Abstract
Purpose
Prostate cancer is a heterogeneous disease with variable clinical outcomes. Despite numerous recent approvals of novel therapies, castration-resistant prostate cancer remains lethal. A “real-world” clinical-genomic database is urgently needed to enhance our characterization of advanced prostate cancer and further enable precision oncology.
Methods
The Prostate Cancer Precision Medicine Multi-Institutional Collaborative Effort (PROMISE) is a consortium whose aims are to establish a repository of de-identified clinical and genomic patient data that are linked to patient outcomes. The consortium structure includes a (1) bio-informatics committee to standardize genomic data and provide quality control, (2) biostatistics committee to independently perform statistical analyses, (3) executive committee to review and select proposals of relevant questions for the consortium to address, (4) diversity/inclusion committee to address important clinical questions pertaining to racial disparities, and (5) patient advocacy committee to understand patient perspectives to improve patients’ quality of care.
Results
The PROMISE consortium was formed by 16 academic institutions in early 2020 and a secure RedCap database was created. The first patient record was entered into the database in April 2020 and over 1000 records have been entered as of early 2021. Data entry is proceeding as planned with the goal to have over 2500 patient records by the end of 2021.
Conclusions
The PROMISE consortium provides a powerful clinical-genomic platform to interrogate and address data gaps that have arisen with increased genomic testing in the clinical management of prostate cancer. The dataset incorporates data from patient populations that are often underrepresented in clinical trials, generates new hypotheses to direct further research, and addresses important clinical questions that are otherwise difficult to investigate in prospective studies.
Similar content being viewed by others
Introduction
Despite advances in treatment, the median overall survival from onset of metastatic castrate-resistant prostate cancer (mCRPC) remains dismal [1]. In this subset of patients, there is a variable response to currently available therapies, and treatment in these men has not historically been guided by molecular biomarkers. Over the last decade, advancements in genomic sequencing technologies have allowed for a deeper understanding of the molecular complexity of this disease. Many potentially actionable alterations are now identified, fueling biomarker-based clinical trials of novel molecularly targeted agents as well as standard therapies. An important example of genomically tailored therapy in prostate cancer is the utility of Poly-(ADP ribose) polymerase (PARP) inhibitors in patients whose tumors harbor Homologous Recombination Repair (HRR) defects. The demonstrated clinical efficacy of olaparib [2] and rucaparib [3] in metastatic CRPC with HRR defects and their subsequent approval by the US Food and Drug Administration (FDA) marked an important milestone in the implementation of precision medicine for this disease.
The approvals of PARP inhibitors make it imperative to obtain somatic sequencing for all men with advanced prostate cancer. In addition, germline testing is also recommended for all men with metastatic prostate cancer per National Comprehensive Cancer Network guidelines, and in selected men with localized disease based on clinical risk and histologic subtypes as well as family history [4]. In fact, germline mutations in HRR genes exists in ~5% of localized prostate cancer [5]. In addition, multiple guideline panels recommend genomic sequencing for all patients with advanced prostate cancer. With this shifting paradigm and advancements in technology, there will be a rise in availability, breadth, and scope of genomic data derived from CLIA-based testing platforms. Linkage of such data with granular patient outcomes can be leveraged to improve our understanding of the molecular mechanisms that lead to variable clinical outcomes across prostate cancer disease states over time, identify additional subgroups of patients whose tumors are more vulnerable to specific therapies, and help guide decisions of optimal sequencing of therapies in prostate cancer.
The Prostate Cancer Precision Medicine Multi-Institutional Collaborative Effort (PROMISE) is a consortium of academic cancer centers with the goal of better defining the clinical-genomic features across the entire prostate cancer disease spectrum. Herein, we outline the rationale, design, and objectives of our multi-institutional, retrospective clinical-genomic database of advanced prostate cancer patients that addresses this important need.
Molecular landscape of advanced prostate cancer
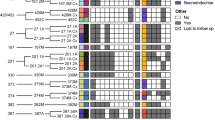
Large-scale genomic analyses of metastatic prostate tumors have demonstrated a high frequency of germline and somatic alterations in several cancer-specific genes that are actionable or currently being investigated as candidate predictive biomarkers (Fig. 1) [6,7,8,9]. Some are more commonly seen in localized disease such as SPOP mutations and ETS family gene fusions [10]. The most common gene alterations enriched in CRPC, compared to earlier disease states, include AR and TP53, which are present in >50% of cases [6, 9]. Other commonly affected pathways with important clinical and therapeutic implications include the PI3K pathway genes such as PTEN-AKT-mTOR, and HRR genes such as BRCA1/2, ATM which are altered in ~45%, ~25%, ~7-10% of CRPC cases, respectively [6, 8]. Additional altered genes in CRPC include those involved in the cell cycle (~30%) such as RB loss, CDKN1B, and CCND1; epigenetic regulator genes (~25%) such as KMT2C, KMT2D, and CHD1; WNT pathway genes (~15%) such as APC and CTNNB1, CDK12 (~7%); and MAP kinase pathway genes (~5%) [6, 8, 9]. These alterations in the context of drug development and clinical decision-making are discussed in further detail in the next section. Many of these alterations occur concurrently, and based on prior reports, about 65–85% of analyzed CRPC tumors harbor potentially actionable alterations beyond AR, defined as the ability to predict response to an available drug based on existing preclinical data [6]. Structural variants in the non-protein-coding regions also have a potential impact on the activity of important regulators of cancer progression [9]. A recent whole-genome sequencing analysis, the first of its kind in CRPC, identified that 81% of cases harbored amplification of a putative enhancer of AR, which may drive androgen resistance [9, 11]. Structural variants were also demonstrated near the MYC, TP53, CDK12, FOXA1, and BRCA2 genes with potential biological and clinical implications.
