Abstract
Ribonucleotides incorporated during DNA replication are removed by RNase H2–dependent ribonucleotide excision repair (RER). In RER-defective yeast, topoisomerase 1 (Top1) incises DNA at unrepaired ribonucleotides, initiating their removal, but this is accompanied by RNA-DNA–damage phenotypes. Here we show that these phenotypes are incurred by a high level of ribonucleotides incorporated by a leading strand–replicase variant, DNA polymerase (Pol) ɛ, but not by orthologous variants of the lagging-strand replicases, Pols α or δ. Moreover, loss of both RNases H1 and H2 is lethal in combination with increased ribonucleotide incorporation by Pol ɛ but not by Pols α or δ. Several explanations for this asymmetry are considered, including the idea that Top1 incision at ribonucleotides relieves torsional stress in the nascent leading strand but not in the nascent lagging strand, in which preexisting nicks prevent the accumulation of superhelical tension.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nick McElhinny, S.A. et al. Abundant ribonucleotide incorporation into DNA by yeast replicative polymerases. Proc. Natl. Acad. Sci. USA 107, 4949–4954 (2010).
Pursell, Z.F., Isoz, I., Lundstrom, E.B., Johansson, E. & Kunkel, T.A. Yeast DNA polymerase epsilon participates in leading-strand DNA replication. Science 317, 127–130 (2007).
Nick McElhinny, S.A., Gordenin, D.A., Stith, C.M., Burgers, P.M. & Kunkel, T.A. Division of labor at the eukaryotic replication fork. Mol. Cell 30, 137–144 (2008).
Wang, J. et al. Crystal structure of a pol alpha family replication DNA polymerase from bacteriophage RB69. Cell 89, 1087–1099 (1997).
Pavlov, Y.I., Shcherbakova, P.V. & Kunkel, T.A. In vivo consequences of putative active site mutations in yeast DNA polymerases alpha, epsilon, delta, and zeta. Genetics 159, 47–64 (2001).
Nick McElhinny, S.A. et al. Genome instability due to ribonucleotide incorporation into DNA. Nat. Chem. Biol. 6, 774–781 (2010).
Eder, P.S., Walder, R.Y. & Walder, J.A. Substrate specificity of human RNase H1 and its role in excision repair of ribose residues misincorporated in DNA. Biochimie 75, 123–126 (1993).
Rydberg, B. & Game, J. Excision of misincorporated ribonucleotides in DNA by RNase H (type 2) and FEN-1 in cell-free extracts. Proc. Natl. Acad. Sci. USA 99, 16654–16659 (2002).
Sparks, J.L. et al. RNase H2-initiated ribonucleotide excision repair. Mol. Cell 47, 980–986 (2012).
Lujan, S.A., Williams, J.S., Clausen, A.R., Clark, A.B. & Kunkel, T.A. Ribonucleotides are signals for mismatch repair of leading-strand replication errors. Mol. Cell 50, 437–443 (2013).
Miyabe, I., Kunkel, T.A. & Carr, A.M. The major roles of DNA polymerases epsilon and delta at the eukaryotic replication fork are evolutionarily conserved. PLoS Genet. 7, e1002407 (2011).
Lujan, S.A. et al. Mismatch repair balances leading and lagging strand DNA replication fidelity. PLoS Genet. 8, e1003016 (2012).
Williams, J.S. et al. Topoisomerase 1-mediated removal of ribonucleotides from nascent leading-strand DNA. Mol. Cell 49, 1010–1015 (2013).
Ghodgaonkar, M.M. et al. Ribonucleotides misincorporated into DNA act as strand-discrimination signals in eukaryotic mismatch repair. Mol. Cell 50, 323–332 (2013).
Vengrova, S. & Dalgaard, J.Z. The wild-type Schizosaccharomyces pombe mat1 imprint consists of two ribonucleotides. EMBO Rep. 7, 59–65 (2006).
Dalgaard, J.Z. Causes and consequences of ribonucleotide incorporation into nuclear DNA. Trends Genet. 28, 592–597 (2012).
Williams, J.S. & Kunkel, T.A. Ribonucleotides in DNA: origins, repair and consequences. DNA Repair (Amst.) 19, 27–37 (2014).
