Abstract
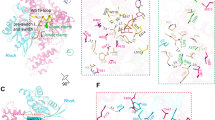
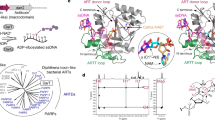
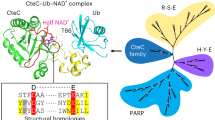
ExoS and ExoT are bifunctional type III cytotoxins of Pseudomonas aeruginosa that contain an N-terminal RhoGAP domain and a C-terminal ADP-ribosylation domain. Although they share 76% amino acid identity, ExoS and ExoT ADP-ribosylate different substrates. Using protein modeling and site-directed mutagenesis, the regions of ExoS and ExoT that define substrate specificity were determined. Regions B (active site loop), C (ARTT motif) and E (PN loop) on ExoS are necessary and sufficient to recognize ExoS targets, whereas regions B, C and E on ExoT are necessary but not sufficient to recognize ExoT targets, such as the Crk proteins. A specific Crk recognition motif on ExoT was defined as region A (helix α1). The electrostatic properties of regions A, B, C and E define the substrate specificity of ExoS and ExoT and these interactions can explain how other bacterial ADP-ribosylating toxins recognize their unique substrates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Krueger, K.M. & Barbieri, J.T. The family of bacterial ADP-ribosylating exotoxins. Clin. Microbiol. Rev. 8, 34–47 (1995).
Bjorn, M.J., Pavlovskis, O.R., Thompson, M.R. & Iglewski, B.H. Production of exoenzyme S during Pseudomonas aeruginosa infections of burned mice. Infect. Immun. 24, 837–842 (1979).
Kulich, S.M., Frank, D.W. & Barbieri, J.T. Purification and characterization of exoenzyme S from Pseudomonas aeruginosa 388. Infect. Immun. 61, 307–313 (1993).
Kulich, S.M., Yahr, T.L., Mende-Mueller, L.M., Barbieri, J.T. & Frank, D.W. Cloning the structural gene for the 49-kDa form of exoenzyme S (exoS) from Pseudomonas aeruginosa strain 388. J. Biol. Chem. 269, 10431–10437 (1994).
Yahr, T.L., Barbieri, J.T. & Frank, D.W. Genetic relationship between the 53- and 49-kilodalton forms of exoenzyme S from Pseudomonas aeruginosa. J. Bacteriol. 178, 1412–1419 (1996).
Goehring, U.M., Schmidt, G., Pederson, K.J., Aktories, K. & Barbieri, J.T. The N-terminal domain of Pseudomonas aeruginosa exoenzyme S is a GTPase-activating protein for Rho GTPases. J. Biol. Chem. 274, 36369–36372 (1999).
Krall, R., Schmidt, G., Aktories, K. & Barbieri, J.T. Pseudomonas aeruginosa ExoT is a Rho GTPase-activating protein. Infect. Immun. 68, 6066–6068 (2000).
Kazmierczak, B.I. & Engel, J.N. Pseudomonas aeruginosa ExoT acts in vivo as a GTPase-activating protein for RhoA, Rac1, and Cdc42. Infect. Immun. 70, 2198–2205 (2002).
Garrity-Ryan, L. et al. The arginine finger domain of ExoT contributes to actin cytoskeleton disruption and inhibition of internalization of Pseudomonas aeruginosa by epithelial cells and macrophages. Infect. Immun. 68, 7100–7113 (2000).
Geiser, T.K., Kazmierczak, B.I., Garrity-Ryan, L.K., Matthay, M.A. & Engel, J.N. Pseudomonas aeruginosa ExoT inhibits in vitro lung epithelial wound repair. Cell Microbiol. 3, 223–236 (2001).
Coburn, J., Dillon, S.T., Iglewski, B.H. & Gill, D.M. Exoenzyme S of Pseudomonas aeruginosa ADP-ribosylates the intermediate filament protein vimentin. Infect. Immun. 57, 996–998 (1989).
Coburn, J. & Gill, D.M. ADP-ribosylation of p21ras and related proteins by Pseudomonas aeruginosa exoenzyme S. Infect. Immun. 59, 4259–4262 (1991).