In metastatic hormone-sensitive prostate cancer (mHSPC), treatment options have rapidly expanded, yet decision-making is based mainly on clinical factors without integration of genomic biomarkers. A recent study describes potential biomarkers of disease aggressiveness [7]. Specifically, alterations involving AR, TP53, cell cycle, and MYC pathways predict for a shorter time to castration resistance, while alterations in SPOP gene and WNT pathway are associated with lower rate and longer time to castration resistance [7]. Furthermore, in tumors with high-volume (≥ 4 bone metastases and/or visceral metastasis) mHSPC, higher fraction of alterations indicative of genomic instability, and alterations involving cell cycle, epigenetic regulation, and NOTCH pathways are seen. Thus, the development of a clinical-genomic model for both mHSPC as well as CRPC represents a major unmet medical need, for which a large number of patients are needed with careful clinical and genomic annotation.
Implementation of precision medicine in prostate cancer
Our understanding of the molecular landscape of advanced prostate cancer provides a platform for the development of targeted therapies, and for optimizing therapeutic combination strategies and sequencing. Excitingly, intense investigation is already taking place to leverage this knowledge and expand therapeutic options for men with advanced prostate cancer. Increasingly, genomic sequencing is also becoming a pre-requisite for clinical trials, thus availability of routine genetic testing becomes imperative to ensure patients have access to biomarker-driven therapies and clinical trials.
Approximately 20–25% of men with mCRPC have germline or somatic alterations involving HRR genes, such as BRCA1/2 and ATM, which may predict for sensitivity to PARP inhibitors and platinum agents [12]. Large clinical trials such as PROfound and TRITON2 have led to the FDA approval of two PARP inhibitors, olaparib and rucaparib, respectively, in men with mCRPC that harbor deleterious somatic or germline mutations in HRR genes [2, 3]. In May 2020, olaparib was approved for men with HRR gene-altered CRPC while the rucaparib approval was restricted to patients with tumors harboring alterations in BRCA1/2. Despite these approvals, many questions remain about the relevance of PARP inhibitors in patients harboring non-BRCA HRR genes. Many of these questions can be answered with a real-world database such as PROMISE. In addition to PARP inhibitors, exceptional responses to platinum-based chemotherapy have also been observed in BRCA2-mutant CRPC cases [13, 14]. However, not much is known in this population about the utility of platinum agents and PARP inhibitors in combination or as sequential therapy. Other HRR gene targets under clinical development include ATR inhibitors and ATM inhibitors. Of specific interest within the HRR pathway is CDK12 loss, which has shown to correspond with high focal tandem duplications and high neoantigen burden, potentially sensitizing CDK12-altered tumors to immune checkpoint blockade [15,16,17]. Thus, knowledge from real-world datasets on the appropriate sequencing and outcomes of taxane chemotherapy with PARP inhibitors in less common subgroups of men with mCRPC such as ATM mutations or less common HRR gene alterations is needed.
About 45% men with mCRPC harbor pathogenic genomic alterations within the PI3K pathway. The majority of alterations involving this pathway occur in the PTEN gene (~40%), which has previously proven to be a difficult target with single agents and is associated with poor prognosis [18]. Preclinical data suggest that PTEN/PI3K and AR pathways have reciprocal crosstalk, thus targeting both pathways in combination may enhance therapeutic efficacy [19]. A phase II study of abiraterone acetate and AKT inhibitor, ipatasertib, showed a clinical benefit particularly in PTEN-deficient mCRPC [20]. In the primary analysis of the phase III study of this combination (IPATential150, NCT03072238), combined AR and AKT blockade provided an improved progression-free survival in patients with mCRPC with PTEN loss, compared to AR blockade alone [21]. Another target within this pathway is AKT1 gene, which occurs in about 1% of mCRPC patients. AKT inhibitors exhibit clinical activity in AKT-mutated breast cancer and other solid tumors [22] and are currently in development for mCRPC (NCT04087174). Other alterations of interest in this pathway involve PIK3CA, PIK3C2B, BRAF, MAP2K1, and KRAS genes. Recently the first PI3KCA inhibitor, alpelisib, was approved for PIK3CA-mutated breast cancer [23], a development that can inform future clinical trial designs and treatment options for advanced prostate cancer as well.
About 3–5% of patients with CRPC also have evidence of DNA mismatch repair deficiency (dMMR) and/or microsatellite instability (MSH)-high that are found through immunohistochemistry to evaluate for loss of MMR protein and/or DNA sequencing [24]. These patients tend to have tumors with a higher tumor mutational burden (TMB), predicting response to immune checkpoint blockade [25,26,27,28]. In 2017, based on data from five multi-cohort, single-arm studies of MSH-H or dMMR advanced solid tumors [29, 30], pembrolizumab received a tumor-agnostic approval for patients who progressed on prior treatment and did not have satisfactory alternative treatment options. Although this study marked the first time in oncology drug development that a therapy was approved based on a specific biomarker irrespective of tumor histology, very few patients with prostate cancer were enrolled. Furthermore, it is not clear whether routine testing of dMMR and/or MSI is being performed in most clinical practices to identify this subset of prostate cancer patients. More recently, tumor-agnostic pembrolizumab approval was expanded to include tumors with high TMB [31].