Caldecott, K.W. Ribose: an internal threat to DNA. Science 343, 260–261 (2014).
Tumbale, P., Williams, J.S., Schellenberg, M.J., Kunkel, T.A. & Williams, R.S. Aprataxin resolves adenylated RNA-DNA junctions to maintain genome integrity. Nature 506, 111–115 (2014).
Sekiguchi, J. & Shuman, S. Site-specific ribonuclease activity of eukaryotic DNA topoisomerase I. Mol. Cell 1, 89–97 (1997).
Kim, N. et al. Mutagenic processing of ribonucleotides in DNA by yeast topoisomerase I. Science 332, 1561–1564 (2011).
Clark, A.B., Lujan, S.A., Kissling, G.E. & Kunkel, T.A. Mismatch repair-independent tandem repeat sequence instability resulting from ribonucleotide incorporation by DNA polymerase epsilon. DNA Repair (Amst.) 10, 476–482 (2011).
Cho, J.E., Kim, N., Li, Y.C. & Jinks-Robertson, S. Two distinct mechanisms of Topoisomerase 1-dependent mutagenesis in yeast. DNA Repair (Amst.) 12, 205–211 (2013).
Lazzaro, F. et al. RNase H and postreplication repair protect cells from ribonucleotides incorporated in DNA. Mol. Cell 45, 99–110 (2012).
Cerritelli, S.M. & Crouch, R.J. Ribonuclease H: the enzymes in eukaryotes. FEBS J. 276, 1494–1505 (2009).
Chon, H. et al. RNase H2 roles in genome integrity revealed by unlinking its activities. Nucleic Acids Res. 41, 3130–3143 (2013).
Nick McElhinny, S.A., Kissling, G.E. & Kunkel, T.A. Differential correction of lagging-strand replication errors made by DNA polymerases α and δ. Proc. Natl. Acad. Sci. USA 107, 21070–21075 (2010).
Clausen, A.R. et al. Tracking replication enzymology in vivo by genome-wide mapping of ribonucleotide incorporation. Nat. Struct. Mol. Biol. 22, 185–191 (2015).
Smith, D.J. & Whitehouse, I. Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Nature 483, 434–438 (2012).
Burgers, P.M. Polymerase dynamics at the eukaryotic DNA replication fork. J. Biol. Chem. 284, 4041–4045 (2009).
Zheng, L. & Shen, B. Okazaki fragment maturation: nucleases take centre stage. J. Mol. Cell Biol. 3, 23–30 (2011).
Balakrishnan, L. & Bambara, R.A. Okazaki fragment metabolism. Cold Spring Harb. Perspect. Biol. 5, a010173 (2013).
Marteijn, J.A., Lans, H., Vermeulen, W. & Hoeijmakers, J.H. Understanding nucleotide excision repair and its roles in cancer and ageing. Nat. Rev. Mol. Cell Biol. 15, 465–481 (2014).
Lujan, S.A. et al. Heterogeneous polymerase fidelity and mismatch repair bias genome variation and composition. Genome Res. 24, 1751–1764 (2014).
Niimi, A. et al. Palm mutants in DNA polymerases alpha and eta alter DNA replication fidelity and translesion activity. Mol. Cell. Biol. 24, 2734–2746 (2004).
Garg, P., Stith, C.M., Sabouri, N., Johansson, E. & Burgers, P.M. Idling by DNA polymerase delta maintains a ligatable nick during lagging-strand DNA replication. Genes Dev. 18, 2764–2773 (2004).
Aguilera, A. & Klein, H.L. Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic characterization of hyper-recombination mutations. Genetics 119, 779–790 (1988).
Potenski, C.J., Niu, H., Sung, P. & Klein, H.L. Avoidance of ribonucleotide-induced mutations by RNase H2 and Srs2-Exo1 mechanisms. Nature 511, 251–254 (2014).
Williams, J.S. et al. Proofreading of ribonucleotides inserted into DNA by yeast DNA polymerase epsilon. DNA Repair (Amst.) 11, 649–656 (2012).
Wang, J.C. Cellular roles of DNA topoisomerases: a molecular perspective. Nat. Rev. Mol. Cell Biol. 3, 430–440 (2002).
