Abstract
Background
Urinary tract obstruction is a common cause of renal failure in children and infants, and the pathophysiological mechanisms of obstructive nephropathy are largely unclear. It has been reported that m6A modulation is involved in renal injury. However, whether m6A RNA modulation is associated with obstructive nephropathy has not yet been reported. The aim of this study was to investigate the m6A epitranscriptome profiles in the kidneys of bilateral ureteral obstruction (BUO) in young rats.
Methods
The total level of m6A in the kidneys was measured by liquid chromatography-tandem mass spectrometry. The mRNAs of related genes were detected by real-time PCR. Methylated RNA immunoprecipitation sequencing was performed to map the epitranscriptome-wide m6A profile.
Results
Global m6A levels were increased after BUO, and the mRNA expression levels of m6A methyltransferases and demethylases were significantly decreased in BUO group rat kidneys; the expression levels of EGFR and Brcal were significantly upregulated, while the mRNA expression levels of Notch1 were downregulated (P < 0.05). A total of 154 genes associated with 163 m6A peaks were identified.
Conclusion
The m6A epitranscriptome was significantly altered in BUO rat kidneys, which is potentially implicated in the pathophysiological processes of obstructive nephropathy.
Impact
-
The m6A RNA modification was associated with the process of renal injury in ureteral obstructive nephropathy by participating in multiple dimensions.
-
The dysregulation of m6A methyltransferases and demethylases may be related to the pathophysiological changes of BUO-induced obstructive nephropathy.
-
The m6A RNA modulation of the genes EGFR, Brca1, and Notch1 that were related to the regulation of aquaporin2 might be the potential mechanism for the polyuresis after ureteral obstruction.
Similar content being viewed by others
Introduction
Ureteral obstruction-induced congenital hydronephrosis is a common cause of renal failure in children and infants.1 It has been reported that developmental urinary tract and renal abnormalities are responsible for approximately 54% of chronic renal insufficiency cases, and most of the developmental abnormalities are congenital obstructive nephropathies.2 Even though this disease has serious impacts on patients and society, the specific pathophysiological mechanisms are largely not yet clear.
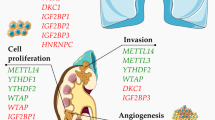
It has been demonstrated that epigenetic regulation, like histone modification and noncoding RNAs, could affect chronic kidney disease and acute kidney injury.3,4 However, the epitranscriptomic modifications of mRNA in the pathological process of bilateral ureteral obstruction (BUO)-induced obstructive nephropathy are largely unknown. N6-methyladenosine (m6A) is the most prevailing internal chemical modification of eukaryotic mRNAs.3,4 The m6A modifications are jointly regulated by the “writer”(e.g., WTAP, METTL14, and METTLl3), “eraser” (e.g., ALKBH5 and FTO), and “reader” (e.g., YTHDF, IGF2BPs, eIF3, and HNRNP).5 Previous studies have demonstrated that the dysregulation of m6A modulation is associated with a number of diseases, such as infertility, obesity, diabetes, etc.6 Recently, it has also been found that the dysregulated m6A in mRNA is associated with cardiac diseases, and the enhanced or diminished m6A RNA methylation could result in cardiac dysfunction.7 These results suggest the possibility of exploring innovative therapeutic methods through the adjustment of m6A in mRNA.
In the realm of renal diseases, it has been demonstrated that the m6A demethylase FTO is relevant to end-stage renal disease8 and can predict the fatality of chronic kidney diseases.9 Li et al. also found that m6A modification played a crucial part in obstructive nephropathy-induced renal interstitial fibrosis.10 In another study, Xu et al. found that METTL14 could promote renal ischemia–reperfusion injury by weakening YAP1-TEAD signaling.11 However, the relationship between m6A modification and BUO-induced renal damage is still not clear. To bridge the gap in the literature, in this study, we examined the relationship between m6A modification of RNAs and obstructive nephropathy.
Materials and methods
All procedures conformed to the Chinese National Guidelines for the Care and Handling of Animals and the published guidelines from the National Institutes of Clinical Medicine, Zhengzhou University, according to the licenses for the use of experimental animals issued by the Chinese Ministry of Justice (2022-KY-0046-002). Studies were performed on male (Sprague–Dawley, SD) rats, initially weighing 160–180 g (Henan Experimental Animal Center, Henan, China). The animals were maintained on a standard rodent diet and had free access to water. During the entire experimental period, rats were kept in individual metabolic cages with a 12/12-h artificial light/dark cycle, a temperature of 21 ± 2 °C, and a humidity of 55 ± 2%.