In addition to potential actionable alterations discussed, multi-institution clinical trials such as Targeted Agent and Profiling Utilization Registry (TAPUR, NCT02693535) by American Society of Clinical Oncology highlight the aims of understanding the safety and efficacy of novel targeted agents in advanced prostate cancer and other malignancies. In addition, large registries of real-world data are being built to identify novel targets. The American Association of Cancer Research (AACR) has implemented Project GENIE (Genomics Evidence Neoplasia Information Exchange), which consist of real-world data among 19 leading cancer centers in the world. Despite the robustness of these pan-cancer platforms, the nuances of different clinical states of prostate cancer will not be captured. To complement these efforts, our consortium was developed to leverage the wealth of existing clinical-genomic data to expand our understanding of prostate cancer biology and to improve current therapeutic approaches. In the remaining sections, we highlight some key clinical questions that are present in routine practice to provide context into the design, structure, and objectives of PROMISE.
Current challenges in prostate cancer and potential solutions
The last two decades have seen significant new advancements in the treatment of metastatic prostate cancer. Much wider availability of next-generation sequencing (NGS) panels has made it possible to individualize treatment options for a subset of prostate cancer patients with specific tumor markers or alterations. These developments have created a plethora of new questions, many of which may be answered though accumulation of clinical and genomic data in a “real-world” dataset. The following examples highlight current challenges in the treatment of advanced prostate cancer as well as potential solutions and strategies to overcome barriers to the realization of the true promise of precision medicine in prostate cancer.
Sequencing of therapies in prostate cancer
With the evolution of multiple novel treatment options in mCRPC, including novel hormonal agents and PARP inhibitors, as well as the emergence of novel therapeutic classes such as PSMA-targeted radioligand therapies, there is increasingly a wealth of options to offer patients in a disease space where only a couple of decades ago there were relatively few [32, 33]. In this setting, the questions about appropriate sequencing of therapies become especially relevant, especially as most patients will not be able to receive all available treatment options and decisions of which therapies to prioritize earlier as opposed to later in the disease course must be made by the treating physician. For instance, in molecularly selected patients able to receive PARP inhibitors, what is the most optimal way to sequence these agents with taxane chemotherapy and should the sequencing of therapies be impacted by mutation status? The PROMISE clinical-genomic platform can help to generate initial data on potential positive and negative predictive biomarkers to guide these types of therapeutic questions.
Novel treatment paradigms in mHSPC and impact on subsequent treatment options
As novel treatment options are also introduced earlier in the disease course of prostate cancer, such as novel androgen receptor signaling inhibitors (ARSIs) [34,35,36,37], the natural history of many patients with mCRPC progressing on these therapies earlier in the disease course may also be altered. Do patients who develop castrate-resistant disease while being treated with ARSIs in the hormone-sensitive space subsequently have more aggressive and rapidly progressing CRPC? Do these patients subsequently respond to the standard of care treatment options available for mCRPC similarly to prior populations of patients not treated with these agents in the hormone-sensitive setting? Are these patients more likely to develop treatment-emergent neuroendocrine or small cell prostate cancer [38]? It is incumbent on the prostate cancer research community to better understand this natural history in light of new treatment paradigms. In this dynamic and rapidly changing treatment space much of the available clinical trial data used to guide treatment decisions may reflect a very different patient population than the patients being treated now. Serial cell-free DNA (cfDNA) is a cutting-edge solution to temporally understand therapeutic resistance mechanisms and is expected to be widely adopted in the future. Therefore, the vast genomic data gathered for each patient will be invaluable in our efforts to answer questions related to treatment exposure and acquisition of resistance.
Natural history and treatment options for novel molecular subtypes in prostate cancer
As treatment for prostate cancer is increasingly individualized and new biological subsets are identified through increased use of NGS, it is also important to better define the natural history of these patient populations. The molecular characterization of exceptional responders and non-responders to standard of care therapies will help better define molecular predictive biomarkers. In addition, as of 2020 there are now two agents, olaparib and rucaparib, specifically approved for the treatment of mCRPC with HRR alterations [2, 3] as well as multiple clinical trial options for patients with advanced prostate cancer and various genomic alterations. This is particularly important since response to standard of care treatments will serve as a benchmark for the efficacy of novel therapies that specifically target these genomic alterations. These questions need to be answered in order for precision medicine to truly fulfill its promise. More individualized treatment, implying the division of the broad diagnosis of prostate cancer into ever smaller molecularly defined subsets, will increasingly help guide the rational design of clinical trials and therapy selection. One early example of this approach is the IND.234 trial (NCT03385655) from the Canadian Clinical Trials group that uses a cfDNA platform to select therapy in prostate cancer. The improved understanding of these molecularly defined subsets through hypothesis-generating retrospective studies can help improve the design of biomarker-driven clinical trials of prostate cancer.
