Abstract
Cotranslational chaperones assist de novo folding of nascent polypeptides, prevent them from aggregating and modulate translation. The ribosome-associated complex (RAC) is unique in that the Hsp40 protein Zuo1 and the atypical Hsp70 chaperone Ssz1 form a stable heterodimer, which acts as a cochaperone for the Hsp70 chaperone Ssb. Here we present the structure of the Chaetomium thermophilum RAC core comprising Ssz1 and the Zuo1 N terminus. We show how the conserved allostery of Hsp70 proteins is abolished and this Hsp70–Hsp40 pair is molded into a functional unit. Zuo1 stabilizes Ssz1 in trans through interactions that in canonical Hsp70s occur in cis. Ssz1 is catalytically inert and cannot adopt the closed conformation, but the substrate binding domain β is completed by Zuo1. Our study offers insights into the coupling of a special Hsp70–Hsp40 pair, which evolved to link protein folding and translation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Grudnik, P., Bange, G. & Sinning, I. Protein targeting by the signal recognition particle. Biol. Chem. 390, 775–782 (2009).
Wild, K. & Sinning, I. RNA gymnastics in mammalian signal recognition particle assembly. RNA Biol. 11, 1330–1334 (2014).
Drazic, A., Myklebust, L.M., Ree, R. & Arnesen, T. The world of protein acetylation. Biochim. Biophys. Acta 1864, 1372–1401 (2016).
Preissler, S. & Deuerling, E. Ribosome-associated chaperones as key players in proteostasis. Trends Biochem. Sci. 37, 274–283 (2012).
Kramer, G., Boehringer, D., Ban, N. & Bukau, B. The ribosome as a platform for co-translational processing, folding and targeting of newly synthesized proteins. Nat. Struct. Mol. Biol. 16, 589–597 (2009).
Mayer, M.P., Brehmer, D., Gässler, C.S. & Bukau, B. Hsp70 chaperone machines. Adv. Protein Chem. 59, 1–44 (2001).
Hartl, F.U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Mayer, M.P. & Bukau, B. Hsp70 chaperones: cellular functions and molecular mechanism. Cell. Mol. Life Sci. 62, 670–684 (2005).
Mayer, M.P. & Kityk, R. Insights into the molecular mechanism of allostery in Hsp70s. Front. Mol. Biosci. 2, 58 (2015).
Yang, J., Nune, M., Zong, Y., Zhou, L. & Liu, Q. Close and allosteric opening of the polypeptide-binding site in a human Hsp70 chaperone BiP. Structure 23, 2191–2203 (2015).
Bertelsen, E.B., Chang, L., Gestwicki, J.E. & Zuiderweg, E.R. Solution conformation of wild-type E. coli Hsp70 (DnaK) chaperone complexed with ADP and substrate. Proc. Natl. Acad. Sci. USA 106, 8471–8476 (2009).
Mayer, M.P., Laufen, T., Paal, K., McCarty, J.S. & Bukau, B. Investigation of the interaction between DnaK and DnaJ by surface plasmon resonance spectroscopy. J. Mol. Biol. 289, 1131–1144 (1999).
Jiang, J. et al. Structural basis of J cochaperone binding and regulation of Hsp70. Mol. Cell 28, 422–433 (2007).
Chen, D.H., Huang, Y., Liu, C., Ruan, Y. & Shen, W.H. Functional conservation and divergence of J-domain-containing ZUO1/ZRF orthologs throughout evolution. Planta 239, 1159–1173 (2014).
Pech, M., Spreter, T., Beckmann, R. & Beatrix, B. Dual binding mode of the nascent polypeptide-associated complex reveals a novel universal adapter site on the ribosome. J. Biol. Chem. 285, 19679–19687 (2010).
Hallstrom, T.C., Katzmann, D.J., Torres, R.J., Sharp, W.J. & Moye-Rowley, W.S. Regulation of transcription factor Pdr1p function by an Hsp70 protein in Saccharomyces cerevisiae. Mol. Cell. Biol. 18, 1147–1155 (1998).