After 3 days of acclimation to the metabolic cages, experimental BUO models were induced by occlusion of both ureters for 24 h. During surgery, experimental rats were anesthetized with inhaled isoflurane (Sigma-Aldrich) and placed on a heated table to maintain the rectal temperature at 37–38 °C. An abdominal median incision exposed the bilateral ureters, and a 5-mm-long piece of bisected polyethylene tube (PE-50) was placed around the importation of each dissected ureter. The ureters were then occluded by tightening the tube with a 5-0 silk ligature. After 24 h, the obstructed ureters were decompressed. This technique completely occludes the ureters for 24 h without evidence of subsequent functional impairment of ureteral function.12 After release of the ureteral obstruction, all rats were followed for 24 h for urine collection and then sacrificed.
As a sham group, rats were subjected to sham operations identical to those used for BUO rats without occlusion of both ureters. Sham-operated rats were monitored in parallel with BUO rats. All rats were sacrificed under light isoflurane anesthesia, during which time the kidneys were rapidly removed, and the plasma was collected from the abdominal aorta.
Rats were divided into the BUO group (n = 8) and the sham group (n = 8).
Clearance studies
The rats were maintained in metabolic cages, allowing quantitative urine collections and measurements of water intake. Urine volume and osmolality were measured. Plasma was used for the measurement of osmolality, creatinine, and urea nitrogen.
Histopathological evaluation
Kidney samples were fixed with 4% paraformaldehyde buffer, washed for 3 × 10 min with phosphate-buffered saline (PBS), and then dehydrated and embedded in wax. The paraffin-embedded tissues were cut into 4-μm sections on a rotary microtome (Leica, RM2016). The sections were dewaxed and rehydrated. The slides were stained with hematoxylin and eosin and evaluated with a Leica light microscope.
Triple staining
Kidneys were fixed with 4% paraformaldehyde buffer, washed three times for 10 min each with PBS buffer, dehydrated, and embedded in wax. The paraffin-embedded tissues were cut into 8-μm sections via a rotary microtome (RM2016, Leica). The sections were dewaxed and rehydrated. To reveal the antigens, the sections were incubated in 1 mmol/L Tris solution (pH 9.0) supplemented with 0.5 mM ethylenediaminetetraacetic acid (EDTA) and heated in a microwave oven for 15 min. Endogenous peroxidase was blocked by 3% H2O2 for 25 min at room temperature. Nonspecific binding of IgG was prevented by incubating the sections in 50 mM NH4Cl for 30 min, followed by blocking in PBS supplemented with 2% bovine serum albumin. The sections were incubated overnight at 4 °C with rabbit anti-aquaporin2 (AQP2) (GB112259, Servicebio, China) diluted in PBS. Then the sections were incubated overnight, after which the sections were incubated with horseradish peroxidase-conjugated secondary antibodies (G23303, goat anti-rabbit immunoglobulin, Servicebio) for 50 min at room temperature. After rinsing with PBS washing buffer, the sites were incubated for 10 min withCY3-TSA. Then the sites were incubated in EDTA and heated in a microwave oven as described above. Then the sections were incubated overnight at 4 °C with rabbit anti-Atp6v1b1 (ab192612, Abcam, UK) diluted in PBS. The sections were cubated overnight, after which the sections were incubated with FITC-conjugated secondary antibodies (G22303, goat anti-rabbit immunoglobulin, Servicebio) for 50 min at room temperature. After rinsing with PBS washing buffer, the sites of antibody-antigen reactions were visualized with 0.05% 3,3’-diaminobenzidine dissolved in distilled water supplemented with 0.1% H2O2. Light microscopy was carried out with a Leica microscope.
Quantitative real-time PCR
Total RNA was isolated and purified with TRIzol reagent (Invitrogen, Carlsbad, CA). RNA was quantitated using a spectrophotometer (ND-1000 V3.5.2 software). cDNA was synthesized using a High Capacity cDNA Reverse Transcription kit (CW BIOTECH, China), and quantitative real-time PCR was performed with the Applied Biosystems 7500 Sequence Detection System, as previously described.13 The primer information is listed in Table 1.
RNA-seq
Total RNA extraction and DNA digestion were performed as described above. Qualified RNAs were finally quantified by Qubit3.0 with a QubitTM RNA Broad Range Assay kit (Life Technologies, Q10210). Two micrograms of total RNA were used for stranded RNA sequencing library preparation using a KC-DigitalTM Stranded mRNA Library Prep Kit for Illumina® (Catalog NO. DR08502, Wuhan Seqhealth Co., Ltd. China) following the manufacturer’s instructions.