Impact of race/ethnicity or socioeconomic status on prostate cancer treatments and outcomes
Many studies focusing on characterizing the molecular alterations in prostate cancer have included a very limited number of ethnically diverse subsets of patients. Understanding how race/ethnicity impacts genomic profile and response to therapy is critical to bridging the disparities gap in men with prostate cancer. Real-world data from this platform can fill this gap. Black men remain underrepresented in phase 3 trials in prostate cancer despite being disproportionately impacted by lethal prostate cancer [39] and despite evidence of perhaps superior outcomes when treated with sipuleucel-T or docetaxel [40, 41]. Do these factors impact the diagnostic decisions made, such as the choice of imaging in advanced prostate cancer [42]? Are certain treatment options chosen preferentially over others and does this create healthcare inequities? Among the treatment options chosen, do certain options work better for certain subsets of patients? Such questions are difficult to answer with clinical trials which historically underrepresent many of the populations in question [43]. However, it is absolutely vital to answer these questions in order to help us improve the care of all patients with advanced prostate cancer. The collection of germline data for the patients in a real-world dataset will also help to both understand the landscape of somatic and germline mutations in underrepresented patients and the general availability of germline testing in this patient population [44, 45].
Utility of real-world data and the unmet need for the PROMISE consortium
All of the above clinical questions and scenarios in advanced prostate cancer illustrate the importance and potential utility of a large and well-maintained real-world database that serves as a repository for clinical and genomic data of prostate cancer patients. There are several pertinent reasons necessitating such a database as an important clinical research tool in advanced prostate cancer.
Prospective randomized clinical trials remain the gold standard in clinical cancer research and shape the current standard of care. While prospective clinical trials capture data on efficacy, observational studies can help to better understand effectiveness of the treatments in the real-world setting. Importantly, most prostate cancer patients do not get their cancer treatment as part of a clinical trial. A recent study estimated that only about 8% of oncology patients enroll in a clinical trial [46]. Consequently, the experiences and treatment outcomes of most prostate cancer patients are not systematically recorded or analyzed. In addition, most clinical trial datasets do not capture the entirety of patient experience from diagnosis until final treatments, focusing rather on one specific treatment outcome, and thus provide only a limited window into the outcomes of patients in the real-world setting. Retrospective studies of real-world patient datasets that include well-annotated clinical and genomic data can shed light on important questions that may be difficult to address in prospective clinical trials.
The inclusion of underrepresented minorities in this dataset is one of the important missions of this consortium. Data from retrospective series can more extensively capture clinical outcomes and other information of more diverse populations, including minority populations underrepresented in clinical trials. The low participation rates of patients who identify as underrepresented minorities in clinical trials of prostate cancer have been well documented and remain a significant challenge in clinical research and the conduct of trials [43]. The reasons for this are complex and multifactorial but it remains a significant challenge that is yet to be addressed. The selection for this consortium of diverse clinical sites representing different geographic regions of the United States that serve unique underrepresented patient populations is certainly an important step toward bridging the cancer genomics disparities gap in prostate cancer. Many of the initially proposed projects focus on questions related to racial disparities.
It is also important to keep in mind that although randomized clinical trials remain the gold standard for generating clinical data, many relevant clinical questions are initially asked in hypothesis-generating retrospective studies. Important descriptive data of the natural history of disease in these specific subgroups can be obtained through retrospective chart review. Many important prognostic and predictive markers and biomarkers of interest can initially be identified through analyses of real-world data and are subsequently validated prospectively. Furthermore, in contrast to prospective clinical trials where treatment options are prespecified, retrospective observational studies can also attempt to capture clinical decision-making, such as decisions made based on the results of genomic testing. In databases with large numbers of patients and well-annotated clinical and genomic data, important insights can be gleaned from this approach.
As treatments are increasingly individualized for molecularly selected subsets of patients with mCRPC, the careful collection of genomic data in large real-world cohorts can also help shed light on tumor biology and help advance questions that produce important validation studies. The availability of multiple tissue or blood samples from the same patient can also be particularly instructive. This information can help better define the evolution of molecular alterations or other biomarkers in the tumor throughout the course of disease progression. The understanding of such changes obtained from serial sampling of either repeat solid biopsies or of repeated cfDNA sampling can also help define the changes in the tumor that happen during progression on specific treatments. Further important comparisons can be drawn in the molecular alterations and other biomarkers detected in solid biopsies and cfDNA in the same patient.
Overall, our long-term goal is to develop a large real-world dataset collected from institutions including both academic and large community-based hospital systems, which by definition will be both heterogeneous and also more representative of real-world trends. Clinical and genomic information from many patients who are unable or unwilling to be treated on a clinical trial due to clinical, logistical or other reasons will also be included. This will make potential findings more generalizable relative to the data obtained from carefully selected clinical trial populations. The patient population included in a real-world dataset will also reflect the diversity of treatment options and strategies employed, diagnostic and genomic testing used, and the diverse patient populations across the different geographic regions.
Development of PROMISE, its aims, and the inclusion of a dedicated bio-informatics committee
The PROMISE consortium was formed with the recognition of the significant clinical and research needs of linking clinical and genomic data to outcomes, in order to better inform treatment decisions and outcomes for patients with advanced prostate cancer. Data collected in this consortium will address significant knowledge gaps that come about as the underlying biology of prostate cancer becomes better understood and multiple novel agents and classes of drugs emerge as treatment options. The structure of the PROMISE consortium including all of the committees and the flow chart of data analysis are shown in Fig. 2.
