Abstract
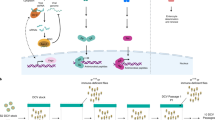
Insects rely solely on innate immune responses to combat a wide array of pathogens. With its powerful genetics, drosophila has proven especially powerful for the study of humoral innate immunity, characterized by the rapid induction of antimicrobial peptides. The two signaling pathways involved, Toll and Imd, have been studied intensely, but other aspects of the drosophila immune response are less well understood. A flurry of reports has focused on the mechanisms of phagocytosis, antiviral immunity and viral pathogenesis in drosophila. These studies have taken advantage of genome-wide RNA-mediated interference screening in drosophila cells, as well as more traditional genetic tools available in the fly. This review discusses advances in these exciting new areas of drosophila immunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Royet, J., Reichhart, J.M. & Hoffmann, J.A. Sensing and signaling during infection in Drosophila. Curr. Opin. Immunol. 17, 11–17 (2005).
Kaneko, T. & Silverman, N. Bacterial recognition and signalling by the Drosophila IMD pathway. Cell. Microbiol. 7, 461–469 (2005).
Kaneko, T. et al. PGRP-LC and PGRP-LE have essential yet distinct functions in the drosophila immune response to monomeric DAP-type peptidoglycan. Nat. Immunol. 7, 715–723 (2006).
Stuart, L.M. & Ezekowitz, R.A. Phagocytosis: elegant complexity. Immunity 22, 539–550 (2005).
Pearson, A.M. et al. Identification of cytoskeletal regulatory proteins required for efficient phagocytosis in Drosophila. Microbes Infect. 5, 815–824 (2003).
Philips, J.A., Rubin, E.J. & Perrimon, N. Drosophila RNAi screen reveals CD36 family member required for mycobacterial infection. Science 309, 1251–1253 (2005).
Agaisse, H. et al. Genome-wide RNAi screen for host factors required for intracellular bacterial infection. Science 309, 1248–1251 (2005).
Stroschein-Stevenson, S.L., Foley, E., O'Farrell, P.H. & Johnson, A.D. Identification of Drosophila gene products required for phagocytosis of Candida albicans. PLoS Biol. 4, e4 (2006).
Franc, N.C., Heitzler, P., Ezekowitz, R.A. & White, K. Requirement for croquemort in phagocytosis of apoptotic cells in Drosophila. Science 284, 1991–1994 (1999).
Hoebe, K. et al. CD36 is a sensor of diacylglycerides. Nature 433, 523–527 (2005).
Savill, J., Hogg, N., Ren, Y. & Haslett, C. Thrombospondin cooperates with CD36 and the vitronectin receptor in macrophage recognition of neutrophils undergoing apoptosis. J. Clin. Invest. 90, 1513–1522 (1992).
Ramet, M. et al. Drosophila scavenger receptor CI is a pattern recognition receptor for bacteria. Immunity 15, 1027–1038 (2001).
Kocks, C. et al. Eater, a transmembrane protein mediating phagocytosis of bacterial pathogens in Drosophila. Cell 123, 335–346 (2005).
Ulvila, J. et al. Double stranded RNA is internalized by scavenger receptor mediated endocytosis in Drosophila S2 cells. J. Biol. Chem. 281, 14370–14375 (2006).
Schmucker, D. & Flanagan, J.G. Generation of recognition diversity in the nervous system. Neuron 44, 219–222 (2004).
Watson, F.L. et al. Extensive diversity of Ig-superfamily proteins in the immune system of insects. Science 309, 1874–1878 (2005).
Dong, Y., Taylor, H.E. & Dimopoulos, G. AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system. PLoS Biol. 4, e229 (2006).
Levashina, E.A. et al. Conserved role of a complement-like protein in phagocytosis revealed by dsRNA knockout in cultured cells of the mosquito, Anopheles gambiae. Cell 104, 709–718 (2001).
Blandin, S. et al. Complement-like protein TEP1 is a determinant of vectorial capacity in the malaria vector Anopheles gambiae. Cell 116, 661–670 (2004).
