Abstract
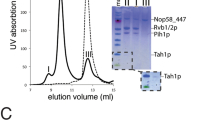
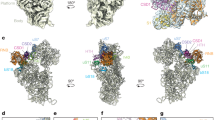
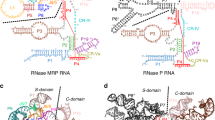
Ribonucleoprotein particles are central to numerous cellular pathways, but their study in vitro is often complicated by heterogeneity and aggregation. We describe a new technique to characterize these complexes trapped as homogeneous species in a nondenaturing gel. Using this technique, in conjunction with phosphorothioate footprinting analysis, we identify the protein-binding site and RNA folding states of ribonuclease P (RNase P), an RNA-based enzyme that, in vivo, requires a protein cofactor to catalyze the 5′ maturation of precursor transfer RNA (pre-tRNA). Our results show that the protein binds to a patch of conserved RNA structure adjacent to the active site and influences the conformation of the RNA near the tRNA-binding site. The data are consistent with a role of the protein in substrate recognition and support a new model of the holoenzyme that is based on a recently solved crystal structure of RNase P RNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Reich, C., Olsen, G.J., Pace, B. & Pace, N.R. Role of the protein moiety of ribonuclease P, a ribonucleoprotein enzyme. Science 239, 178–181 (1988).
Kurz, J.C., Niranjanakumari, S. & Fierke, C.A. Protein component of Bacillus subtilis RNase P specifically enhances the affinity for precursor-tRNAAsp. Biochemistry 37, 2393–2400 (1998).
Buck, A.H., Dalby, A.B., Poole, A., Kazantsev, A.V. & Pace, N.R. Protein activation of a ribozyme: the role of bacterial RNase P protein. EMBO J. published online 15 September 2005 (10.1038/sj.emboj.7600805).
Kazantsev, A.V. et al. Crystal structure of a bacterial ribonuclease P RNA. Proc. Natl. Acad. Sci. USA 102, 13392–13397 (2005).
Loria, A., Niranjanakumari, S., Fierke, C.A. & Pan, T. Recognition of a pre-tRNA substrate by the Bacillus subtilis RNase P holoenzyme. Biochemistry 37, 15466–15473 (1998).
Rox, C., Feltens, R., Pfeiffer, T. & Hartmann, R.K. Potential contact sites between the protein and RNA subunit in the Bacillus subtilis RNase P holoenzyme. J. Mol. Biol. 315, 551–560 (2002).
Day-Storms, J.J., Niranjanakumari, S. & Fierke, C.A. Ionic interactions between PRNA and P protein in Bacillus subtilis RNase P characterized using a magnetocapture-based assay. RNA 10, 1595–1608 (2004).
Loria, A. & Pan, T. Modular construction for function of a ribonucleoprotein enzyme: the catalytic domain of Bacillus subtilis RNase P complexed with B. subtilis RNase P protein. Nucleic Acids Res. 29, 1892–1897 (2001).
Waugh, D.S. & Pace, N.R. Complementation of an RNase P RNA (rnpB) gene deletion in Escherichia coli by homologous genes from distantly related eubacteria. J. Bacteriol. 172, 6316–6322 (1990).
Vioque, A., Arnez, J. & Altman, S. Protein-RNA interactions in the RNase P holoenzyme from Escherichia coli. J. Mol. Biol. 202, 835–848 (1988).
Talbot, S.J. & Altman, S. Gel retardation analysis of the interaction between C5 protein and M1 RNA in the formation of the ribonuclease P holoenzyme from Escherichia coli. Biochemistry 33, 1399–1405 (1994).
Westhof, E., Wesolowski, D. & Altman, S. Mapping in three dimensions of regions in a catalytic RNA protected from attack by an Fe(II)-EDTA reagent. J. Mol. Biol. 258, 600–613 (1996).
Sharkady, S.M. & Nolan, J.M. Bacterial ribonuclease P holoenzyme crosslinking analysis reveals protein interaction sites on the RNA subunit. Nucleic Acids Res. 29, 3848–3856 (2001).
Biswas, R., Ledman, D.W., Fox, R.O., Altman, S. & Gopalan, V. Mapping RNA-protein interactions in ribonuclease P from Escherichia coli using disulfide-linked EDTA-Fe. J. Mol. Biol. 296, 19–31 (2000).
