE (ANR-10-LABX-73), including a post-doctoral grant to JEdL and a PhD fellowship to L.M.
Author information
Authors and Affiliations
Contributions
J.E.d.L. designed, performed, and analyzed the majority of the experiments. R.B.A., L.M., and Y.L. performed and analyzed experiments, contributing equally to this work. B.D.L. performed experiments. F.J.D. supervised experiments. F.R. designed and supervised experiments and oversaw the project. J.E.d.L. and F.R. wrote the manuscript. R.B.A., L.M., Y.L., B.D.L., and F.J.D. read and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Pier Lorenzo Puri, Terence Partridge and Vittorio Sartorelli for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Esteves de Lima, J., Bou Akar, R., Machado, L. et al. HIRA stabilizes skeletal muscle lineage identity. Nat Commun 12, 3450 (2021). https://doi.org/10.1038/s41467-021-23775-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-021-23775-9
This article is cited by
-
Histone chaperone HIRA complex regulates retrotransposons in embryonic stem cells
Stem Cell Research & Therapy (2022)
-
Tcf12 is required to sustain myogenic genes synergism with MyoD by remodelling the chromatin landscape
Communications Biology (2022)
-
Master regulators of skeletal muscle lineage development and pluripotent stem cells differentiation
Cell Regeneration (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.