Garver, L.S., Wu, J. & Wu, L.P. The peptidoglycan recognition protein PGRP-SC1a is essential for Toll signaling and phagocytosis of Staphylococcus aureus in Drosophila. Proc. Natl. Acad. Sci. USA 103, 660–665 (2006).
Kawai, T. & Akira, S. Innate immune recognition of viral infection. Nat. Immunol. 7, 131–137 (2006).
Akira, S., Uematsu, S. & Takeuchi, O. Pathogen recognition and innate immunity. Cell 124, 783–801 (2006).
Hiscott, J., Lin, R., Nakhaei, P. & Paz, S. MasterCARD: a priceless link to innate immunity. Trends Mol. Med. 12, 53–56 (2006).
Clemens, M.J. Translational control in virus-infected cells: models for cellular stress responses. Semin. Cell Dev. Biol. 16, 13–20 (2005).
Li, H.W. & Ding, S.W. Antiviral silencing in animals. FEBS Lett. 579, 5965–5973 (2005).
Grumbling, G. & Strelets, V. FlyBase: anatomical data, images and queries. Nucleic Acids Res. 34, D484–D488 (2006).
Meister, G. & Tuschl, T. Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343–349 (2004).
Haasnoot, J. & Berkhout, B. RNA interference: its use as antiviral therapy. Handb. Exp. Pharmacol., 173, 117–150 (2006).
Miyano-Kurosaki, N. & Takaku, H. Gene silencing of virus replication by RNA interference. Handb. Exp. Pharmacol. 173, 151–171 (2006).
Xie, Q. & Guo, H.S. Systemic antiviral silencing in plants. Virus Res. 118, 1–6 (2005).
Li, W.X. et al. Interferon antagonist proteins of influenza and vaccinia viruses are suppressors of RNA silencing. Proc. Natl. Acad. Sci. USA 101, 1350–1355 (2004).
Li, H., Li, W.X. & Ding, S.W. Induction and suppression of RNA silencing by an animal virus. Science 296, 1319–1321 (2002).
Galiana-Arnoux, D., Dostert, C., Schneemann, A., Hoffmann, J.A. & Imler, J.L. Essential function in vivo for Dicer-2 in host defense against RNA viruses in drosophila. Nat. Immunol. 7, 590–597 (2006).
Wang, X.H. et al. RNA interference directs innate immunity against viruses in adult Drosophila. Science 312, 452–454 (2006).
Zambon, R.A., Vakharia, V.N. & Wu, L.P. RNAi is an antiviral immune response against a dsRNA virus in Drosophila melanogaster. Cell. Microbiol. 8, 880–889 (2006).
Pal-Bhadra, M., Bhadra, U. & Birchler, J.A. RNAi related mechanisms affect both transcriptional and posttranscriptional transgene silencing in Drosophila. Mol. Cell 9, 315–327 (2002).
Sarot, E., Payen-Groschene, G., Bucheton, A. & Pelisson, A. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics 166, 1313–1321 (2004).
Kalmykova, A.I., Klenov, M.S. & Gvozdev, V.A. Argonaute protein PIWI controls mobilization of retrotransposons in the Drosophila male germline. Nucleic Acids Res. 33, 2052–2059 (2005).
Berkhout, B. & Haasnoot, J. The interplay between virus infection and the cellular RNA interference machinery. FEBS Lett. 580, 2896–2907 (2006).
Reavy, B., Dawson, S., Canto, T. & MacFarlane, S.A. Heterologous expression of plant virus genes that suppress post-transcriptional gene silencing results in suppression of RNA interference in Drosophila cells. BMC Biotechnol. 4, 18 (2004).
Obbard, D.J., Jiggins, F.M., Halligan, D.L. & Little, T.J. Natural selection drives extremely rapid evolution in antiviral RNAi genes. Curr. Biol. 16, 580–585 (2006).
Brun, G. & Plus, N. in Genetics and Biology of Drosophila (eds. Ashburner, M. & Wright, T.R.F.) 625–702 (Academic, New York, 1980).