Tsai, H.Y., Masquida, B., Biswas, R., Westhof, E. & Gopalan, V. Molecular modeling of the three-dimensional structure of the bacterial RNase P holoenzyme. J. Mol. Biol. 325, 661–675 (2003).
Fang, X.W. et al. The Bacillus subtilis RNase P holoenzyme contains two RNase P RNA and two RNase P protein subunits. RNA 7, 233–241 (2001).
Haas, E.S., Banta, A.B., Harris, J.K., Pace, N.R. & Brown, J.W. Structure and evolution of ribonuclease P RNA in Gram-positive bacteria. Nucleic Acids Res. 24, 4775–4782 (1996).
Brown, J.W. & Pace, N.R. Ribonuclease P RNA and protein subunits from bacteria. Nucleic Acids Res. 20, 1451–1456 (1992).
Kazantsev, A.V. et al. High-resolution structure of RNase P protein from Thermotoga maritima. Proc. Natl. Acad. Sci. USA 100, 7497–7502 (2003).
Stams, T., Niranjanakumari, S., Fierke, C.A. & Christianson, D.W. Ribonuclease P protein structure: evolutionary origins in the translational apparatus. Science 280, 752–755 (1998).
Gish, G. & Eckstein, F. DNA and RNA sequence determination based on phosphorothioate chemistry. Science 240, 1520–1522 (1988).
Winkler, W.C., Nahvi, A., Roth, A., Collins, J.A. & Breaker, R.R. Control of gene expression by a natural metabolite-responsive ribozyme. Nature 428, 281–286 (2004).
Christian, E.L., Zahler, N.H., Kaye, N.M. & Harris, M.E. Analysis of substrate recognition by the ribonucleoprotein endonuclease RNase P. Methods 28, 307–322 (2002).
Chen, J.L., Nolan, J.M., Harris, M.E. & Pace, N.R. Comparative photocross-linking analysis of the tertiary structures of Escherichia coli and Bacillus subtilis RNase P RNAs. EMBO J. 17, 1515–1525 (1998).
Kufel, J. & Kirsebom, L.A. The P15-loop of Escherichia coli RNase P RNA is an autonomous divalent metal ion binding domain. RNA 4, 777–788 (1998).
Oh, B.K., Frank, D.N. & Pace, N.R. Participation of the 3′-CCA of tRNA in the binding of catalytic Mg2+ ions by ribonuclease P. Biochemistry 37, 7277–7283 (1998).
Kazakov, S. & Altman, S. Site-specific cleavage by metal ion cofactors and inhibitors of M1 RNA, the catalytic subunit of RNase P from Escherichia coli. Proc. Natl. Acad. Sci. USA 88, 9193–9197 (1991).
Massire, C., Jaeger, L. & Westhof, E. Derivation of the three-dimensional architecture of bacterial ribonuclease P RNAs from comparative sequence analysis. J. Mol. Biol. 279, 773–793 (1998).
Niranjanakumari, S., Stams, T., Crary, S.M., Christianson, D.W. & Fierke, C.A. Protein component of the ribozyme ribonuclease P alters substrate recognition by directly contacting precursor tRNA. Proc. Natl. Acad. Sci. USA 95, 15212–15217 (1998).
Crary, S.M., Niranjanakumari, S. & Fierke, C.A. The protein component of Bacillus subtilis ribonuclease P increases catalytic efficiency by enhancing interactions with the 5′ leader sequence of pre-tRNAAsp. Biochemistry 37, 9409–9416 (1998).
Pan, T. & Sosnick, T.R. Intermediates and kinetic traps in the folding of a large ribozyme revealed by circular dichroism and UV absorbance spectroscopies and catalytic activity. Nat. Struct. Biol. 4, 931–938 (1997).
Fang, X., Pan, T. & Sosnick, T.R. A thermodynamic framework and cooperativity in the tertiary folding of a Mg2+-dependent ribozyme. Biochemistry 38, 16840–16846 (1999).
Haas, E.S., Morse, D.P., Brown, J.W., Schmidt, F.J. & Pace, N.R. Long-range structure in ribonuclease P RNA. Science 254, 853–856 (1991).