Zhang, S., Lockshin, C., Herbert, A., Winter, E. & Rich, A. Zuotin, a putative Z-DNA binding protein in Saccharomyces cerevisiae. EMBO J. 11, 3787–3796 (1992).
Yan, W. et al. Zuotin, a ribosome-associated DnaJ molecular chaperone. EMBO J. 17, 4809–4817 (1998).
Gautschi, M. et al. RAC, a stable ribosome-associated complex in yeast formed by the DnaK-DnaJ homologs Ssz1p and zuotin. Proc. Natl. Acad. Sci. USA 98, 3762–3767 (2001).
Fiaux, J. et al. Structural analysis of the ribosome-associated complex (RAC) reveals an unusual Hsp70/Hsp40 interaction. J. Biol. Chem. 285, 3227–3234 (2010).
Peisker, K., Chiabudini, M. & Rospert, S. The ribosome-bound Hsp70 homolog Ssb of Saccharomyces cerevisiae. Biochim. Biophys. Acta 1803, 662–672 (2010).
Hundley, H. et al. The in vivo function of the ribosome-associated Hsp70, Ssz1, does not require its putative peptide-binding domain. Proc. Natl. Acad. Sci. USA 99, 4203–4208 (2002).
Conz, C. et al. Functional characterization of the atypical Hsp70 subunit of yeast ribosome-associated complex. J. Biol. Chem. 282, 33977–33984 (2007).
Pfund, C. et al. The molecular chaperone Ssb from Saccharomyces cerevisiae is a component of the ribosome-nascent chain complex. EMBO J. 17, 3981–3989 (1998).
Leidig, C. et al. Structural characterization of a eukaryotic chaperone—the ribosome-associated complex. Nat. Struct. Mol. Biol. 20, 23–28 (2013).
Zhang, Y. et al. Structural basis for interaction of a cotranslational chaperone with the eukaryotic ribosome. Nat. Struct. Mol. Biol. 21, 1042–1046 (2014).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G.J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
van Noort, V. et al. Consistent mutational paths predict eukaryotic thermostability. BMC Evol. Biol. 13, 7 (2013).
Amlacher, S. et al. Insight into structure and assembly of the nuclear pore complex by utilizing the genome of a eukaryotic thermophile. Cell 146, 277–289 (2011).
Kityk, R., Kopp, J., Sinning, I. & Mayer, M.P. Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones. Mol. Cell 48, 863–874 (2012).
Kabsch, W., Kabsch, H. & Eisenberg, D. Packing in a new crystalline form of glutamine synthetase from Escherichia coli. J. Mol. Biol. 100, 283–291 (1976).
Kabsch, W. & Holmes, K.C. The actin fold. FASEB J. 9, 167–174 (1995).
Flaherty, K.M., DeLuca-Flaherty, C. & McKay, D.B. Three-dimensional structure of the ATPase fragment of a 70K heat-shock cognate protein. Nature 346, 623–628 (1990).
Woo, H.J., Jiang, J., Lafer, E.M. & Sousa, R. ATP-induced conformational changes in Hsp70: molecular dynamics and experimental validation of an in silico predicted conformation. Biochemistry 48, 11470–11477 (2009).
Qi, R. et al. Allosteric opening of the polypeptide-binding site when an Hsp70 binds ATP. Nat. Struct. Mol. Biol. 20, 900–907 (2013).
Gumiero, A. et al. Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain. Nat. Commun. 7, 13563 (2016).
Bhattacharya, A. et al. Allostery in Hsp70 chaperones is transduced by subdomain rotations. J. Mol. Biol. 388, 475–490 (2009).
Gautschi, M., Mun, A., Ross, S. & Rospert, S. A functional chaperone triad on the yeast ribosome. Proc. Natl. Acad. Sci. USA 99, 4209–4214 (2002).
Raue, U., Oellerer, S. & Rospert, S. Association of protein biogenesis factors at the yeast ribosomal tunnel exit is affected by the translational status and nascent polypeptide sequence. J. Biol. Chem. 282, 7809–7816 (2007).
Gragerov, A. & Gottesman, M.E. Different peptide binding specificities of hsp70 family members. J. Mol. Biol. 241, 133–135 (1994).