A Structure consisting of administrative headquarters, bio-informatics committee, executive committee, data committee, diversity & inclusion committee, and patient advocacy committee. Responsibilities of each committed listed. B Project workflow from data collection to release of data in abstract/manuscript form.
PROMISE is currently a consortium of 16 academic institutions with the common goal of understanding the real-world data in patients with advanced prostate cancer. The goal is to expand the reach of this database to include academic institutions and community practices. The consortium has two main aims:
(1) To establish a large, diverse, inclusive, and well-annotated repository of completely de-identified clinical and genomic patient data that is linked to clinical characteristics and disease-related outcomes.
(2) To investigate prognostic and potentially predictive biomarkers in patients with advanced prostate cancer treated with approved therapies including genomic, transcriptomic, and proteomic biomarkers.
Inclusion criteria for patients in this dataset consist of having advanced prostate cancer with either mHSPC or castrate-resistant prostate cancer (CRPC). In addition, patients should have undergone prior germline and/or somatic tumor molecular profiling studies including, but not limited to, one or more of the following blood or tissue-based assays: tumor DNA sequencing (e.g., cell-free circulating tumor DNA or tumor tissue sequencing), germline DNA sequencing and transcript profiling studies (e.g., RNA-seq, qRT-PCR). In total, consecutive patients with available clinical and genomic data at each site will be included to limit selection bias. Clinical and treatment data is collected from the time of initial prostate cancer diagnosis (Table 1). All data are maintained in a secure RedCap database and is completely deidentified without any protected health information. Genomic information entered into the database from both solid tissue and liquid biopsy testing and inclusive of both somatic and germline alterations are formatted to standardized criteria and reviewed by the bio-informatics committee. We will only include the entries that have all available data related to clinical outcomes and genomic sequencing.
PROMISE consortium institutions, data acquisition plan, and mission statement
The PROMISE consortium participating institutions are listed in Table 2. The consortium was formed in the first half of 2020 and data entry to the consortium has since risen steadily as sites have come onboard and over 1000 patient records are anticipated by early 2021 (Fig. 3). Genomic data are being audited for accuracy at each institution.
The mission of the PROMISE consortium is to investigate the outcomes of patients with advanced prostate cancer who have genomic and molecular profiling information and therefore to bridge the knowledge gap between real-world genomic data and clinical outcomes in patients with advanced prostate cancer. Consortium structure will also provide significant opportunities for mentorship of junior investigators and trainees. The consortium will plan to pursue multiple projects addressing specific questions which are proposed by the participating site investigators and selected through scientific review by the executive committee with priority given to projects proposed by junior investigators and trainees. Given the important potential contribution of evaluating outcomes in a more diverse population than that represented in clinical trials, a subcommittee for diversity and inclusion has been created with a plan to implement strategies to increase diversity if the database fails to maintain a balanced population.
Future directions
In a short period of time, we have gathered vast amounts of clinico-genomic information to answer important clinical questions. We are looking forward to expand our database to include many more academic institutions and community practices to make our patient population more representative of the real-world setting. In addition, once the database becomes robust for large-scale analyses, these data may be made publicly available for future research, following efforts such as cBioportal. Lastly, there is a long-term plan to develop infrastructure for blood- and tissue biorepository to perform molecular analyses.
Conclusions
Precision medicine holds significant promise for helping to individualize treatment approaches for patients with advanced prostate cancer and to help understand the underlying mechanisms of this heterogeneous disease. The PROMISE consortium is an important new collaboration among leading academic centers in the United States that aims to bridge the gap between clinical and genomic information for patients with metastatic prostate cancer and available genomic data from commercial and institutional assays. The well-annotated, de-identified patient data collected for this consortium will be leveraged to help answer specific questions and guide projects proposed by the consortium members. Genomic data entered into the database will be vetted by a bioinformatician and individual projects approved by the executive committee as part of the consortium structure. This organized approach will help to address important questions at the intersection of clinical and genomic data that are most optimally addressed using a real-world dataset. It is our hope that this approach will eventually help to improve outcomes for patients with advanced prostate cancer.
References
Beer TM, Armstrong AJ, Rathkopf D, Loriot Y, Sternberg CN, Higano CS, et al. Enzalutamide in men with chemotherapy-naïve metastatic castration-resistant prostate cancer: extended analysis of the phase 3 PREVAIL study. Eur Urol. 2017;71:151–4.
de Bono J, Mateo J, Fizazi K, Saad F, Shore N, Sandhu S, et al. Olaparib for metastatic castration-resistant prostate cancer. N Engl J Med. 2020;382:2091–102.
Abida W, Patnaik A, Campbell D, Shapiro J, Bryce AH, McDermott R, et al. Rucaparib in men with metastatic castration-resistant prostate cancer harboring a BRCA1 or BRCA2 gene alteration. J Clin Oncol. 2020;38:3763–72.
Cheng HH, Sokolova AO, Schaeffer EM, Small EJ, Higano CS. Germline and somatic mutations in prostate cancer for the clinician. J Natl Compr Canc Netw. 2019;17:515–21.
