Abstract
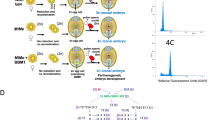
Traditionally, hybrid seeds are produced by crossing selected inbred lines. Here we provide a proof of concept for reverse breeding, a new approach that simplifies meiosis such that homozygous parental lines can be generated from a vigorous hybrid individual. We silenced DMC1, which encodes the meiotic recombination protein DISRUPTED MEIOTIC cDNA1, in hybrids of A. thaliana, so that non-recombined parental chromosomes segregate during meiosis. We then converted the resulting gametes into adult haploid plants, and subsequently into homozygous diploids, so that each contained half the genome of the original hybrid. From 36 homozygous lines, we selected 3 (out of 6) complementing parental pairs that allowed us to recreate the original hybrid by intercrossing. In addition, this approach resulted in a complete set of chromosome-substitution lines. Our method allows the selection of a single choice offspring from a segregating population and preservation of its heterozygous genotype by generating homozygous founder lines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Chen, Z.J. Molecular mechanisms of polyploidy and hybrid vigor. Trends Plant Sci. 15, 57–71 (2010).
van Dijk, P. & van Damme, J. Apomixis technology and the paradox of sex. Trends Plant Sci. 5, 81–84 (2000).
Dirks, R. et al. Reverse breeding: A novel breeding approach based on engineered meiosis. Plant Biotechnol. J. 7, 837–845 (2009).
Ma, H. A molecular portrait of Arabidopsis meiosis. Arabidopsis Book 4, e0095 (2006).
Couteau, F. et al. Random chromosome segregation without meiotic arrest in both male and female meiocytes of a dmc1 mutant of Arabidopsis. Plant Cell 11, 1623–1634 (1999).
Hartung, F. et al. The catalytically active tyrosine residues of both SPO11–1 and SPO11–2 are required for meiotic double-strand break induction in Arabidopsis. Plant Cell 19, 3090–3099 (2007).
Forster, B.P., Heberle-Bors, E., Kasha, K.J. & Touraev, A. The resurgence of haploids in higher plants. Trends Plant Sci. 12, 368–375 (2007).
Ravi, M. & Chan, S.W.L. Haploid plants produced by centromere-mediated genome elimination. Nature 464, 615–618 (2010).
Drouaud, J. et al. Sex-specific crossover distributions and variations in interference level along Arabidopsis thaliana chromosome 4. PLoS Genet. 3, e106 (2007).
Toyota, M., Matsuda, K., Kakutani, T., Terao Morita, M. & Tasaka, M. Developmental changes in crossover frequency in Arabidopsis. Plant J. 65, 589–599 (2011).
Alonso-Blanco, C. et al. Development of an AFLP based linkage map of Ler, Col and Cvi Arabidopsis thaliana ecotypes and construction of a Ler/Cvi recombinant inbred line population. Plant J. 14, 259–271 (1998).
Hauge, B.M. et al. An integrated genetic/RFLP map of the Arabidopsis thaliana genome. Plant J. 3, 745–754 (1993).
Nadeau, J.H., Singer, J.B., Matin, A. & Lander, E.S. Analysing complex traits with chromosome substitution strains. Nat. Genet. 24, 221–225 (2000).
Koumproglou, R. et al. STAIRS: A new genetic resource for functional genomic studies of Arabidopsis. Plant J. 31, 355–364 (2002).
Marimuthu, M.P.A. et al. Synthetic clonal reproduction through seeds. Science 331, 876 (2011).
Mercier, R. & Grelon, M. Meiosis in plants: Ten years of gene discovery. Cytogenet. Genome Res. 120, 281–290 (2008).
Tester, M. & Langridge, P. Breeding technologies to increase crop production in a world. Science 327, 818–822 (2010).
Gleave, A.P. A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol. Biol. 20, 1203–1207 (1992).
Clough, S.J. & Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M.K. & Scheible, W.-R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005).
Ross, K.J., Fransz, P. & Jones, G.H. A light microscopic atlas of meiosis in Arabidopsis thaliana. Chromosome Res. 4, 507–516 (1996).
Ross, K.J. et al. Cytological characterization of four meiotic mutants of Arabidopsis isolated from T-DNA–transformed lines. Chromosome Res. 5, 551–559 (1997).
Tang, X. et al. Cross-species bacterial artificial chromosome–fluorescence in situ hybridization painting of the tomato and potato chromosome 6 reveals undescribed chromosomal rearrangements. Genetics 180, 1319–1328 (2008).
Warthmann, N., Fitz, J. & Weigel, D. MSQT for choosing SNP assays from multiple DNA alignments. Bioinformatics 23, 2784–2787 (2007).
Stam, P. Construction of integrated genetic linkage maps by means of a new computer package: JoinMap. Plant J. 3, 739–744 (1993).
Lister, C. & Dean, C. Recombinant inbred lines for mapping RFLP and phenotypic markers in Arabidopsis thaliana. Plant J. 4, 745–750 (1993).
Singer, T. et al. A high-resolution map of Arabidopsis recombinant inbred lines by whole-genome exon array hybridization. PLoS Genet. 2, e144 (2006).
Bikard, D. et al. Divergent evolution of duplicate genes leads to genetic incompatibilities within A. thaliana. Science 323, 623–626 (2009).
Acknowledgements
We thank T. Stoker, G. Stunnenberg, C. Pillen and H. Blankestijn for help in the greenhouse, J. van Ooijen for advice in using Joinmap, and L. Mlynarova, Y.-F. Lin and D.Z. Agudelo for their help with real-time PCR. We thank M. Koornneef, B. Zwaan and P.J. Flood for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
K.v.D., C.L.C.L., H.d.J. and R.D. conceived the research. E.W. performed cytology and crosses, C.B.d.S. constructed vectors and performed genotyping and N.S.N. performed cytology and FISH. E.W. analyzed data with the help of C.B.d.S., J.J.B.K., M.R. and S.W.L.C. E.W. wrote the manuscript and made the figures with substantial contributions by J.J.B.K., M.R., S.W.L.C. and H.d.J. All authors except N.S.N. were involved in planning and design of experiments, and read and improved the manuscript.
Corresponding author
Ethics declarations
Competing interests
E. W., K.v.D., C.B.d.S., C.L.C.L. and R.D. are employees of Rijk Zwaan. H.d.J. has received research funding from Rijk Zwaan in recent years.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1, 3 and 4 (PDF 735 kb)
Supplementary Table 2
Genotypes of WT and RB haploids and reconstructed heterozygotes. (XLSX 134 kb)
Rights and permissions
About this article
Cite this article
Wijnker, E., van Dun, K., de Snoo, C. et al. Reverse breeding in Arabidopsis thaliana generates homozygous parental lines from a heterozygous plant. Nat Genet 44, 467–470 (2012). https://doi.org/10.1038/ng.2203
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2203
This article is cited by
-
Androgenesis in soybean (Glycine max (L.) Merr.): a critical revisit
In Vitro Cellular & Developmental Biology - Plant (2024)
-
A simple and highly efficient strategy to induce both paternal and maternal haploids through temperature manipulation
Nature Plants (2023)
-
Repair of DNA double-strand breaks in plant meiosis: role of eukaryotic RecA recombinases and their modulators
Plant Reproduction (2023)
-
Fifty years of a public cassava breeding program: evolution of breeding objectives, methods, and decision-making processes
Theoretical and Applied Genetics (2021)
-
In vitro androgenesis: spontaneous vs. artificial genome doubling and characterization of regenerants
Plant Cell Reports (2020)