Schlecht, R., Erbse, A.H., Bukau, B. & Mayer, M.P. Mechanics of Hsp70 chaperones enables differential interaction with client proteins. Nat. Struct. Mol. Biol. 18, 345–351 (2011).
Kityk, R., Vogel, M., Schlecht, R., Bukau, B. & Mayer, M.P. Pathways of allosteric regulation in Hsp70 chaperones. Nat. Commun. 6, 8308 (2015).
Zhu, X. et al. Structural analysis of substrate binding by the molecular chaperone DnaK. Science 272, 1606–1614 (1996).
Willmund, F. et al. The cotranslational function of ribosome-associated Hsp70 in eukaryotic protein homeostasis. Cell 152, 196–209 (2013).
Lee, K., Sharma, R., Shrestha, O.K., Bingman, C.A. & Craig, E.A. Dual interaction of the Hsp70 J-protein cochaperone Zuotin with the 40S and 60S ribosomal subunits. Nat. Struct. Mol. Biol. 23, 1003–1010 (2016).
Kaschner, L.A., Sharma, R., Shrestha, O.K., Meyer, A.E. & Craig, E.A. A conserved domain important for association of eukaryotic J-protein co-chaperones Jjj1 and Zuo1 with the ribosome. Biochim. Biophys. Acta 1853, 1035–1045 (2015).
Huang, P., Gautschi, M., Walter, W., Rospert, S. & Craig, E.A. The Hsp70 Ssz1 modulates the function of the ribosome-associated J-protein Zuo1. Nat. Struct. Mol. Biol. 12, 497–504 (2005).
Rakwalska, M. & Rospert, S. The ribosome-bound chaperones RAC and Ssb1/2p are required for accurate translation in Saccharomyces cerevisiae. Mol. Cell. Biol. 24, 9186–9197 (2004).
Halic, M. et al. Structure of the signal recognition particle interacting with the elongation-arrested ribosome. Nature 427, 808–814 (2004).
Greber, B.J. et al. Insertion of the Biogenesis Factor Rei1 Probes the Ribosomal Tunnel during 60S Maturation. Cell 164, 91–102 (2016).
Greber, B.J., Boehringer, D., Montellese, C. & Ban, N. Cryo-EM structures of Arx1 and maturation factors Rei1 and Jjj1 bound to the 60S ribosomal subunit. Nat. Struct. Mol. Biol. 19, 1228–1233 (2012).
Meyer, A.E., Hung, N.J., Yang, P., Johnson, A.W. & Craig, E.A. The specialized cytosolic J-protein, Jjj1, functions in 60S ribosomal subunit biogenesis. Proc. Natl. Acad. Sci. USA 104, 1558–1563 (2007).
Meyer, A.E., Hoover, L.A. & Craig, E.A. The cytosolic J-protein, Jjj1, and Rei1 function in the removal of the pre-60 S subunit factor Arx1. J. Biol. Chem. 285, 961–968 (2010).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P.R. & Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Afonine, P.V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
The PyMOL molecular graphics system, version 1.8. (Schrödinger LLC, 2015).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Dolinsky, T.J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005).
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010).
Celniker, G. et al. ConSurf: using evolutionary data to raise testable hypotheses about protein function. Isr. J. Chem. 53, 199–206 (2013).
Brachmann, C.B. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Knop, M. et al. Epitope tagging of yeast genes using a PCR-based strategy: more tags and improved practical routines. Yeast 15, 963–972 (1999).
Wach, A., Brachat, A., Pöhlmann, R. & Philippsen, P. New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 10, 1793–1808 (1994).
Acknowledgements
We thank J. Kopp, C. Siegmann and G. Müller from the BZH-Cluster of Excellence:CellNetworks crystallization platform for protein crystallization, and acknowledge access to the beamlines at the European Synchrotron Radiation Facility (ESRF) in Grenoble and the support of the beamline scientists. We acknowledge A. Hendricks for expert technical assistance and K. Wild for discussions. We would like to thank E. Hurt, E. Thomson and M. Thoms (Heidelberg University Biochemistry Center) for supplying the C. thermophilum cDNA and S. cerevisiae strains, and G. Stier (Heidelberg University Biochemistry Center) for the modified pET24d and pET16b expression vectors and the SUMO protease. We thank R. Kityk and M. Mayer (Heidelberg University Molecular Biology Center) for providing purified DnaK, for advice for fluorescence anisotropy experiments and for discussions. This work was supported by the Deutsche Forschungs-gemeinschaft (DFG) (through FOR967, GRK1188 and the Leibniz programme) and by HBIGS fellowships (to G.V.G. and F.A.W.). I.S. is an investigator of the Cluster of Excellence:CellNetworks.
