Abstract
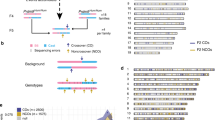
There is considerable interest in understanding patterns of linkage disequilibrium (LD) in the human genome, to aid investigations of human evolution and facilitate association studies in complex disease1,2,3,4,5. The relative influences of meiotic crossover distribution and population history on LD remain unclear, however5. In particular, it is uncertain to what extent crossovers are clustered into 'hot spots,6,7,8 that might influence LD patterns. As a first step to investigating the relationship between LD and recombination, we have analyzed a 216-kb segment of the class II region of the major histocompatibility complex (MHC) already characterized for familial crossovers9. High-resolution LD analysis shows the existence of extended domains of strong association interrupted by patchwork areas of LD breakdown. Sperm typing shows that these areas correspond precisely to meiotic crossover hot spots. All six hot spots defined share a remarkably similar symmetrical morphology but vary considerably in intensity, and are not obviously associated with any primary DNA sequence determinants of hot-spot activity. These hot spots occur in clusters and together account for almost all crossovers in this region of the MHC. These data show that, within the MHC at least, crossovers are far from randomly distributed at the molecular level and that recombination hot spots can profoundly affect LD patterns.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Collins, A., Lonjou, C. & Morton, N.E. Genetic epidemiology of single-nucleotide polymorphisms. Proc. Natl Acad. Sci. USA 96, 15173–15177 (1999).
Kruglyak, L. Prospects for whole-genome linkage disequilibrium mapping of common disease genes. Nature Genet. 22, 139–144 (1999).
Jorde, L.B. Linkage disequilibrium and the search for complex disease genes. Genome Res. 10, 1435–1444 (2000).
Ott, J. Predicting the range of linkage disequilibrium. Proc. Natl Acad. Sci. USA 97, 2–3 (2000).
Reich, D.E. et al. Linkage disequilibrium in the human genome. Nature 411,199–204 (2001).
Lichten, M. & Goldman, A.S.H. Meiotic recombination hotspots. Annu. Rev. Genet. 29, 423–444 (1995).
Shiroishi, T., Koide, T., Yoshino, M., Sagai, T. & Morikawi, K. Hotspots of homologous recombination in mouse meiosis. Adv. Biophys. 31, 119–132 (1995).
Petes, T.D. Meiotic recombination hot spots and cold spots. Nature Rev. Genet. 2, 360–369 (2001).
Cullen, M. et al. Characterization of recombination in the HLA class II region. Am. J. Hum. Genet. 60, 397–407 (1997).
Beck, S. et al. Complete sequence and gene map of a human major histocompatibility complex. Nature 401, 921–923 (1999).
Cullen, M., Erlich, H., Klitz, W. & Carrington, M. Molecular mapping of a recombination hotspot located in the second intron of the human TAP2 locus. Am. J. Hum. Genet. 56, 1350–1358 (1995).
Jeffreys, A.J., Ritchie, A. & Neumann, R. High-resolution analysis of haplotype diversity and meiotic crossover in the human TAP2 recombination hotspot. Hum. Mol. Genet. 9, 725–733 (2000).
Sachidanandam, R. et al. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001).
Jeffreys, A.J., Murray, J. & Neumann, R. High-resolution mapping of crossovers in human sperm defines a minisatellite-associated recombination hotspot. Mol. Cell 2, 267–273 (1998).
Borts, R.H. & Haber, J.E. Meiotic recombination in yeast—alteration by multiple heterozygosities. Science 237, 1459–1465 (1987).
Gyapay, G. et al. The 1993–94 Généthon human genetic linkage map. Nature Genet. 7, 246–339 (1994).
Steinmetz, M., Stephan, D. & Lindahl, K.F. Gene organization and recombinational hotspots in the murine major histocompatibility complex. Cell 44, 895–904 (1986).
Snoek, M., Teuscher, C. & van Vugt, H. Molecular analysis of the major MHC recombinational hot spot located within the G7c gene of the murine class III region that is involved in disease susceptibility. J. Immunol. 160, 266–272 (1998).