Pritchard CC, Mateo J, Walsh MF, De Sarkar N, Abida W, Beltran H, et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N Engl J Med. 2016;375:443–53.
Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161:1215–28.
Stopsack KH, Nandakumar S, Wibmer AG, Haywood S, Weg ES, Barnett ES, et al. Oncogenic genomic alterations, clinical phenotypes, and outcomes in metastatic castration-sensitive prostate cancer. Clin Cancer Res. 2020;26:3230–8.
Abida W, Cyrta J, Heller G, Prandi D, Armenia J, Coleman I, et al. Genomic correlates of clinical outcome in advanced prostate cancer. Proc Natl Acad Sci USA. 2019;116:11428–36.
Quigley DA, Dang HX, Zhao SG, Lloyd P, Aggarwal R, Alumkal JJ, et al. Genomic hallmarks and structural variation in metastatic prostate. Cancer Cell. 2018;174:758–69.e9.
Cancer Genome Atlas Research Network. The Molecular Taxonomy of Primary Prostate Cancer. Cell. 2015;163:1011–25.
Viswanathan SR, Ha G, Hoff AM, Wala JA, Carrot-Zhang J, Whelan CW, et al. Structural alterations driving castration-resistant prostate cancer revealed by linked-read genome sequencing. Cell. 2018;174:433–47.e19.
Mota JM, Barnett E, Nauseef JT, Nguyen B, Stopsack KH, Wibmer A, et al. Platinum-based chemotherapy in metastatic prostate cancer with DNA repair gene alterations. JCO Precis Oncol. 2020;4:355–66.
Cheng HH, Pritchard CC, Boyd T, Nelson PS, Montgomery B. Biallelic inactivation of BRCA2 in platinum-sensitive metastatic castration-resistant prostate cancer. Eur Urol. 2016;69:992–5.
Zafeiriou Z, Bianchini D, Chandler R, Rescigno P, Yuan W, Carreira S, et al. Genomic analysis of three metastatic prostate cancer patients with exceptional responses to carboplatin indicating different types of DNA repair deficiency. Eur Urol. 2019;75:184–92.
Wu YM, Cieślik M, Lonigro RJ, Vats P, Reimers MA, Cao X, et al. Inactivation of CDK12 delineates a distinct immunogenic class of advanced prostate. Cancer Cell. 2018;173:1770–82.e14.
Nguyen B, Mota JM, Nandakumar S, Stopsack KH, Weg E, Rathkopf D, et al. Pan-cancer analysis of <em>CDK12</em> alterations identifies a subset of prostate cancers with distinct genomic and clinical characteristics. Eur Urol. 2020;78:671–9.
Schweizer MT, Ha G, Gulati R, Brown LC, McKay RR, Dorff T, et al. CDK12-mutated prostate cancer: clinical outcomes with standard therapies and immune checkpoint blockade. JCO Precis Oncol. 2020;4:382–92.
George DJ, Halabi S, Healy P, Jonasch D, Anand M, Rasmussen J, et al. Phase 2 clinical trial of TORC1 inhibition with everolimus in men with metastatic castration-resistant prostate cancer. Urol Oncol. 2020;38:79.e15–79.e22.
Crumbaker M, Khoja L, Joshua AM. AR signaling and the PI3K pathway in prostate cancer. Cancers. 2017;9:34.
de Bono JS, De Giorgi U, Rodrigues DN, Massard C, Bracarda S, Font A, et al. Randomized Phase II study evaluating Akt blockade with ipatasertib, in combination with abiraterone, in patients with metastatic prostate cancer with and without PTEN loss. Clin Cancer Res. 2019;25:928–36.
de Bono JS, Bracarda S, Sternberg CN, Chi KN, Olmos D, Sandhu S, et al. LBA4 IPATential150: Phase III study of ipatasertib (ipat) plus abiraterone (abi) vs placebo (pbo) plus abi in metastatic castration-resistant prostate cancer (mCRPC). Ann Oncol. 2020;31:S1153–S4.
Hyman DM, Smyth LM, Donoghue MTA, Westin SN, Bedard PL, Dean EJ, et al. AKT inhibition in solid tumors with AKT1 mutations. J Clin Oncol. 2017;35:2251–9.
André F, Ciruelos E, Rubovszky G, Campone M, Loibl S, Rugo HS, et al. Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N Engl J Med. 2019;380:1929–40.
Nava Rodrigues D, Rescigno P, Liu D, Yuan W, Carreira S, Lambros MB, et al. Immunogenomic analyses associate immunological alterations with mismatch repair defects in prostate cancer. J Clin Investig. 2018;128:4441–53.
Graham LS, Montgomery B, Cheng HH, Yu EY, Nelson PS, Pritchard C, et al. Mismatch repair deficiency in metastatic prostate cancer: response to PD-1 blockade and standard therapies. PLoS ONE. 2020;15:e0233260.
Ritch E, Fu SYF, Herberts C, Wang G, Warner EW, Schönlau E, et al. Identification of hypermutation and defective mismatch repair in ctDNA from metastatic prostate cancer. Clin Cancer Res. 2020;26:1114–25.