Author information
Authors and Affiliations
Contributions
F.A.W., A.G., G.V.G., K.L. and I.S. designed the experiments, analyzed the data and wrote the manuscript. F.A.W., A.G., G.V.G. and K.L. performed experiments. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
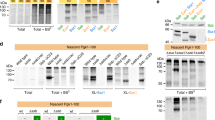
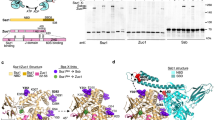
Supplementary Figure 1 Pulldown analysis of Ssz1 with Zuo1 or Zuo1N and purification of the Ssz1–Zuo1N complex for crystallization.
(a) MBP-pulldown analysis of Ssz1 with MBP-Zuo1 or MBP-Zuo1N and of the coexpressed Ssz1-MBP-Zuo1N complex. (b) Uncropped SDS-PAGE of Fig. 1c showing the SEC fractions of the Ssz1-Zuo1N complex.
Supplementary Figure 2 Packing of the two molecules in the asymmetric unit.
Within the two molecules (Ssz1: gray and purple, Zuo1N: orange; cartoon representation) in the asymmetric unit, several regions are disordered. The disordered regions are depicted as dotted lines and the residues numbered. The N-, C-terminus and domains are indicated. ATP is represented as sticks and Mg2+ is shown as a pink sphere.
Supplementary Figure 3 Electron density map of typical regions in the Ssz1–Zuo1N complex and B factor plot for the Ssz1–Zuo1N structure.
(a) Stereoview of the Zuo1N EPVG motif (orange, yellow sticks). The 2Fo-Fc electron density map is contoured at 1 σ. (b) Stereoview of the nucleotide bound to Ssz1. ATP (sticks), Mg2+ (green sphere) and water molecules (red spheres). The 2Fo-Fc electron density map is contoured at 2 σ. (c) Ribbon representation of the Ssz1-Zuo1N complex with Ssz1 (left) and Zuo1N (right) colored according to B-factors, scaled from 50 (blue) to 250 (red) Å2. ATP is represented as sticks. NBD and SBDβ are indicated.
Supplementary Figure 4 Surface analyses of Hsp70 proteins (with bound ATP).
Comparison of electrostatic surface potentials of (a) Ssz1, (b) Ssb1 (Gumiero, A. et al. Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain. Nat. Commun. 7,13563 (2016)) and (c) DnaK (Kityk, R., Kopp, J., Sinning, I. & Mayer, M.P. Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones. Mol. Cell 48, 863-74 (2012)). The structures have been superimposed on the NBD and are shown side by side. The positively charged patch observed in Ssz1 is absent in canonical Hsp70s. (d) Close-up view of the positively charged patch in Ssz1. Surface charge spans from -4 kT/e (deep red) to +4kT/e (deep blue). (e) and (f) Conservation surface mapping of the Ssz1 patch shown in (d) as reported by ConSurf (Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299-302 (2005)), from variable (cyan) to conserved (deep red). Figures were produced using PDB2PQR (Dolinsky, T.J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522-5 (2007); Dolinsky, T.J., Nielsen, J.E., McCammon, J.A. & Baker, N.A. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res. 32, W665-7 (2004)) and APBS (Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. U S A 98, 10037-41 (2001)) in PyMOL (Schrödinger, L.L.C. The PyMOL Molecular Graphics System, Version 1.8. (2015)).
Supplementary Figure 5 Comparison of Ssz1–Zuo1N and DnaK to show the different domain arrangement.
