Abstract
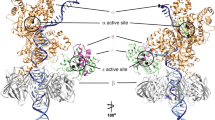
The ability or inability of a DNA polymerase to extend a mispair directly affects the establishment of genomic mutations. We report here kinetic analyses of the ability of Dpo4, a Y-family polymerase from Sulfolobus solfataricus, to extend from all mispairs opposite a template G or T. Dpo4 is equally inefficient at extending these mispairs, which include, surprisingly, a G·T mispair expected to conform closely to Watson-Crick geometry. To elucidate the basis of this, we solved the structure of Dpo4 bound to G·T-mispaired primer template in the presence of an incoming nucleotide. As a control, we also determined the structure of Dpo4 bound to a matched A-T base pair at the primer terminus. The structures offer a basis for the low efficiency of Dpo4 in extending a G·T mispair: a reverse wobble that deflects the primer 3′-OH away from the incoming nucleotide.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Goodman, M.F. Error-prone repair DNA polymerases in prokaryotes and eukaryotes. Annu. Rev. Biochem. 71, 17–50 (2002).
Prakash, S. & Prakash, L. Translesion DNA synthesis in eukaryotes: a one- or two-polymerase affair. Genes Dev. 16, 1872–1883 (2002).
Doublie, S., Sawaya, M.R. & Ellenberger, T. An open and closed case for all polymerases. Structure 7, R31–R35 (1999).
Mendelman, L.V., Petruska, J. & Goodman, M.F. Base mispair extension kinetics. Comparison of DNA polymerase α and reverse transcriptase. J. Biol. Chem. 265, 2338–2346 (1990).
Huang, M.-M., Arnheim, N. & Goodman, M.F. Extension of base mispairs by Taq DNA polymerase: implications for single nucleotide discrimination in PCR. Nucleic Acids Res. 20, 4567–4573 (1992).
Joyce, C.M., Sun, X.C. & Grindley, N.D. Reactions at the polymerase active site that contribute to the fidelity of Escherichia coli DNA polymerase I (Klenow fragment). J. Biol. Chem. 267, 24485–24500 (1992).
Washington, M.T., Johnson, R.E., Prakash, S. & Prakash, L. Mismatch extension ability of yeast and human DNA polymerase η. J. Biol. Chem. 276, 2263–2266 (2001).
Vaisman, A., Tissier, A., Frank, E.G., Goodman, M.F. & Woodgate, R. Human DNA polymerase ι promiscuous mismatch extension. J. Biol. Chem. 276, 30615–30622 (2001).
Washington, M.T., Johnson, R.E., Prakash, L. & Prakash, S. Human DINB1-encoded DNA polymerase κ is a promiscuous extender of mispaired primer termini. Proc. Nat. Acad. Sci. USA 99, 1910–1914 (2002).
Haracska, L., Prakash, L. & Prakash, S. Role of human DNA polymerase κ as an extender in translesion synthesis. Proc. Nat. Acad. Sci. USA 99, 16000–16005 (2002).
Ling, H., Boudsocq, F., Woodgate, R. & Yang, W. Crystal structure of a Y-family DNA polymerase in action: a mechanism for error-prone and lesion-bypass replication. Cell 107, 91–102 (2001).
Ling, H., Boudsocq, F., Plosky, B.S., Woodgate, R. & Yang, W. Replication of cis-syn thymine dimer at atomic resolution. Nature 242, 1083–1087 (2003).
Steitz, T.A. DNA polymerases: structural diversity and common mechanisms. J. Biol. Chem. 274, 17395–17398 (1999).
Trincao, J. et al. Structure of the catalytic core of S. cerevisiae DNA polymerase η: implications for translesion DNA synthesis. Mol. Cell 8, 417–426 (2001).
Zhou, B.-L., Pata, J.D. & Steitz, T.A. Crystal structure of a DinB lesion bypass DNA polymerase catalytic fragment reveals a classic polymerase catalytic domain. Mol. Cell 8, 427–437 (2001).
Silvian, L.F., Toth, E.A., Pham, P., Goodman, M.F. & Ellenberger, T. Crystal structure of a DinB family error-prone DNA polymerase from Sulfolobus solfataricus . Nat. Struct. Biol. 8, 984–989 (2001).