Ganesan, A.K., Vincent, T.S., Olson, J.C. & Barbieri, J.T. Pseudomonas aeruginosa exoenzyme S disrupts Ras-mediated signal transduction by inhibiting guanine nucleotide exchange factor-catalyzed nucleotide exchange. J. Biol. Chem. 274, 21823–21829 (1999).
Coburn, J., Kane, A.V., Feig, L. & Gill, D.M. Pseudomonas aeruginosa exoenzyme S requires a eukaryotic protein for ADP-ribosyltransferase activity. J. Biol. Chem. 266, 6438–6446 (1991).
Fu, H., Coburn, J. & Collier, R.J. The eukaryotic host factor that activates exoenzyme S of Pseudomonas aeruginosa is a member of the 14-3-3 protein family. Proc. Natl. Acad. Sci. USA 90, 2320–2324 (1993).
Liu, S., Kulich, S.M. & Barbieri, J.T. Identification of glutamic acid 381 as a candidate active site residue of Pseudomonas aeruginosa exoenzyme S. Biochemistry 35, 2754–2758 (1996).
Radke, J., Pederson, K.J. & Barbieri, J.T. Pseudomonas aeruginosa exoenzyme S is a biglutamic acid ADP-ribosyltransferase. Infect. Immun. 67, 1508–1510 (1999).
Liu, S., Yahr, T.L., Frank, D.W. & Barbieri, J.T. Biochemical relationships between the 53-kilodalton (Exo53) and 49-kilodalton (ExoS) forms of exoenzyme S of Pseudomonas aeruginosa. J. Bacteriol. 179, 1609–1613 (1997).
Sundin, C., Henriksson, M.L., Hallberg, B., Forsberg, A. & Frithz-Lindsten, E. Exoenzyme T of Pseudomonas aeruginosa elicits cytotoxicity without interfering with Ras signal transduction. Cell Microbiol. 3, 237–246 (2001).
Sun, J. & Barbieri, J.T. Pseudomonas aeruginosa ExoT ADP-ribosylates CT10 regulator of kinase (Crk) proteins. J. Biol. Chem. 278, 32794–32800 (2003).
Bell, C.E. & Eisenberg, D. Crystal structure of diphtheria toxin bound to nicotinamide adenine dinucleotide. Biochemistry 35, 1137–1149 (1996).
Sixma, T.K. et al. Refined structure of Escherichia coli heat-labile enterotoxin, a close relative of cholera toxin. J. Mol. Biol. 230, 890–918 (1993).
Sixma, T.K., Stein, P.E., Hol, W.G. & Read, R.J. Comparison of the B-pentamers of heat-labile enterotoxin and verotoxin-1: two structures with remarkable similarity and dissimilarity. Biochemistry 32, 191–198 (1993).
Wedekind, J.E. et al. Refined crystallographic structure of Pseudomonas aeruginosa exotoxin A and its implications for the molecular mechanism of toxicity. J. Mol. Biol. 314, 823–837 (2001).
Allured, V.S., Collier, R.J., Carroll, S.F. & McKay, D.B. Structure of exotoxin A of Pseudomonas aeruginosa at 3.0-Angstrom resolution. Proc. Natl. Acad. Sci. USA 83, 1320–1324 (1986).
Li, M., Dyda, F., Benhar, I., Pastan, I. & Davies, D.R. Crystal structure of the catalytic domain of Pseudomonas exotoxin A complexed with a nicotinamide adenine dinucleotide analog: implications for the activation process and for ADP ribosylation. Proc. Natl. Acad. Sci. USA 93, 6902–6906 (1996).
Stein, P.E. et al. The crystal structure of pertussis toxin. Structure 2, 45–57 (1994).
Han, S., Craig, J.A., Putnam, C.D., Carozzi, N.B. & Tainer, J.A. Evolution and mechanism from structures of an ADP-ribosylating toxin and NAD complex. Nat. Struct. Biol. 6, 932–936 (1999).
Han, S., Arvai, A.S., Clancy, S.B. & Tainer, J.A. Crystal structure and novel recognition motif of rho ADP-ribosylating C3 exoenzyme from Clostridium botulinum: structural insights for recognition specificity and catalysis. J. Mol. Biol. 305, 95–107 (2001).