MeRIP-seq and data analysis
Total RNA was extracted as described above. Fifty micrograms of total RNA were used for polyadenylated RNA enrichment by VAHTS mRNA Capture Beads (VAHTS, cat. No. N401-01/02). Then, 20 mM ZnCl2 was added to mRNA and incubated at 95 °C for 5–10 min until the RNA fragments were mainly distributed in 100–200 nt. Then, 10% of the RNA fragments were saved as “input” data, and the remaining RNA fragments were used for m6A immunoprecipitation (IP). A specific anti-m6A antibody (Synaptic Systems, 202203) was applied for m6A IP. The stranded RNA sequencing library was constructed by a KC-DigitalTM Stranded mRNA Library Prep Kit for Illumina® (Catalog No. DR08502, Wuhan Seqhealth Co., Ltd. China) following the manufacturer’s instructions.
Data analysis
Raw sequencing data were first filtered by Trimmomatic (version 0.36), low-quality reads were discarded, and the reads contaminated with adaptor sequences were trimmed. Clean reads were further treated with in-house scripts to eliminate duplication bias introduced in library preparation and sequencing. ExomePeak (Version 3.8) software was used for peak calling. The m6A peaks were annotated using bedtools (Version 2.25.0). DeepTools (version 2.4.1) was used for peak distribution analysis. The differential m6A peaks were identified by a Python script using Fisher’s test. Sequence motifs enriched in m6A peak regions were identified using Homer (version 4.10). The m6A-methylated RNAs with |fold changes| ≥ 2 and P values ≤0.05 between the control and BUO groups were identified as having significantly differential expression. Gene Ontology (GO) is a bioinformatic method for annotating genes, gene products, and sequences to the underlying biological activities (Gene Ontology Consortium). Pathway analysis based on Kyoto Encyclopedia of Genes and Genomes (KEGG) was utilized to identify the pathways significantly enriched in the parental genes of the differentially expressed m6A-methylated RNAs.
Quantification of m6A in total RNA and mRNA by LC-MS-MS
Total RNA was isolated as described above. Adding buffer, S1 nuclease, Alkaline Phosphatase, and Phosphodiesterase I into 1 μg RNA, then the mixture was incubated at 37 °C. After the RNA was digested into nucleosides completely, the mixture was extracted with chloroform. The resulting aqueous layer was collected for analysis with LC-ESI-MS/MS. The sample extracts were analyzed using a UPLC-ESI-MS/MS system. The effluent was alternatively connected to an ESI-triple quadrupole-linear ion trap (QTRAP)-MS. Linear ion trap and triple quadrupole scans were acquired on a triple QTRAP, QTRAP® 6500+ LC-MS/MS System, equipped with an ESI Turbo Ion-Spray interface, operating in positive ion mode and controlled by Analyst 1.6.3 software. RNA modifications were analyzed using scheduled multiple reaction monitoring. Data acquisitions were performed using Analyst 1.6.3 software (Sciex). Multiquant 3.0.3 software (Sciex) was used to quantify all metabolites. Mass spectrometer parameters including the declustering potentials (DP) and collision energies (CE) for individual MRM transitions were done with further DP and CE optimization. A specific set of MRM transitions was monitored for each period according to the metabolites eluted within this period.
Statistical analysis
Statistical analyses were performed with SPSS version 21 (SPSS, Inc., IBM) and GraphPad Prism version 5.0 (GraphPad Software, San Diego). All the values are presented as the mean ± SD. Comparisons between two groups were conducted according to unpaired t-tests. P < 0.05 was considered significant.
Results
BUO-induced renal function insufficiency with impaired renal water handling
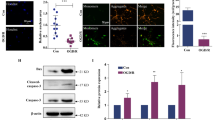
Rats treated with 24-h BUO showed a significant increase in plasma concentrations of urea and creatinine (Table 2). The urine osmolality was significantly lower than that in the sham group, and the urine volume was significantly higher than that in the sham group (Table 2).
HE staining and cellular composition of the collecting duct
HE staining showed that the renal tubules were flattened and that the interstitium was infiltrated with inflammatory cells (Fig. 1a, b). Immunofluorescence staining showed that the ratio of ATP6V1B1/AQP2, which are markers of intercalated cells (ICs) and principal cells (PCs), respectively, positive cells was higher in the BUO group than in the sham group (Fig. 1c, d).