Antonarakis ES, Shaukat F, Isaacsson Velho P, Kaur H, Shenderov E, Pardoll DM, et al. Clinical features and therapeutic outcomes in men with advanced prostate cancer and DNA mismatch repair gene mutations. Eur Urol. 2019;75:378–82.
Abida W, Cheng ML, Armenia J, Middha S, Autio KA, Vargas HA, et al. Analysis of the prevalence of microsatellite instability in prostate cancer and response to immune checkpoint blockade. JAMA Oncol. 2019;5:471–8.
Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science. 2017;357:409–13.
Marcus L, Lemery SJ, Keegan P, Pazdur R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin Cancer Res. 2019;25:3753–8.
Marabelle A, Fakih M, Lopez J, Shah M, Shapira-Frommer R, Nakagawa K, et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 2020;21:1353–65.
Hofman MS, Emmett L, Violet J, AYZ, Lawrence NJ, Stockler M, et al. TheraP: a randomized phase 2 trial of (177) Lu-PSMA-617 theranostic treatment vs cabazitaxel in progressive metastatic castration-resistant prostate cancer (Clinical Trial Protocol ANZUP 1603). BJU Int. 2019;124 Suppl 1:5–13.
Hofman MS, Violet J, Hicks RJ, Ferdinandus J, Thang SP, Akhurst T, et al. [(177)Lu]-PSMA-617 radionuclide treatment in patients with metastatic castration-resistant prostate cancer (LuPSMA trial): a single-centre, single-arm, phase 2 study. Lancet Oncol. 2018;19:825–33.
Armstrong AJ, Szmulewitz RZ, Petrylak DP, Holzbeierlein J, Villers A, Azad A, et al. ARCHES: a randomized, phase III study of androgen deprivation therapy with enzalutamide or placebo in men with metastatic hormone-sensitive prostate cancer. J Clin Oncol. 2019;37:2974–86.
Chi KN, Agarwal N, Bjartell A, Chung BH, Pereira de Santana Gomes AJ, Given R, et al. Apalutamide for metastatic, castration-sensitive prostate cancer. N Engl J Med. 2019;381:13–24.
Fizazi K, Tran N, Fein L, Matsubara N, Rodriguez-Antolin A, Alekseev BY, et al. Abiraterone plus prednisone in metastatic, castration-sensitive prostate cancer. N Engl J Med. 2017;377:352–60.
James ND, de Bono JS, Spears MR, Clarke NW, Mason MD, Dearnaley DP, et al. Abiraterone for prostate cancer not previously treated with hormone therapy. N Engl J Med. 2017;377:338–51.
Aggarwal R, Huang J, Alumkal JJ, Zhang L, Feng FY, Thomas GV, et al. Clinical and genomic characterization of treatment-emergent small-cell neuroendocrine prostate cancer: a multi-institutional prospective study. J Clin Oncol. 2018;36:2492–503.
Spratt DE, Chen YW, Mahal BA, Osborne JR, Zhao SG, Morgan TM, et al. Individual patient data analysis of randomized clinical trials: impact of black race on castration-resistant prostate cancer outcomes. Eur Urol Focus. 2016;2:532–9.
Sartor O, Armstrong AJ, Ahaghotu C, McLeod DG, Cooperberg MR, Penson DF, et al. Survival of African-American and Caucasian men after sipuleucel-T immunotherapy: outcomes from the PROCEED registry. Prostate Cancer Prostatic Dis. 2020;23:517–26.
Halabi S, Dutta S, Tangen CM, Rosenthal M, Petrylak DP, Thompson IM Jr., et al. Overall survival of Black and White men with metastatic castration-resistant prostate cancer treated with docetaxel. J Clin Oncol. 2019;37:403–10.
Bucknor MD, Lichtensztajn DY, Lin TK, Borno HT, Gomez SL, Hope TA. Disparities in PET imaging for prostate cancer at a tertiary academic medical center. J Nucl Med. 2020;62:695–9.
Balakrishnan AS, Palmer NR, Fergus KB, Gaither TW, Baradaran N, Ndoye M, et al. Minority recruitment trends in phase III prostate cancer clinical trials (2003 to 2014): progress and critical areas for improvement. J Urol. 2019;201:259–67.
Kwon DH, Borno HT, Cheng HH, Zhou AY, Small EJ. Ethnic disparities among men with prostate cancer undergoing germline testing. Urol Oncol. 2020;38:80.e1–e7.
Mahal BA, Alshalalfa M, Kensler KH, Chowdhury-Paulino I, Kantoff P, Mucci LA, et al. Racial differences in genomic profiling of prostate cancer. N Engl J Med. 2020;383:1083–5.
Unger JM, Vaidya R, Hershman DL, Minasian LM, Fleury ME. Systematic review and meta-analysis of the magnitude of structural, clinical, and physician and patient barriers to cancer clinical trial participation. J Natl Cancer Inst. 2019;111:245–55.
Author contributions
VK, VP, AA, and RM article conceptualization; all authors including VK, VP, AA, and RM manuscript writing.