Johnson, R.E., Trincao, J., Aggarwal, A.K., Prakash, S. & Prakash, L. Deoxynucleotide triphosphate binding mode conserved in Y family DNA polymerases. Mol. Cell. Biol. 23, 3008–3012 (2003).
Leontis, N.B., Stombaugh, J. & Westhof, E. The non-Watson-Crick base pairs and their associated isostericity matrices. Nucleic Acids Res. 30, 3497–3531 (2002).
Hunter, W.N., Kneale, G., Brown, T., Rabinovich, D. & Kennard, O. Refined crystal structure of an actanucleotide duplex with G·T mismatched base-pairs. J. Mol. Biol. 190, 605–618 (1986).
Hare, D., Shapiro, L. & Patel, D.J. Wobble dG × dT pairing in right-handed DNA: solution conformation of the d(C-G-T-G-A-A-T-T-C-G-C-G) duplex deduced from distance geometry analysis of nuclear Overhauser effect spectra. Biochemistry 25, 7445–7456 (1986).
Mooers, B.H., Eichman, B.F. & Ho, P.S. The structures and relative stabilities of d(G × G) reverse Hoogsteen, d(G × T) reverse wobble, and d(G × C) reverse Watson-Crick base-pairs in DNA crystals. J. Mol. Biol. 269, 796–810 (1997).
Perrino, F.W. & Loeb, L.A. Differential extension of 3′ mispairs is a major contribution to the high fidelity of calf thymus DNA polymerase α. J. Biol. Chem. 264, 2898–2905 (1989).
Varani, G., Cheong, C. & Tinoco, I. Jr. Structure of an unusually stable RNA hairpin. Biochemistry 30, 3280–3289 (1991).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science 289, 905–920 (2000).
Bullock, T.L., Sherlin, L.D. & Perona, J.J. Tertiary core rearrangements in a tight binding transfer RNA aptamer. Nat. Struct. Biol. 7, 497–504 (2000).
Klinck, R. et al. A potential RNA drug target in the hepatitis C virus internal ribosomal entry site. RNA 6, 1423–1431 (2000).
Johnson, R.E., Yu, S.-L., Prakash, S. & Prakash, L. Yeast DNA polymerase zeta (ζ) is essential for error-free replication past thymine glycol. Genes Dev. 17, 77–87 (2003).
Johnson, R.E., Washington, M.T., Prakash, S. & Prakash, L. Fidelity of human DNA polymerase η. J. Biol. Chem. 275, 7447–7450 (2000).
Aggarwal, A.K. Crystallization of DNA binding proteins with oligodeoxynucleotides. Methods 1, 83–90 (1990).
Brunger, A.T. et al. Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, A.T., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Koradi, R., Billeter, M. & Wuthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996).
Acknowledgements
We thank the staff at the Advanced Photon Source and the National Synchrotron Light Source for facilitating X-ray data collection. We thank T. Edwards for help with data collection and processing. This was supported was supported by US National Institutes of Health grant CA094006 (A.K.A. and L.P.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Trincao, J., Johnson, R., Wolfle, W. et al. Dpo4 is hindered in extending a G·T mismatch by a reverse wobble. Nat Struct Mol Biol 11, 457–462 (2004). https://doi.org/10.1038/nsmb755
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb755
This article is cited by
-
Structures of LIG1 that engage with mutagenic mismatches inserted by polβ in base excision repair
Nature Communications (2022)
-
Effect of ionic liquid on the long-term structural and chemical stability of basidiomycetes DNAs integrated within Schottky-like junctions
Applied Physics A (2021)
-
Biological and nanotechnological applications using interactions between ionic liquids and nucleic acids
Biophysical Reviews (2018)
-
Comparable Stability of Hoogsteen and Watson–Crick Base Pairs in Ionic Liquid Choline Dihydrogen Phosphate
Scientific Reports (2014)
-
A nucleotide binding rectification Brownian ratchet model for translocation of Y-family DNA polymerases
Theoretical Biology and Medical Modelling (2011)