Yu, H. & Droge, P. Replication-induced supercoiling: a neglected DNA transaction regulator? Trends Biochem. Sci. 39, 219–220 (2014).
Nilsen, H. et al. Uracil-DNA glycosylase (UNG)-deficient mice reveal a primary role of the enzyme during DNA replication. Mol. Cell 5, 1059–1065 (2000).
Bermejo, R. et al. Top1- and Top2-mediated topological transitions at replication forks ensure fork progression and stability and prevent DNA damage checkpoint activation. Genes Dev. 21, 1921–1936 (2007).
Gambus, A. et al. GINS maintains association of Cdc45 with MCM in replisome progression complexes at eukaryotic DNA replication forks. Nat. Cell Biol. 8, 358–366 (2006).
Georgescu, R.E. et al. Mechanism of asymmetric polymerase assembly at the eukaryotic replication fork. Nat. Struct. Mol. Biol. 21, 664–670 (2014).
Burgers, P.M. & Gerik, K.J. Structure and processivity of two forms of Saccharomyces cerevisiae DNA polymerase delta. J. Biol. Chem. 273, 19756–19762 (1998).
Shcherbakova, P.V. & Kunkel, T.A. Mutator phenotypes conferred by MLH1 overexpression and by heterozygosity for mlh1 mutations. Mol. Cell. Biol. 19, 3177–3183 (1999).
Pavlov, Y.I., Newlon, C.S. & Kunkel, T.A. Yeast origins establish a strand bias for replicational mutagenesis. Mol. Cell 10, 207–213 (2002).
Acknowledgements
We thank K. Bebenek, C. Orebaugh and S. Williams for helpful comments on the manuscript and all members of the Kunkel laboratory for thoughtful discussions. We acknowledge the US National Institute of Environmental Health Sciences (NIEHS) Molecular Genetics Core Facility for sequence analysis of 5-FOA–resistant mutants and the NIEHS Flow Cytometry Center for fluorescence-activated cell-sorting analysis. This work was supported by project Z01 ES065070 to T.A.K. from the Division of Intramural Research of the US National Institutes of Health (NIH), NIEHS, by the Swedish Cancer Society to A.C. and by US NIH grant GM032431 to P.M.B.
Author information
Authors and Affiliations
Contributions
J.S.W., A.R.C., S.A.L., L.M., A.B.C. and A.C. designed and performed experiments. P.M.B. provided reagents. All authors were involved in data analysis. J.S.W. and T.A.K. wrote the manuscript, with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
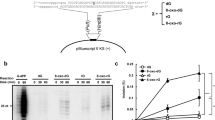
Supplementary Figure 1 Probing for ribonucleotides in nascent leading-strand DNA.
(a) The annealing locations of radiolabeled leading strand-specific probes on the URA3 reporter in OR1 or OR2. (b) Detection of alkali-sensitive sites in nascent leading strand DNA synthesized by Pol ɛ was performed as described13. Smaller DNA fragments observed for the α-LM rnh201Δ and δ-LM rnh201Δ mutants when using probes that anneal to the nascent leading strand (lanes 4, 6, 12 and 14) may be related to the close proximity of URA3 to ARS306 (1.6 kb). These alkali-sensitive sites may arise during ribonucleotide incorporation by Pols α or δ into the nascent lagging strand during bidirectional synthesis proceeding from this origin in the opposite direction (to the left of the origin in panel a). In addition, these small fragments that hybridize to the ‘leading strand’ probe may be generated by synthesis performed by L868M Pol α or L612M Pol δ as they replicate from the adjacent ARS307 origin. The same explanation applies for small DNA fragments observed for the ɛ-MG rnh201Δ mutant in Figure 2 (main text) when using probes that anneal to the nascent lagging strand (lanes 8 and 16). (c) The average fraction of alkali-sensitive fragments along the membrane was determined by quantifying the radioactive signal using data from in panel b (for both Probes A and B) from two independent experiments.
Supplementary Figure 2 Workflow for calculating fractional replication-strand bias from HydEn-seq end counts without background subtraction or internal standards.