BUO increased the total m6A level and modulated the gene expression of m6A-related enzymes in rat kidneys
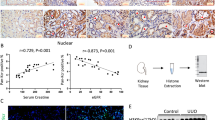
To reveal whether the observed renal damage was related to the modification of m6A after obstruction, the total m6A in mRNA and the mRNA levels of three methyltransferases (METTL3, METTL14, and ALKBH5) and two demethylases (FTO and WTAP) in the kidney were measured in the sham and BUO groups. As depicted in Fig. 2a, m6A was higher in the BUO group than in the Sham group, and the mRNA levels of the five examined enzymes were generally decreased in BUO rat kidneys compared with those of the sham group (P < 0.05) (Fig. 2b).
Overview of epitranscriptome profile and conjoint analysis of m6A-RIP-seq and RNA-seq data
In Fig. 3a, the volcano plots display significantly differentiated genes (BUO vs. sham). There were 2031 downregulated and 1401 upregulated genes (P < 0.05). The corresponding differentially expressed mRNAs between the BUO and sham groups are presented by hierarchical clustering in Fig. 3b. The volcano plots in Fig. 3c show that there were 220 peaks that were significantly hypomethylated and 315 peaks that were significantly hypermethylated for the m6A MeRIP-enriched regions. In total, there were 154 genes associated with 163 significantly differentially methylated transcripts of the renal m6A epitranscriptome. Detailed information on the top 20 significantly differentially methylated peaks after BUO is presented in Table 3. Further conjoint analysis of MeRIP-seq and RNA-seq data revealed 79 hypermethylated and 84 hypomethylated m6A peaks. For the hypermethylated m6A peaks, 61 transcripts were significantly downregulated, and 18 were upregulated; for the hypomethylated m6A peaks, 32 transcripts were significantly upregulated, and 52 were downregulated (Fig. 3d). Detailed information on the top three genes in each quadrant of Fig. 3d is listed in Table 4.
a Volcano plots showing the mRNAs that were differentially expressed between BUO and sham groups with significant difference (p < 0.05). b Renal clustering analysis of the differentially expressed mRNA (p < 0.05). c Volcano plots displaying the distinct m6A peaks (p < 0.05). d Four quadrant graph showing the distribution of transcripts with significant changes in both m6A-modified level and corresponding mRNA expression after 24-h BUO.
Distribution and pathway analysis of significantly differentially methylated genes
The m6A peaks were predominantly enriched in the CDS and 3’UTR regions (Fig. 4a, b). Enrichment analysis for the conserved motifs shared among the identified m6A peaks supported the m6A RRACH ([R:G/A][H: U/A/C]) consensus sequence (Fig. 4c). Figure 4d shows that genes corresponding to significantly differential m6A peaks were widely distributed among all chromosomes (chr), particularly chr1, chr2, and chr10 (Fig. 4d).
a Density curve shows the accumulation of differentially methylated m6A peaks of transcripts in each sample. Each transcript is divided into three parts including 5’UTR, 3’UTR, and CDS. b Percentage pile-up bar chart shows the distribution of these peaks in three parts. c The top motifs enriched across m6A peaks of the BUO group and the sham group. d Distribution of significantly altered methylated m6A sites with significance in chromosomes of rats in BUO group and sham group. 3’UTR 3’untranslated regions, CDS coding DNA sequence, 5’UTR 5’untranslated region.
GO and KEGG analyses to reveal the biological function of identified differentiated m6A genes
Figure 5a, c shows that the differentially methylated genes were associated with biological processes, the cell cycle, DNA metabolic processes and cellular processes, and the regulation of cell communication, signal transduction, and cell projection. KEGG pathway analyses showed that the differentially methylated genes were mainly significantly related to Notch signaling pathways (for upregulated genes) and steroid biosynthesis signaling pathways (for downregulated genes) (Fig. 5b, d).
Table 5 presents the information of our genes of interest, which are associated with the expression of AQP2 and water reabsorption. We further examined the mRNA expression of these genes with RT–PCR. The mRNA expression levels of EGF receptor (EGFR) and Brcal were significantly upregulated, while the mRNA expression levels of Notch1 were downregulated compared with those in the sham group (P < 0.05) (Table 5).