Ribbon representation of (a) the Ssz1 inter-domain linker (dark blue), Ssz1 SBDb (pale cyan), Zuo1N (orange), and (b)DnaK SBDβ (green, 4B9Q (Kityk, R., Kopp, J., Sinning, I. & Mayer, M.P. Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones. Mol. Cell 48, 863-74 (2012)). Both structures have been superimposed on the NBD. NBD and SBDβ are indicated and the NBDs are shown in surface representation (gray). SBDb binds to different lobes of the NBD.
Supplementary Figure 6 Superposition of the SBDs of Ssz1 (with bound Zuo1N) and DnaK (ADP, substrate).
Ssz1-Zuo1N and DnaK are shown in ribbon representation, with Zuo1N in orange and Ssz1 SBDβ in pale cyan, and DnaK in green (bound-substrate is in yellow; 1DKY (Zhu, X. et al. Structural analysis of substrate binding by the molecular chaperone DnaK. Science 272, 1606-14 (1996)). The Ssz1 specific insertion in β4–β5 would interfere with the lid closure observed in canonical Hsp70s.
Supplementary Figure 7 Comparison of the substrate binding domains from Ssz1, DnaK (closed and open) and Ssb.
(a) Schematic representation of the twisted β-sandwich, (b) ribbon representation of the structures of Ssz1 SBDβ–Zuo1N (pale cyan, orange), DnaK-ADP SBDβ (Zhu, X. et al. Structural analysis of substrate binding by the molecular chaperone DnaK. Science 272, 1606-14 (1996); green, sand) with bound substrate (yellow), DnaK-ATP SBDβ (Kityk, R., Kopp, J., Sinning, I. & Mayer, M.P. Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones. Mol. Cell 48, 863-74 (2012); green, sand), and Ssb1-ATP SBDβ (Gumiero, A. et al. Interaction of the cotranslational Hsp70 Ssb with ribosomal proteins and rRNA depends on its lid domain. Nat. Commun. 7, 13563 (2016); dark red). (c) Corresponding residues in the putative substrate-binding pocket of Ssz1 SBDβ and the SBDs of DnaK and Ssb1 are shown. The β-strands are labeled, the residue are shown as sticks.
Supplementary Figure 8 Binding assays of RAC to known Hsp70 substrates.
Fluorescence anisotropy binding assay of (a) EcDnaK, Ssz1 / Zuo1N or RAC with the NR peptide (NRLLLTG) (Schlecht, R., Erbse, A.H., Bukau, B. & Mayer, M.P. Mechanics of Hsp70 chaperones enables differential interaction with client proteins. Nat. Struct. Mol. Biol. 18, 345-51 (2011); Gragerov, A. & Gottesman, M.E. Different peptide binding specificities of hsp70 family members. J Mol Biol 241, 133-5 (1994)) and (b) CtSsb, Ssz1 / Zuo1N or RAC with the CoxIV peptide (Pfund, C. et al. The molecular chaperone Ssb from Saccharomyces cerevisiae is a component of the ribosome-nascent chain complex. EMBO J. 17, 3981-3989 (1998)). (c) Binding constants (KD) values for peptide substrates binding with EcDnaK or CtSsb. Each value is the average of three technical replicates ± standard deviation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Notes 1–3 (PDF 2265 kb)
Rights and permissions
About this article
Cite this article
Weyer, F., Gumiero, A., Gesé, G. et al. Structural insights into a unique Hsp70-Hsp40 interaction in the eukaryotic ribosome-associated complex. Nat Struct Mol Biol 24, 144–151 (2017). https://doi.org/10.1038/nsmb.3349
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3349
This article is cited by
-
Structural inventory of cotranslational protein folding by the eukaryotic RAC complex
Nature Structural & Molecular Biology (2023)
-
Structural remodeling of ribosome associated Hsp40-Hsp70 chaperones during co-translational folding
Nature Communications (2022)
-
Pathway of Hsp70 interactions at the ribosome
Nature Communications (2021)
-
A functional interaction between GRP78 and Zika virus E protein
Scientific Reports (2021)
-
Intracellular HSP70L1 inhibits human dendritic cell maturation by promoting suppressive H3K27me3 and H2AK119Ub1 histone modifications
Cellular & Molecular Immunology (2020)