Funding
Currently, each site principal investigator is funding their own efforts for the consortium. In the future, we will be evaluating outside funding opportunities from organizations such as the Prostate Cancer Foundation and industry sources.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
VSK received research funding to his institution from Janssen, Nektar, Endocyte, Clovis. He has received compensation as a member of the scientific advisory board of Janssen, Clovis, Pfizer, Seattle Genetics/Astellas, Dendreon, AstraZeneca, EMD Serono. All funding received is for work performed outside the scope of this study. MAB received research funding to his institution from Xencor, Bayer, Bristol-Myers Squibb, Genentech/Roche, Seattle Genetics, Incyte, Nektar, AstraZeneca, Tricon Pharmaceuticals, Genome & Company, AAA, Peloton Therapeutics, and Pfizer. He has received compensation as a member of the scientific advisory board or as paid consultant of Exelixis, Bayer, BMS, Eisai, Pfizer, AstraZeneca, Janssen, Calithera Biosciences, Genomic Health, Nektar, and Sanofi. All funding received is for work performed outside the scope of this study. CP has consulted for AstraZaneca and received compensation for work performed outside the scope of this study. AT received research funding to his institution from EMD Serono, Aravive Inc, and Wndmil therapeutics. He has received compensation as a member of the scientific advisory board of Exilixis, EMD Serono, Genzyme, Foundation, and Pfizer. All funding received is for work performed outside the scope of this study. PB received research funding to his institution from AstraZeneca, Merck, Blue Earth Diagnostics. He has received compensation as a member of the scientific advisory board of Bristol Myers Squibb, Exelexis, Janseen, Pfizer, Sanofi, Dendreon, Caris, Bayer, EMD Serono. All funding received is for work performed outside the scope of this study. CH received research funding to his institution from AstraZeneca, Bayer, Dendreon, Merck. She has stock ownership in Johnson & Johnson. All funding received is for work performed outside the scope of this study. WO is an employee of Sema4 as Chief Medical Science Officer. He has consulted for and received funding from Amgen, Astellas, Bayer, AstraZeneca, Genzyme, Janssen, and Pfizer. All funding received is for work performed outside the scope of this study. EH received research funding to her institution from Caris. She has consulted for and received compensation from Caris. All funding received is for work performed outside the scope of this study. CHM is an employee of McGraw Hill. She has consulted for and received compensation from Dendreon and Bayer. All funding received is for work performed outside the scope of this study. STT received research funding to his institution from Sanofi, Medivation, Astellas, Janssen, rogenics, Dendreon, Newlink, Inovio, Immunomedics, Atlab, Boehringer Ingelheim, Millennium, Bayer, Merck, Abbvie, Karyopharm, Endocyte, Clovis, and Novartis. He has received personal honoraria from Sanofi, Medivation/Astellas, Dendreon, Janssen, Genentech, Bayer, Endocyte, Eisai, Immunomedics, Karyopharm, Abbvie, Tolmar, Seattle Genetics, Amgen, Clovis, QED, Pfizer, AAA/Novartis, Genomic Health, POINT Biopharma, Blue Earth Diagnostics. All funding received is for work performed outside the scope of this study. MTS received research funding to his institution from Zenith Epigenetics, Bristol Myers Squibb, Merck, Immunomedics, Janssen, AstraZeneca, Pfizer, Madison Vaccines, Tmunity and Hoffman-La Roche. He has consulted for and received compensation from Janssen and Resverlogix. All funding received is for work performed outside the scope of this study. A Armstrong received research funding to his institution from Merck, Bayer, Pfizer, Astellas, Janssen, Genentech/Roche, Constellation, Beigene, BMS, AstraZeneca. He consulted for and received compensation from Merck, Bayer, Pfizer, Astellas, Janssen, AstraZeneca, and Clovis. All funding received is for work performed outside the scope of this study. A Alva received research funding to his institution from MSD, AstraZeneca, Bristol-Myers Squibb, Astellas, Seattle Genetics, Genentech, Pfizer, Progenics, Prometheus, Eli Lilly, ASCO, Celgene, and Harpoon Therapeutics. He has received compensation as a member of the advisory board and as paid consultant for MSD, Bristol-Myers Squibb, AstraZeneca, Genentech, Roche, Pfizer, Progenics, and Prometheus. All funding received is for work performed outside the scope of this study. RM received research funding to her institution from Bayer, Pfizer, and Tempus. She has received compensation as a member of the advisory board for AstraZeneca, Bayer, Bristol Myers Squibb, Calithera, Exelixis, Janssen, Merck, Novartis, Pfizer, Sanofi, and Tempus. She consulted for and received compensation for Dendreon and Vividion. She serves on the molecular tumor board at Caris. All funding received is for work performed outside the scope of this study. VGP, A Ali, DR, JJP, OK, MC, JS, AC, LB, ML, LSG, SN, AJ, SRC, RG, LA, AP, TJ, DN, CN, DK, MP, BT, FC, AM, SH, and TD have no competing interests to disclose.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koshkin, V.S., Patel, V.G., Ali, A. et al. PROMISE: a real-world clinical-genomic database to address knowledge gaps in prostate cancer. Prostate Cancer Prostatic Dis 25, 388–396 (2022). https://doi.org/10.1038/s41391-021-00433-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41391-021-00433-1
This article is cited by
-
Clinical implications of AR alterations in advanced prostate cancer: a multi-institutional collaboration
Prostate Cancer and Prostatic Diseases (2024)
-
Identification of Chemical–Disease Associations Through Integration of Molecular Fingerprint, Gene Ontology and Pathway Information
Interdisciplinary Sciences: Computational Life Sciences (2022)