Discussion
Congenital ureteral obstruction-induced hydronephrosis is one of the most important challenges faced by pediatric urologists. However, the pathogenesis of this disease is largely not yet clear.14 Thus, it is observed that obstructive nephropathy is not a simple result of mechanical injury to urine flow but the consequence of a complex series of pathophysiological changes, including interstitial inflammatory cell infiltration, tubular atrophy, tubulointerstitial fibrosis, etc., which leads to the impairment of all renal functions.15,16,17 Lately, it has been exposed that m6A modifications in RNAs are associated with many diseases, like diabetes mellitus and coronary heart disease.18 However, the relationship between m6A RNA modulation and BUO-induced obstructive nephropathy is still unknown. According to the results of this research, the m6A expression pattern of the obstructive kidneys was different from that of the control group. First, the m6A level was higher than that in controls. In line with this result, the mRNA levels of FTO and ALKBH5 declined. Of greater interest is the inverse trend of the mRNA levels of METTL3, METTL14, and ALKBH5 compared with the change in total m6A levels. Moreover, there were a total of 618 significantly differentially regulated mRNAs, which might be implicated in multiple pathways and biological processes. These results may help us to understand the role of m6A RNA modification in BUO-induced obstructive nephropathy.
Increasing confirmation manifests that m6A modulation is associated with renal diseases. Recent research demonstrated that the m6A-methylated RNA levels of kidney biopsies from patients with AKI were significantly higher than those of kidney biopsies from patients without AKI.11 Liu et al. discovered that m6A modification played a crucial role in the process of renal fibrosis.10 Ramalingam et al. demonstrated that m6A modification followed the development of polycystic kidney disease.19 Taken together, these studies suggest that m6A methylation is involved in scores of renal diseases. In the present study, m6A methylation levels in total RNA in BUO rat kidneys were higher than that of controls and the expression patterns of m6A-modified mRNAs were significantly different from that of the control groups. Similar to our results, Li et al. found the expression pattern of renal m6A methylation changed significantly after ureteral obstruction.10 By virtue of the differences in the animal model (UUO vs. BUO) and experimental animal species (C57BL/6 mice vs. SD rats), the results in the two studies did not entirely correspond to each other. However, the conclusions are the same, confirming that m6A methylation may have a close relationship with the pathophysiologic process of obstructive nephropathy.
m6A modification is modulated by the “writer”, including WTAP, METTL3, and METTL14, and the “eraser”, including ALKBH5 and FTO.5 In the current study, FTO, and ALKBH5 were discovered to be downregulated in BUO rat kidneys, which was in parallel with increased m6A modification. However, the mRNA levels of WTAP, METTL3, and METTL14 were decreased in BUO rat kidneys, which was inconsistent with the trend of changes in the total m6A levels. Similarly, Li et al. found that METTL3 and WTAP protein levels were in parallel with the increased m6A modification in cisplatin-induced AKI.10 In contrast, the METTL14 protein levels show the opposite trend compared to that of total m6A levels. Taken together, these results suggest that the prosperity of m6A has partially merged with the expression level with the regulation of their methyltransferases and demethylases. Thus, our results indicated that the effects of the downregulation of the demethylases (FTO and ALKBH5) may transcend the effects of decreased methyltransferases (WTAP, METTL3, and METTL14) expression, thus jointly leading to the upregulation of m6A modification in BUO rats. However, the potential mechanisms are still not yet clear.
Previous studies have verified that the adenosine methyltransferase complex is associated with many biological processes, such as metabolism, the inflammatory response, cell apoptosis, the cell cycle, and cell proliferation.20 Harini Ramalingam and colleagues found that METTL3 activation promotes proliferative signaling and cyst growth in autosomal dominant polycystic kidney disease (ADPKD), and the deletion of METTL3 could improve ADPKD in mouse models.19 Instead, suppressing the expression of METTL3 reduced the apoptosis of RK-52E cells.21 Apart from METTL3, METTL14 was also found to be involved in the progression of many renal diseases. Li et al. found that METTL14 was highly expressed in the kidneys of diabetic nephropathy patients and high glucose-induced human renal glomerular endothelial cells (HRGECs). Overexpression of METTL14 increased apoptosis in HRGECs. In contrast, METTL14 silencing decreased cell apoptosis.22 Hence, the essential members of m6A methyltransferases METTL14 and METTL3 might be involved in the development of BUO-induced nephropathy.
