Abstract
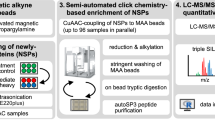
Regulation of mRNA translation has a pivotal role in modulating protein levels, and the genome-wide identification of proteins synthesized at a given time is indispensable to our understanding of gene expression. This protocol describes the mass-spectrometric analysis of newly synthesized proteins from cultured cells or whole tissues by using a biotinylated derivative of puromycin, which becomes incorporated into nascent polypeptide chains by ribosome catalysis. In this method, termed puromycin-associated nascent chain proteomics (PUNCH-P), intact ribosome-nascent chain complexes are first recovered from cells by ultracentrifugation, followed by biotin-puromycin labeling of newly synthesized proteins, streptavidin affinity purification and liquid chromatography–tandem mass spectrometry (LC-MS/MS). Unlike methods that require in vivo labeling, the sensitivity and coverage of PUNCH-P depend only on the amount of starting material and not on the duration of labeling, thus enabling the measurement of rapid fluctuations in protein synthesis. The protocol requires 3 d for sample preparation and analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Calkhoven, C.F., Muller, C. & Leutz, A. Translational control of gene expression and disease. Trends Mol. Med. 8, 577–583 (2002).
Holcik, M. & Sonenberg, N. Translational control in stress and apoptosis. Nat. Rev. Mol. Cell Biol. 6, 318–327 (2005).
Vogel, C. & Marcotte, E.M. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 13, 227–232 (2012).
Ingolia, N.T., Ghaemmaghami, S., Newman, J.R. & Weissman, J.S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Schwanhausser, B., Gossen, M., Dittmar, G. & Selbach, M. Global analysis of cellular protein translation by pulsed SILAC. Proteomics 9, 205–209 (2009).
Dieterich, D.C., Link, A.J., Graumann, J., Tirrell, D.A. & Schuman, E.M. Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc. Natl. Acad. Sci. USA 103, 9482–9487 (2006).
Howden, A.J. et al. QuaNCAT: quantitating proteome dynamics in primary cells. Nat. Methods 10, 343–346 (2013).
Aviner, R., Geiger, T. & Elroy-Stein, O. Novel proteomic approach (PUNCH-P) reveals cell cycle-specific fluctuations in mRNA translation. Genes Dev. 27, 1834–1844 (2013).
Pestka, S. Inhibitors of ribosome functions. Annu. Rev. Microbiol. 25, 487–562 (1971).
Baliga, B.S., Cohen, S.A. & Munro, H.N. Effect of cycloheximide on the reaction of puromycin with polysome-bound peptidyl-tRNA. FEBS Lett. 8, 249–252 (1970).
David, A. et al. Nuclear translation visualized by ribosome-bound nascent chain puromycylation. J. Cell Biol. 197, 45–57 (2012).
Kramer, G. et al. Identification and quantitation of newly synthesized proteins in Escherichia coli by enrichment of azidohomoalanine-labeled peptides with diagonal chromatography. Mol. Cell Proteomics 8, 1599–1611 (2009).
Vaughan, M.H. Jr., Pawlowski, P.J. & Forchhammer, J. Regulation of protein synthesis initiation in HeLa cells deprived of single essential amino acids. Proc. Natl. Acad. Sci. USA 68, 2057–2061 (1971).
Schwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Liu, J., Xu, Y., Stoleru, D. & Salic, A. Imaging protein synthesis in cells and tissues with an alkyne analog of puromycin. Proc. Natl. Acad. Sci. USA 109, 413–418 (2012).
Schmidt, E.K., Clavarino, G., Ceppi, M. & Pierre, P. SUnSET, a nonradioactive method to monitor protein synthesis. Nat. Methods 6, 275–277 (2009).
Starck, S.R., Green, H.M., Alberola-Ila, J. & Roberts, R.W. A general approach to detect protein expression in vivo using fluorescent puromycin conjugates. Chem. Biol. 11, 999–1008 (2004).