Tsuge, H. et al. Crystal structure and site-directed mutagenesis of enzymatic components from Clostridium perfringens iota-toxin. J. Mol. Biol. 325, 471–483 (2003).
Evans, H.R. et al. The crystal structure of C3stau2 from Staphylococcus aureus and its complex with NAD. J. Biol. Chem. 278, 45924–45930 (2003).
Menetrey, J. et al. NAD binding induces conformational changes in Rho ADP-ribosylating Clostridium botulinum C3 exoenzyme. J. Biol. Chem. 277, 30950–30957 (2002).
Van Ness, B.G., Howard, J.B. & Bodley, J.W. ADP-ribosylation of elongation factor 2 by diphtheria toxin. Isolation and properties of the novel ribosyl-amino acid and its hydrolysis products. J. Biol. Chem. 255, 10717–10720 (1980).
Iglewski, B.H. & Kabat, D. NAD-dependent inhibition of protein synthesis by Pseudomonas aeruginosa toxin. Proc. Natl. Acad. Sci. USA 72, 2284–2288 (1975).
Gierschik, P. ADP-ribosylation of signal-transducing guanine nucleotide-binding proteins by pertussis toxin. Curr. Top. Microbiol. Immunol. 175, 69–96 (1992).
Moss, J. & Richardson, S.H. Activation of adenylate cyclase by heat-labile Escherichia coli enterotoxin. Evidence for ADP-ribosyltransferase activity similar to that of choleragen. J. Clin. Invest. 62, 281–285 (1978).
Aktories, K. et al. Botulinum C2 toxin ADP-ribosylates actin. Nature 322, 390–392 (1986).
Schering, B., Barmann, M., Chhatwal, G.S., Geipel, U. & Aktories, K. ADP-ribosylation of skeletal muscle and non-muscle actin by Clostridium perfringens iota toxin. Eur. J. Biochem. 171, 225–229 (1988).
Aktories, K. & Frevert, J. ADP-ribosylation of a 21–24 kDa eukaryotic protein(s) by C3, a novel botulinum ADP-ribosyltransferase, is regulated by guanine nucleotide. Biochem. J. 247, 363–368 (1987).
Wilde, C., Chhatwal, G.S., Schmalzing, G., Aktories, K. & Just, I. A novel C3-like ADP-ribosyltransferase from Staphylococcus aureus modifying RhoE and Rnd3. J. Biol. Chem. 276, 9537–9542 (2001).
Wilde, C., Just, I. & Aktories, K. Structure-function analysis of the Rho-ADP-ribosylating exoenzyme C3stau2 from Staphylococcus aureus. Biochemistry 41, 1539–1544 (2002).
Schwede, T., Kopp, J., Guex, N. & Peitsch, M.C. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res. 31, 3381–3385 (2003).
Guex, N. & Peitsch, M.C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18, 2714–2723 (1997).
Acknowledgements
This study was supported by a grant from the US National Institutes of Health (AI-30162) to J.T.B.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Indicated toxins were incubated with GST-Crk-I or moesin with the presence of NAD/P32-NAD and FAS, and reaction rates (V) were calculated as described in Methods. The data from at least three independent experiments were fit to Michaelis-Menten equation in both SigmaPlot and EnzFitter, where the values of Km and Vmax were calculated. (PDF 84 kb)
Rights and permissions
About this article
Cite this article
Sun, J., Maresso, A., Kim, JJ. et al. How bacterial ADP-ribosylating toxins recognize substrates. Nat Struct Mol Biol 11, 868–876 (2004). https://doi.org/10.1038/nsmb818
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb818
This article is cited by
-
Molecular basis of threonine ADP-ribosylation of ubiquitin by bacterial ARTs
Nature Chemical Biology (2024)
-
14-3-3 proteins activate Pseudomonas exotoxins-S and -T by chaperoning a hydrophobic surface
Nature Communications (2018)
-
Insights into the biogenesis, function, and regulation of ADP-ribosylation
Nature Chemical Biology (2018)
-
Novel bacterial ADP-ribosylating toxins: structure and function
Nature Reviews Microbiology (2014)
-
ADP-ribosylation of arginine
Amino Acids (2011)