In addition to the adenosine methyltransferase, the demethylases of m6A were also found to be connected with a variety of renal diseases. FTO was found to be an independent predictor of all-cause mortality in patients with CKD of various severities.23,24 As one of the demethylases of m6A, FTO can selectively remove the methyl group from target RNAs, which may be associated with the regulation of specific genes m6A modification. Yan et al. found that FTO-dependent m6A demethylation intensifies the mRNA stability of proliferation/survival transcripts bearing m6A and afterward could enhance cell survival via promoting the expression of antiapoptotic genes.22 Another study demonstrated that reduced FTO expression accelerated apoptosis in cisplatin-treated HK-2 cells; conversely, FTO protein overexpression inhibited apoptosis.4 It has also been found that FTO is increased in ureteral obstruction-induced renal fibrosis.10,25 In the present study, renal FTO was diminished after BUO, which was correlated with the increment of total m6A level in mRNA. Accordingly, FTO and the succeeding m6A modification might be concerned with the pathological process of BUO-induced obstructive nephropathy. ALKBH5, another primary m6A demethylase, has been demonstrated to play a pivotal role in many pathophysiological processes, including apoptosis, proliferation, and autophagy.26 Zhu and Lu found that ALKBH5 knockdown inhibited cell viability, induced cell apoptosis, and declined inflammatory cytokine production in HK-2 cells.27 Zhang et al. demonstrated that ALKBH5 could stimulate cell proliferation in renal cell carcinoma.28 Ning et al. also found that ALKBH5 was associated with UUO-induced renal fibrosis.29 Therefore, the dysregulation of m6A-related adenosine methyltransferase and demethylases may be involved in the pathophysiological processes of BUO-induced obstructive nephropathy.
In this research, we also found that many host genes for the m6A-modified transcripts were involved in many important pathways and bioprocesses revealed by KEGG and GO analyses. Specifically, many of the methylated genes were involved in biological processes, the cell cycle, DNA metabolic processes and cellular processes, and the adjustment of cell communication, signal transduction, and cell projection, which may be associated with the pathophysiological changes induced by BUO. By KEGG pathway analyses the significantly differently methylated genes were considerably linked to the Notch and steroid biosynthesis signaling pathways. The Notch signaling pathway is an evolutionarily conserved cell–cell communication mechanism and is essential for cell fate choice.30 For renal collecting duct cells, it has been demonstrated that inactivation of Notch signaling during kidney collecting duct development aberrantly favors the differentiation of collecting duct cells into ICs instead of PCs, which can express the water reabsorption-related proteins AQP2.31 The growth of an inadequate number of PCs could reduce the renal urine concentrating capacity. Previous studies found that lithium treatment could induce PCs into ICs through the Notch signaling pathway.32 In this study, we also found that the ratio of ICs was higher in BUO rat collecting ducts than in sham rats, while the expression level of Notch1 was dropped compared with that in the sham group, which indicates that the Notch signaling pathway-induced conversion between PCs and ICs may also exist in BUO-induced obstructive nephropathy and could be one of the potential mechanisms for ureteral obstruction-induced nephrogenic diabetes insipidus (NDI).
In addition, according to the conjoint analyses of MeRIP-seq and RNA-seq, we discovered that several genes that were associated with the regulation of AQP2 and renal water reabsorption were changed significantly. EGFR has been found to play an important role in water reabsorption. The suppression of EGFR reinforced AQP2 membrane accumulation in mouse kidney PCs and ameliorated lithium-induced NDI.33 In this study, we found that EGFR was demethylated 3.7-fold (3’UTR) compared with that in the sham group, while the mRNA expression was 1.33-fold that in the sham group. Since EGFR is a negative regulator of AQP2 expression, the high expression of EGFR paralleled the decreased renal water reabsorption ability; thus, EGFR mRNA m6A modification may be the potential mechanism for the dysregulation of EGFR-induced renal water reabsorption disability in BUO rats. In addition to EGFR, Brca1, one of the E3 ubiquitin (Ub)-protein ligases (E3s), was demonstrated to promote the degradation of AQP2 and thus affect urine concentration, which may have similar effects on AQP2.34 In this study, we found that Brcal was demethylated 2.72-fold (CDS) in the BUO group, and the mRNA expression was 1.9-fold that of the sham group. We speculate that the m6A modulation of Brcal mRNA leads to high expression of the Brcal gene, which induces the decreased expression of AQP2 and ultimately contributes to the impairment of urinary concentration. The disclosure of m6A modification information of the obstructive rat kidneys suggests that m6A modification adjusts the course of obstructive nephropathy by engaging in multiple dimensions.
Considering that the BUO model could simulate the pathological changes of human congenital ureteral pelvic junction obstruction-induced obstructive nephropathy, in the clinic, the treatment of ureteral obstruction-induced hydronephrosis mainly focuses on the surgical operation.35 In the current study, we found that the dysregulation of m6A modification may be connected with renal injury induced by BUO. The identified methylases, demethylases, and several functionally altered m6A transcripts could be potential therapeutic targets. In addition, there are chemicals that have been used in pathophysiological mechanism studies of m6A,36 including methylation inhibitors (e.g., 3-deazaadenosine37). The specific m6A-modified genes found in this study will provide a novel direction for the expansion of therapeutic drugs. However, a huge gap lies between laboratory research and clinical situations. Thereby, further studies are necessary to test whether changing the status of m6A will be the very target in the treatment of obstructive nephropathy.