Gerton, J.L. et al. Global mapping of meiotic recombination hotspots and coldspots in the yeast Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 97, 11383–11390 (2000).
Baudat, F. & Nicolas, A. Clustering of meiotic double-strand breaks on yeast chromosome III. Proc. Natl Acad. Sci. USA 94, 5213–5218 (1997).
Ohta, K., Shibata, T. & Nicolas, A. Changes in chromatin structure at recombination initiation sites during yeast meiosis. EMBO J. 13, 5754–5763 (1994).
Wu, T.C. & Lichten, M. Meiosis-induced double-strand break sites determined by yeast chromatin structure. Science 263, 515–518 (1994).
Fan, Q.Q. & Petes, T.D. Relationship between nuclease-hypersensitive sites and meiotic recombination hot spot activity at the HIS4 locus of Saccharomyces cerevisiae. Mol. Cell. Biol. 16, 2037–2043 (1996).
Hedrick, P.W. Inference of recombinational hotspots using gametic disequilibrium values. Heredity 60, 435–438 (1988).
Daly, M.J., Rioux, J.D., Schaffner, S.F., Hudson, T.J. & Lander, E.S. High-resolution haplotype structure in the human genome. Nature Genet. 29, 229–232 (2001).
Johnson, G.C.L. et al. Haplotype tagging for the identification of common disease genes. Nature Genet. 29,233–237 (2001).
Lewontin, R.C. The interaction of selection and linkage. I. General considerations; heterotic models. Genetics 49, 49–67 (1984).
Hill, W.G. & Robertson, A. Linkage disequilibrium in finite populations. Theor. Appl. Genet. 38, 226–231 (1968).
Jeffreys, A.J., MacLeod, A., Tamaki, K., Neil, D.L. & Monckton, D.G. Minisatellite repeat coding as a digital approach to DNA typing. Nature 354, 204–209 (1991).
Jeffreys, A.J. et al. Complex gene conversion events in germline mutation at human minisatellites. Nature Genet. 6, 136–145 (1994).
Cheng, S., Fockler, C., Barnes, W.M. & Higuchi, R. Effective amplification of long targets from cloned inserts and human genomic DNA. Proc. Natl Acad. Sci. USA 91, 5695–5699 (1994).
Su, X., Wu, Y., Sifri, C.D. & Wellems, T.E. Reduced extension temperatures required for PCR amplification of extremely A+T-rich DNA. Nucleic Acids Res. 24, 1574–1575 (1996).
Morton, N.E. Outline of Genetic Epidemiology (Karger, Basel, 1982).
Sved, J.A. Linkage disequilibrium and homozygosity of chromosomal segments in finite populations. Theor. Popul. Biol. 2, 125–141 (1971).
Acknowledgements
We thank J. Blower and numerous volunteers for supplying semen and blood samples, K. Lilley for assistance with automated sequencing and oligonucleotide synthesis, R. Dalgleish for advice on PCR, J. Stead for web site construction and other colleagues for helpful discussions. This work was supported by grants to L.K. from the Instrumentarium Science Foundation and the Finnish Cultural Foundation and to A.J.J. from the Medical Research Council, Wellcome Trust and Royal Society.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jeffreys, A., Kauppi, L. & Neumann, R. Intensely punctate meiotic recombination in the class II region of the major histocompatibility complex. Nat Genet 29, 217–222 (2001). https://doi.org/10.1038/ng1001-217
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1001-217
This article is cited by
-
Fine human genetic map based on UK10K data set
Human Genetics (2022)
-
Identification of Y chromosome markers in the eastern three-lined skink (Bassiana duperreyi) using in silico whole genome subtraction
BMC Genomics (2020)
-
Heterogeneity in the extent of linkage disequilibrium among exonic, intronic, non-coding RNA and intergenic chromosome regions
European Journal of Human Genetics (2019)
-
Inverse PCR to perform long-distance haplotyping: main applications to improve preimplantation genetic diagnosis in hemophilia
European Journal of Human Genetics (2019)
-
Heterogeneous transposable elements as silencers, enhancers and targets of meiotic recombination
Chromosoma (2019)