Abstract
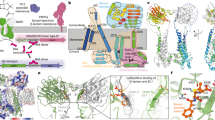
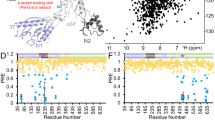
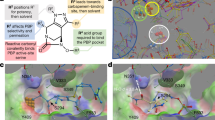
The multiple antibiotic resistance of methicillin-resistant strains of Staphylococcus aureus (MRSA) has become a major clinical problem worldwide. The key determinant of the broad-spectrum β-lactam resistance in MRSA strains is the penicillin-binding protein 2a (PBP2a). Because of its low affinity for β-lactams, PBP2a provides transpeptidase activity to allow cell wall synthesis at β-lactam concentrations that inhibit the β-lactam-sensitive PBPs normally produced by S. aureus. The crystal structure of a soluble derivative of PBP2a has been determined to 1.8 Å resolution and provides the highest resolution structure for a high molecular mass PBP. Additionally, structures of the acyl-PBP complexes of PBP2a with nitrocefin, penicillin G and methicillin allow, for the first time, a comparison of an apo and acylated resistant PBP. An analysis of the PBP2a active site in these forms reveals the structural basis of its resistance and identifies features in newly developed β-lactams that are likely important for high affinity binding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Diekema, D.J. et al. Clin. Infect. Dis. 32, S114–S132 (2001).
Hiramatsu, K. et al. Lancet 350, 1670–1673 (1997).
Hiramatsu, K., Cui, L., Kuroda, M. & Ito, T. Trends Microbiol. 9, 486–493 (2001).
Hiramatsu, K. Microbiol. Immunol. 39, 531–543 (1995).
Chambers, H.F. Clin. Microbiol. Rev. 10, 781–791 (1997).
Ghuysen, J.-M. Int. J. Antimicrob. Agents 8, 45–60 (1997).
Kuroda, M. et al. Lancet 357, 1225–1240 (2001).
Goffin, C. & Ghuysen, J.-M. Microbiol. Mol. Biol. Rev. 62, 1079–1093 (1998).
Hartman, B.J. & Tomasz, A. J. Bacteriol. 158, 513–516 (1984).
Reynolds, P.E. & Brown, D.F. FEBS Lett. 192, 28–32 (1985).
Pinho, M.G., Filipe, S.R., De Lencastre, H. & Tomasz, A. J. Bacteriol. 183, 6525–6531 (2001).
Wu, C.Y., Hoskins, J., Blaszczak, L.C., Preston, D.A. & Skatrud, P.L. Antimicrob. Agents Chemother. 36, 533–539 (1992).
Lu, W.-P. et al. Biochemistry 38, 6537–6546 (1999).
Hendrickson, W.A. Science 254, 51–58 (1991).
Holm, L. & Sander, C. J. Mol. Biol. 233, 123–138 (1993).
Chen, J.M., Xu, S.L., Wawrzak, Z., Basarab, G.S. & Jordan, D.B. Biochemistry 37, 17735–17744 (1998).
Lundqvist, T. et al. Structure 2, 937–944 (1994).
Fribourg, S., Braun, I.C., Izaurralde, E. & Conti, E. Mol. Cell 8, 645–656 (2001).
Wu, Z.R. et al. Science 276, 415–418 (1997).
Massova, I. & Mobashery, S. Antimicrob. Agents Chemother. 42, 1–17 (1998).
Paetzel, M. et al. Nature Struct. Biol. 7, 918–925 (2000).
Pares, S., Mouz, N., Petillot, Y., Hakenbeck, R. & Dideberg, O. Nature Struct. Biol. 3, 284–289 (1996).
Gordon, E., Mouz, N., Duee, E. & Dideberg, O. J. Mol. Biol. 299, 477–485 (2000).
Kelly, J.A. et al. J. Biol. Chem. 260, 6449–6458 (1985).
Fonze, E., Vermeire, M., Nguyen-Disteche, M., Brasseur, R. & Charlier, P. J. Biol. Chem. 274, 21853–21860 (1999).
Davies, C., White, S.W. & Nicholas, R.A. J. Biol. Chem. 276, 616–623 (2001).
Strynadka, N.C.J. et al. Nature, 359, 700–705 (1992).
Ishiguro, M. & Imajo, S. J. Med. Chem. 39, 2207–2218 (1996).
Frere, J.M., Ghuysen, J.M. & Iwatsubo, M. Eur. J. Biochem. 57, 343–351 (1975).
Ghuysen, J.M., Frere, J.M., Leyh-Bouille, M., Nguyen-Disteche, M. & Coyette, J. Biochem J. 235, 159–165 (1986).