Limitations of the study
In this study, we found that m6A modulation may be related to the dysregulation of genes and pathways involved in BUO-induced nephropathy, but we have not examined the direct effect of N6 methyltransferase dysregulation on this disease. In the future, we will conduct further research, including in vitro models with up- or downregulation of methylated/demethylated RNA species, to explore the direct effect of N6 methyltransferase on the regulation of specific genes and renal functions in BUO-induced obstructive nephropathy.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Shimada, K., Matsumoto, F., Kawagoe, M. & Matsui, F. Urological emergency in neonates with congenital hydronephrosis. Int J. Urol. 14, 388–392 (2007).
Ardissino, G. et al. Epidemiology of chronic renal failure in children: data from the ItalKid project. Pediatrics 111, e382–e387 (2003).
Kato, M. & Natarajan, R. Epigenetics and epigenomics in diabetic kidney disease and metabolic memory. Nat. Rev. Nephrol. 15, 327–345 (2019).
Zhou, P., Wu, M., Ye, C., Xu, Q. & Wang, L. Meclofenamic acid promotes cisplatin-induced acute kidney injury by inhibiting fat mass and obesity-associated protein-mediated m6A abrogation in RNA. J. Biol. Chem. 294, 16908–16917 (2019).
Shi, H., Wei, J. & He, C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol. Cell 74, 640–650 (2019).
Wei, W., Ji, X., Guo, X. & Ji, S. Regulatory role of N6-methyladenosine (m6A) methylation in RNA processing and human diseases. J. Cell Biochem. 118, 2534–2543 (2017).
Dorn, L. E. et al. The N6-methyladenosine mRNA methylase METTL3 controls cardiac homeostasis and hypertrophy. Circulation 139, 533–545 (2019).
Hubacek, J. A. et al. The FTO gene polymorphism is associated with end-stage renal disease: two large independent case-control studies in a general population. Nephrol. Dial. Transplant. 27, 1030–1035 (2012).
Spoto, B. et al. The fat-mass and obesity-associated gene (FTO) predicts mortality in chronic kidney disease of various severity. Nephrol. Dial. Transplant. 4(Suppl), iv58–iv62 (2012).
Li, X., Fan, X., Yin, X., Liu, H. & Yang, Y. Alteration of N6-methyladenosine epitranscriptome profile in unilateral ureteral obstructive nephropathy. Epigenomics 12, 1157–1173 (2020).
Xu, Y. et al. The N6-methyladenosine mRNA methylase METTL14 promotes renal ischemic reperfusion injury via suppressing YAP1. J. Cell Biochem. 121, 524–533 (2020).
Li, C., Wang, W., Knepper, M. A., Nielsen, S. & Frøkiaer, J. Downregulation of renal aquaporins in response to unilateral ureteral obstruction. Am. J. Physiol. Ren. Physiol. 284, F1066–F1079 (2003).
Feng, J. J. et al. Aquaporin1-3 expression in normal and hydronephrotic kidneys in the human fetus. Pediatr. Res. 86, 595–602 (2019).
Chevalier, R. L. Chronic partial ureteral obstruction and the developing kidney. Pediatr. Radiol. 38(Suppl 1), S35–S40 (2008).
Manucha, W. Biochemical-molecular markers in unilateral ureteral obstruction. Biocell 31, 1–12 (2007).
Hosohata, K., Jin, D., Takai, S. & Iwanaga, K. Vanin-1 in renal pelvic urine reflects kidney injury in a rat model of hydronephrosis. Int. J. Mol. Sci. 19, 3186 (2018).
Devocelle, A. et al. IL-15 prevents renal fibrosis by inhibiting collagen synthesis: a new pathway in chronic kidney disease? Int. J. Mol. Sci. 22, 11698 (2021).
Wei, W., Ji, X., Guo, X. & Ji, S. Regulatory role of N6-methyladenosine (m6A) methylation in RNA processing and human diseases. J. Cell Biochem. 118, 2534–2543 (2017).
Ramalingam, H. et al. A methionine-Mettl3-N6-methyladenosine axis promotes polycystic kidney disease. Cell Metab. 33, 1234–1247.e7 (2021).
Liu, S. et al. METTL3 plays multiple functions in biological processes. Am. J. Cancer Res. 10, 1631–1646 (2020).
Meng, F. et al. METTL3 contributes to renal ischemia-reperfusion injury by regulating Foxd1 methylation. Am. J. Physiol. Ren. Physiol. 319, F839–F847 (2020).