Each set of arrows indicates the application of the equation shown to the left. Gray data points represent bins of 20 base pairs (bp). Labels above each graph are relate the variables in the equations to the values for which they are color-coded. Solid lines are moving averages over 25 bins (500 bp). (a) HydEn-seq end counts for reads mapping to the forward (black) and reverse (orange) strands from RNase H2-deficient pol1-L868M (α-LM; left), pol2-M644G (ɛ-MG; middle), and pol3-L612M (δ-LM; right) strains. (b) The observed strand bias, expressed as a log-ratio between forward- and reverse-strand end counts. (c) An approximation of the true replication strand bias log-ratio. Extra-replicative contributions were removed by comparing strains with opposite biases. The greater the biases in the chosen strains, the closer the approximation. (d) The fraction of replication events in which the bottom strand originates as the nascent lagging strand.
Supplementary Figure 3 Alkali-sensitive Okazaki fragment–sized DNA is not observed in the α-LM rnh201Δ or δ-LM rnh201D strains.
Purified genomic DNA was subjected to alkaline hydrolysis, alkaline-agarose electrophoresis and stained with SYBR® Gold Nucleic Acid Gel Stain, a sensitive fluorescent stain that can be used for detection of single strand DNA. The fraction of alkali-sensitive fragments was calculated by dividing the fluorescence intensity (arbitrary units) at each position along the gel by the total intensity for each lane. The experiment was performed in duplicate, and a representative gel image and quantitation is displayed. We note that Okazaki fragment-sized DNA (e.g. 150-300 base pair) fragments were not observed among the products of the alkaline hydrolysis of genomic DNA from the α-LM rnh201Δ or δ-LM rnh201Δ strains.
Supplementary Figure 4 Asymmetric phenotypes are associated with unrepaired ribonucleotides incorporated into nascent leading- versus lagging-strand DNA.
(a) Cell cycle progression is not affected by loss of RNH201 in the pol1-LM or the pol3-LM mutator strains with an increased number of unrepaired ribonucleotides in the nascent lagging strand. Cells were grown to mid-log phase at 30°C and processed for flow cytometry as described in13. The experiment was performed in duplicate and data are displayed as the mean % ± standard error. (b) The increase in total dNTP abundance conferred by loss of RNH201 is not significantly enhanced in the pol1-LM or the pol3-LM lagging strand polymerase mutator strains. Data for total dNTP abundance (dCTP, dATP, dTTP and dGTP) is displayed as the mean ± standard error. Each strain genotype was independently analyzed twice.
Supplementary Figure 5 URA3 mutation spectra for pol1 -L868M rnh201 Δ and pol3-L612M rnh201 Δ.
The coding strand of the 804 base pair URA3 open reading frame is shown. The sequence changes observed in independent ura3 mutants are depicted above the coding sequence for pol1-L868M rnh201∆ in red and below the coding sequence for pol3-L612M rnh201∆ in blue. Letters indicate single base substitutions, open triangles indicate single base deletions, and short lines above or below the coding sequence indicate multibase base deletions of between 2 and 5 base pairs. All strains had the URA3 reporter in OR2.
Supplementary information
Supplementary Figures and Text
Supplementary Figures 1–5 and Supplementary Tables 1–3 (PDF 5273 kb)
Supplementary Data Set 1
Original gel images for Figures 1–3 (PDF 28958 kb)
Rights and permissions
About this article
Cite this article
Williams, J., Clausen, A., Lujan, S. et al. Evidence that processing of ribonucleotides in DNA by topoisomerase 1 is leading-strand specific. Nat Struct Mol Biol 22, 291–297 (2015). https://doi.org/10.1038/nsmb.2989
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2989
This article is cited by
-
High-fidelity DNA ligation enforces accurate Okazaki fragment maturation during DNA replication
Nature Communications (2021)
-
RNases H1 and H2: guardians of the stability of the nuclear genome when supply of dNTPs is limiting for DNA synthesis
Current Genetics (2020)
-
Roles for DNA polymerase δ in initiating and terminating leading strand DNA replication
Nature Communications (2019)
-
Evidence that DNA polymerase δ contributes to initiating leading strand DNA replication in Saccharomyces cerevisiae
Nature Communications (2018)
-
Genome-wide mapping of embedded ribonucleotides and other noncanonical nucleotides using emRiboSeq and EndoSeq
Nature Protocols (2015)