Lu, W.P., Kincaid, E., Sun, Y., Bauer, M.D. J. Biol. Chem. 276, 31494–31501 (2001).
de Jonge, B.L.M. & Tomasz, A. Antimicrob. Agents Chemother. 37, 342–346 (1993).
Kondo, N., Kuwahara-Arai, K., Kuroda-Murakami, H., Tateda-Suzuki, E. & Hiramatsu, K. Antimicrob. Agents Chemother. 45, 815–824 (2001).
Mouz, N. et al. Proc. Natl. Acad. Sci. USA 95, 13403–13406 (1998).
Dessen, A., Mouz, N., Gordon, E., Hopkins, J. & Dideberg, O. J. Biol. Chem. 276, 45106–45112 (2001).
Graves-Woodward, K. & Pratt, R.F. Biochem J. 332, 755–761 (1998).
Pratt, R.F., Krishnaraj, R. & Xu, H. Biochem. J. 286, 857–862 (1992).
Entenza, J.M., Hohl, P., Heinze-Krauss, I., Glauser, M.P. & Moreillon, P. Antimicrob. Agents Chemother. 46, 171–177 (2002).
Fung-Tomc, J.C. et al. Antimicrob. Agents Chemother. 46, 971–976 (2002).
Toney, J.H. et al. Anal. Biochem. 255, 113–119 (1998).
Roychoudhury, S., Dotzlaf, J.E., Ghag, S. & Yeh, W.K. J. Biol. Chem. 269, 12067–12073 (1994).
Frank, L.J. et al. Prot. Express. Purif. 6, 671–678 (1995).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–325 (1997).
Collaborative Computational Project, Number 4. Acta Crystallogr. D. 50, 760–763 (1994).
Weeks, C.M. & Miller, R. Acta Crystallogr. D 55, 492–500 (1999).
Terwilliger, T.C. & Berendzen J. Acta Crystallogr. D 55, 849–861 (1999).
de La Fortelle, E. & Bricogne, G. Methods Enzymol. 276, 472–494 (1997).
Terwilliger, T.C. Acta Crystallogr. D 56, 965–972 (2000).
McRee, D.E. J. Mol. Graph. 10, 44–47 (1992).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Cohen, G.E. J. Appl. Crystallogr. 30, 1160–1161 (1997).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Acta Crystallogr. D 50, 869–873 (1994).
Read, R.J. Acta Crystallogr. A 42, 140–149 (1986).
Nicholls, A., Bharadwaj, R. & Honig, B. Biophys. J. 64, A166–A166 (1993).
Sauvage, E. et al. Cell. Mol. Life Sci. 59, 1223–1232 (2002).
Acknowledgements
We thank L.C. Blaszczak and P.L. Skatrud (Infectious Disease Research, Eli Lilly and Company) for providing the initial MRSA strain 27r PBP2a* construct, J. Berendzen and L. Flaks for assistance during data collection at beam line X8-C at the NSLS, S.-M. He for ESMS analysis, Y. Luo and L. De Castro for technical assistance, and M. Bains for DNA sequencing. N.C.J.S. is supported by the Canadian Institutes of Health Research (CIHR), the Howard Hughes Medical Institute (HHMI) and the Canadian Bacterial Diseases Network. N.C.J.S. is a CIHR Investigator, a Burroughs Wellcome New Investigator and a HHMI International Scholar. D.L. is the recipient of a CIHR doctoral research award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lim, D., Strynadka, N. Structural basis for the β lactam resistance of PBP2a from methicillin-resistant Staphylococcus aureus. Nat Struct Mol Biol 9, 870–876 (2002). https://doi.org/10.1038/nsb858
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb858
This article is cited by
-
Computational modelling of the antimicrobial peptides Cruzioseptin-4 extracted from the frog Cruziohyla calcarifer and Pictuseptin-1 extracted from the frog Boana picturata
Scientific Reports (2024)
-
Cloning, Purification, and Biophysical Characterization of FemB Protein from Methicillin-Resistant Staphylococcus aureus and Inhibitors Screening
Applied Biochemistry and Biotechnology (2023)
-
Graphene oxide-ZnO nanorods for efficient dye degradation, antibacterial and in-silico analysis
Applied Nanoscience (2022)
-
Mechanism of drug resistance in bacteria: efflux pump modulation for designing of new antibiotic enhancers
Folia Microbiologica (2021)
-
Performance of the Hologic Panther Fusion® MRSA Assay for the nasal screening of methicillin-sensitive and methicillin-resistant Staphylococcus aureus carriage
European Journal of Clinical Microbiology & Infectious Diseases (2020)