Li, M., Deng, L. & Xu, G. METTL14 promotes glomerular endothelial cell injury and diabetic nephropathy via m6A modification of α-klotho. Mol. Med. 27, 106 (2021).
Franceschini, N. et al. The association of genetic variants of type 2 diabetes with kidney function. Kidney Int. 82, 220–225 (2012).
Chen, G. et al. Association study of genetic variants of 17 diabetes-related genes/loci and cardiovascular risk and diabetic nephropathy in the Chinese She population. J. Diabetes 5, 136–145 (2013).
Wang, J. et al. METTL3/m6A/miRNA-873-5p attenuated oxidative stress and apoptosis in colistin-induced kidney injury by modulating Keap1/Nrf2 pathway. Front. Pharm. 10, 517 (2019).
Wang, J. et al. The biological function of m6A demethylase ALKBH5 and its role in human disease. Cancer Cell Int. 20, 347 (2020).
Zhu, S. & Lu, Y. Dexmedetomidine suppressed the biological behavior of HK-2 cell treated with LPS by down-regulating ALKBH5. Inflammation 43, 2256–2263 (2020).
Zhang, X. et al. ALKBH5 promotes the proliferation of renal cell carcinoma by regulating AURKB expression in an m6A-dependent manner. Ann. Transl. Med. 8, 646 (2020).
Ning, Y. et al. Genistein ameliorates renal fibrosis through regulation Snail via m6A RNA demethylase ALKBH5. Front. Pharm. 11, 579265 (2020).
Mukherjee, M., Fogarty, E., Janga, M. & Surendran, K. Notch signaling in kidney development, maintenance, and disease. Biomolecules 9, 692 (2019).
Mukherjee, M. et al. Endogenous Notch signaling in adult kidneys maintains segment-specific epithelial cell types of the distal tubules and collecting ducts to ensure water homeostasis. J. Am. Soc. Nephrol. 30, 110–126 (2019).
Trepiccione, F., Capasso, G., Nielsen, S. & Christensen, B. M. Evaluation of cellular plasticity in the collecting duct during recovery from lithium-induced nephrogenic diabetes insipidus. Am. J. Physiol. Ren. Physiol. 305, F919–F929 (2013).
Cheung, P. W. et al. EGF receptor inhibition by erlotinib increases aquaporin 2-mediated renal water reabsorption. J. Am. Soc. Nephrol. 27, 3105–3116 (2016).
Lee, Y. J. et al. E3 ubiquitin-protein ligases in rat kidney collecting duct: response to vasopressin stimulation and withdrawal. Am. J. Physiol. Ren. Physiol. 301, F883–F896 (2011).
Mittal, S. & Srinivasan, A. Robotics in pediatric urology: evolution and the future. Urol. Clin. North Am. 48, 113–125 (2021).
Nakano, M., Ondo, K., Takemoto, S., Fukami, T. & Nakajima, M. Methylation of adenosine at the N6position post-transcriptionally regulates hepatic P450s expression. Biochem. Pharm. 171, 113697 (2020).
Chiang, P. K. Conversion of 3T3-L1 fibroblasts to fat cells by an inhibitor of methylation: effect of 3-deazaadenosine. Science 211, 1164–1166 (1981).
Acknowledgements
We would like to thank Keke Ma, who was the administrator of the Laboratory Animal Center of Henan Province for helping us to feed the animals.
Funding
This study was funded by the Medical Science and Technology Research-related joint construction project of Henan Province (7220).
Author information
Authors and Affiliations
Contributions
J.F., Y.Z., J.W., Y.C., J.T., S.Y., Z.Z., B.D., Y.L. and X.Z. made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data. J.F., Y.Z., J.W., J.T., Y.F. and L.L. made substantial contributions to make animal model. J.F., X.Z., Y.Z., J.W., Y.C. and J.T. drafted the article or revised it critically for important intellectual content. X.Z. made the final approval of the version to be published. J.F., Y.Z., and J.W. made equal contribution to the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All procedures conformed to the Chinese National Guidelines for the Care and Handling of Animals and the published guidelines from the National Institutes of Clinical Medicine, Zhengzhou University, according to the licenses for use of experimental animals issued by the Chinese Ministry of Justice (2022-KY-0046-002).
Consent for publication
The submission of this manuscript for publication is approved by all authors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, J., Zhang, Y., Wen, J. et al. Alteration of N6-methyladenosine epitranscriptome profiles in bilateral ureteral obstruction-induced obstructive nephropathy in juvenile rats. Pediatr Res 93, 1509–1518 (2023). https://doi.org/10.1038/s41390-022-02228-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-022-02228-z