Beebe, J.A. & Fierke, C.A. A kinetic mechanism for cleavage of precursor tRNAAsp catalyzed by the RNA component of Bacillus subtilis ribonuclease P. Biochemistry 33, 10294–10304 (1994).
Smith, D. & Pace, N.R. Multiple magnesium ions in the ribonuclease P reaction mechanism. Biochemistry 32, 5273–5281 (1993).
Talbot, S.J. & Altman, S. Kinetic and thermodynamic analysis of RNA-protein interactions in the RNase P holoenzyme from Escherichia coli. Biochemistry 33, 1406–1411 (1994).
Williamson, J.R. Induced fit in RNA-protein recognition. Nat. Struct. Biol. 7, 834–837 (2000).
Kurz, J.C. & Fierke, C.A. The affinity of magnesium binding sites in the Bacillus subtilis RNase P × pre-tRNA complex is enhanced by the protein subunit. Biochemistry 41, 9545–9558 (2002).
Frank, D.N. & Pace, N.R. In vitro selection for altered divalent metal specificity in the RNase P RNA. Proc. Natl. Acad. Sci. USA 94, 14355–14360 (1997).
Christian, E.L., Kaye, N.M. & Harris, M.E. Helix P4 is a divalent metal ion binding site in the conserved core of the ribonuclease P ribozyme. RNA 6, 511–519 (2000).
Crary, S.M., Kurz, J.C. & Fierke, C.A. Specific phosphorothioate substitutions probe the active site of Bacillus subtilis ribonuclease P. RNA 8, 933–947 (2002).
Brannvall, M., Fredrik Pettersson, B.M. & Kirsebom, L.A. The residue immediately upstream of the RNase P cleavage site is a positive determinant. Biochimie 84, 693–703 (2002).
LaGrandeur, T.E., Huttenhofer, A., Noller, H.F. & Pace, N.R. Phylogenetic comparative chemical footprint analysis of the interaction between ribonuclease P RNA and tRNA. EMBO J. 13, 3945–3952 (1994).
Zahler, N.H., Christian, E.L. & Harris, M.E. Recognition of the 5′ leader of pre-tRNA substrates by the active site of ribonuclease P. RNA 9, 734–745 (2003).
Zuleeg, T. et al. Correlation between processing efficiency for ribonuclease P minimal substrates and conformation of the nucleotide −1 at the cleavage position. Biochemistry 40, 3363–3369 (2001).
Christian, E.L. & Harris, M.E. The track of the pre-tRNA 5′ leader in the ribonuclease P ribozyme-substrate complex. Biochemistry 38, 12629–12638 (1999).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brown, J.W. The ribonuclease P database. Nucleic Acids Res. 27, 314 (1999).
Acknowledgements
We thank A. Krivenko for cloning and initial characterization of the catalytic-domain constructs. We thank R. Batey for helpful comments regarding this work. The research was supported by The US National Institutes of Health (grant GM-34527 to N.R.P. and Molecular Biophysics Training Grant T32 GM-65103 to A.H.B.) through the University of Colorado at Boulder.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Phosphorothioate-I2 footprint of RNase P RNAs in the presence of RNase P proteins. (PDF 25 kb)
Supplementary Table 2
Difference in phosphorothioate-I2 cleavage of residues in the I and N folding states of RNase P RNA. (PDF 40 kb)
Supplementary Video 1
Full rotational view of the ternary complex model. (MOV 1780 kb)
Rights and permissions
About this article
Cite this article
Buck, A., Kazantsev, A., Dalby, A. et al. Structural perspective on the activation of RNase P RNA by protein. Nat Struct Mol Biol 12, 958–964 (2005). https://doi.org/10.1038/nsmb1004
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1004
This article is cited by
-
NMR resonance assignments of RNase P protein from Thermotoga maritima
Biomolecular NMR Assignments (2018)
-
Essential is Not Irreplaceable: Fitness Dynamics of Experimental E. coli RNase P RNA Heterologous Replacement
Journal of Molecular Evolution (2014)
-
The ancient history of the structure of ribonuclease P and the early origins of Archaea
BMC Bioinformatics (2010)
-
Structure of a bacterial ribonuclease P holoenzyme in complex with tRNA
Nature (2010)
-
Hfq stimulates the activity of the CCA-adding enzyme
BMC Molecular Biology (2007)