Wayne, M.L., Contamine, D. & Kreitman, M. Molecular population genetics of ref(2)P, a locus which confers viral resistance in Drosophila. Mol. Biol. Evol. 13, 191–199 (1996).
Zambon, R.A., Nandakumar, M., Vakharia, V.N. & Wu, L.P. The Toll pathway is important for an antiviral response in Drosophila. Proc. Natl. Acad. Sci. USA 102, 7257–7262 (2005).
Thoetkiattikul, H., Beck, M.H. & Strand, M.R. Inhibitor κB-like proteins from a polydnavirus inhibit NF-κB activation and suppress the insect immune response. Proc. Natl. Acad. Sci. USA 102, 11426–11431 (2005).
Sabatier, L. et al. Pherokine-2 and -3. Eur. J. Biochem. 270, 3398–3407 (2003).
Roxstrom-Lindquist, K., Terenius, O. & Faye, I. Parasite-specific immune response in adult Drosophila melanogaster: a genomic study. EMBO Rep. 5, 207–212 (2004).
Dostert, C. et al. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of drosophila. Nat. Immunol. 6, 946–953 (2005).
Garcia-Sastre, A. & Biron, C.A. Type 1 interferons and the virus-host relationship: a lesson in detente. Science 312, 879–882 (2006).
Brun, S. et al. The MAPKKK Mekk1 regulates the expression of Turandot stress genes in response to septic injury in Drosophila. Genes Cells 11, 397–407 (2006).
Ekengren, S. & Hultmark, D. A family of Turandot-related genes in the humoral stress response of Drosophila. Biochem. Biophys. Res. Commun. 284, 998–1003 (2001).
Cherry, S. & Perrimon, N. Entry is a rate-limiting step for viral infection in a Drosophila melanogaster model of pathogenesis. Nat. Immunol. 5, 81–87 (2004).
Cherry, S. et al. Genome-wide RNAi screen reveals a specific sensitivity of IRES-containing RNA viruses to host translation inhibition. Genes Dev. 19, 445–452 (2005).
Dean, M. et al. Genetic restriction of HIV-1 infection and progression to AIDS by a deletion allele of the CKR5 structural gene. Hemophilia Growth and Development Study, Multicenter AIDS Cohort Study, Multicenter Hemophilia Cohort Study, San Francisco City Cohort, ALIVE Study. Science 273, 1856–1862 (1996).
Liu, R. et al. Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell 86, 367–377 (1996).
Moore, J.P. & Doms, R.W. The entry of entry inhibitors: a fusion of science and medicine. Proc. Natl. Acad. Sci. USA 100, 10598–10602 (2003).
Gwack, Y. et al. A genome-wide Drosophila RNAi screen identifies DYRK-family kinases as regulators of NFAT. Nature 441, 646–650 (2006).
Feske, S. et al. A mutation in Orai1 causes immune deficiency by abrogating CRAC channel function. Nature 441, 179–185 (2006).
Acknowledgements
We thank R. Doms, M. Tudor, E. Lien and members of the Silverman lab for comments and insights; and B. Graveley for the Dscam figure (Fig. 3). Supported by the National Institutes of Health (AI060025 to N.S.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Cherry, S., Silverman, N. Host-pathogen interactions in drosophila: new tricks from an old friend. Nat Immunol 7, 911–917 (2006). https://doi.org/10.1038/ni1388
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1388
This article is cited by
-
Lipopolysaccharide induced altered signaling pathways in various neurological disorders
Naunyn-Schmiedeberg's Archives of Pharmacology (2022)
-
Insect antimicrobial peptides: potential weapons to counteract the antibiotic resistance
Cellular and Molecular Life Sciences (2021)
-
Serine protease SP105 activates prophenoloxidase in Asian corn borer melanization, and is regulated by serpin-3
Scientific Reports (2017)
-
Symbiosis with Francisella tularensis provides resistance to pathogens in the silkworm
Scientific Reports (2016)
-
The genome- and transcriptome-wide analysis of innate immunity in the brown planthopper, Nilaparvata lugens
BMC Genomics (2013)