Croons, V., Martinet, W., Herman, A.G. & De Meyer, G.R. Differential effect of the protein synthesis inhibitors puromycin and cycloheximide on vascular smooth muscle cell viability. J. Pharmacol. Exp. Ther. 325, 824–832 (2008).
Wharton, S.A. & Hipkiss, A.R. Abnormal proteins of shortened length are preferentially degraded in the cytosol of cultured MRC5 fibroblasts. FEBS Lett. 168, 134–138 (1984).
Dissous, C., Caner, F. & Krembel, J. Isopycnic centrifugation of free and membrane-bound polysomes from rat liver. Eur. J. Biochem. 44, 161–170 (1974).
Tagwerker, C. et al. A tandem affinity tag for two-step purification under fully denaturing conditions: application in ubiquitin profiling and protein complex identification combined with in vivo cross-linking. Mol. Cell Proteomics 5, 737–748 (2006).
Holmberg, A. et al. The biotin-streptavidin interaction can be reversibly broken using water at elevated temperatures. Electrophoresis 26, 501–510 (2005).
Ong, S.E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat. Protoc. 1, 2650–2660 (2006).
Scopes, R.K. & Smith, J.A. Analysis of proteins. Curr. Protoc. Mol. Biol. 90, 10.0.1–10.0.23 (2010).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Acknowledgements
O.E.-S. acknowledges support from the Israel Science Foundation (grant 1036/12) and The Legacy Heritage Bio-Medical Program of the Israel Science Foundation (grant no. 1629/13). T.G. acknowledges support from the Israel Science Foundation (grant 1617/12) and the Israeli Centers of Research Excellence (I-CORE), Gene Regulation in Complex Human Disease, Center no. 41/11.
Author information
Authors and Affiliations
Contributions
R.A., T.G. and O.E.-S. designed the experiments. R.A. performed the biochemical experiments. T.G. performed the mass spectrometry experiments and data analysis. R.A., T.G. and O.E.-S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Comparison of puromycin and biotin-puromycin labeling efficiencies under different conditions.
Intact cultured HeLa cells, lysed cells or isolated polysomes (denoted ‘In-culture’, ‘In-lysate’ or ‘On-polysome’, respectively) were treated with 1μM puromycin (Puro) or 1 μM biotin-puromycin (Biot-PU) for 30 min at 37 °C or left untreated. 20 μg of total protein from each sample were resolved on a 12% SDS-PAGE; the membrane was stained with Ponceau S as loading control and probed sequentially with a puromycin antibody (clone 12D10) and streptavidin-HRP. Note that the puromycin antibody detects only underivatized puromycin and not biotin-puromycin, probably due to epitope masking by biotin. The dashed red box delineates the advantage of ribosome purification for high labeling efficiency using biotin-puromycin.
Supplementary information
Supplementary Figure 1
Comparison of puromycin and biotin-puromycin labeling efficiencies under different conditions.. (PDF 252 kb)
Supplementary Figure 2
Densitometry quantification related to Figure 2. (PDF 186 kb)
Rights and permissions
About this article
Cite this article
Aviner, R., Geiger, T. & Elroy-Stein, O. Genome-wide identification and quantification of protein synthesis in cultured cells and whole tissues by puromycin-associated nascent chain proteomics (PUNCH-P). Nat Protoc 9, 751–760 (2014). https://doi.org/10.1038/nprot.2014.051
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.051
This article is cited by
-
Utilities of Isolated Nerve Terminals in Ex Vivo Analyses of Protein Translation in (Patho)physiological Brain States: Focus on Alzheimer’s Disease
Molecular Neurobiology (2024)
-
Nascent Ribo-Seq measures ribosomal loading time and reveals kinetic impact on ribosome density
Nature Methods (2021)
-
Axonal mRNA localization and translation: local events with broad roles
Cellular and Molecular Life Sciences (2021)
-
Huntington’s Chorea—a Rare Neurodegenerative Autosomal Dominant Disease: Insight into Molecular Genetics, Prognosis and Diagnosis
Applied Biochemistry and Biotechnology (2021)
-
Cysteinyl-tRNA synthetase governs cysteine polysulfidation and mitochondrial bioenergetics
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.