Abstract
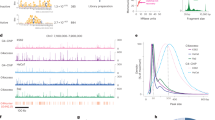
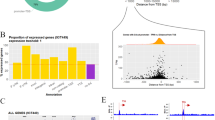
G4 motifs are greatly enriched near promoters, suggesting that quadruplex structures may be targets of transcriptional regulation. Here we show, by ChIP-Seq analysis of human cells, that 40% of the binding sites of the transcription-associated helicases, XPB and XPD, overlap with G4 motifs. The highly significant overlap of XPB and XPD binding sites with G4 motifs cannot be explained by GC richness or parameters of the genomewide analysis, but instead suggests that these proteins are recruited to quadruplex structures that form in genomic DNA (G4 DNA). Biochemical analysis demonstrates that XPD is a robust G4 DNA helicase and that XPB binds G4 DNA. XPB and XPD are enriched near the transcription start site at 20% of genes, especially highly transcribed genes. XPB and XPD enrichment at G4 motifs characterizes specific signaling pathways and regulatory pathways associated with specific cancers. These results identify new candidate pathways for therapies targeted to quadruplexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Gellert, M., Lipsett, M.N. & Davies, D.R. Helix formation by guanylic acid. Proc. Natl. Acad. Sci. USA 48, 2013–2018 (1962).
Sen, D. & Gilbert, W. Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 334, 364–366 (1988).
Maizels, N. & Gray, L.T. The G4 genome. PLoS Genet. 9, e1003468 (2013).
Eddy, J. Gene function correlates with potential for G4 DNA formation in the human genome. Nucleic Acids Res. 34, 3887–3896 (2006).
Huppert, J.L. & Balasubramanian, S. G-quadruplexes in promoters throughout the human genome. Nucleic Acids Res. 35, 406–413 (2007).
Du, Z., Zhao, Y. & Li, N. Genome-wide colonization of gene regulatory elements by G4 DNA motifs. Nucleic Acids Res. 37, 6784–6798 (2009).
Eddy, J. & Maizels, N. Selection for the G4 DNA motif at the 5′ end of human genes. Mol. Carcinog. 48, 319–325 (2009).
Duquette, M.L., Handa, P., Vincent, J.A., Taylor, A.F. & Maizels, N. Intracellular transcription of G-rich DNAs induces formation of G-loops, novel structures containing G4 DNA. Genes Dev. 18, 1618–1629 (2004).
Balasubramanian, S., Hurley, L.H. & Neidle, S. Targeting G-quadruplexes in gene promoters: a novel anticancer strategy? Nat. Rev. Drug Discov. 10, 261–275 (2011).
Renaud de la Faverie, A. et al. Nucleic acids targeted to drugs: SELEX against a quadruplex ligand. Biochimie 93, 1357–1367 (2011).
González, V. & Hurley, L.H. The C-terminus of nucleolin promotes the formation of the c-MYC G-quadruplex and inhibits c-MYC promoter activity. Biochemistry 49, 9706–9714 (2010).
Wei, D., Parkinson, G.N., Reszka, A.P. & Neidle, S. Crystal structure of a c-kit promoter quadruplex reveals the structural role of metal ions and water molecules in maintaining loop conformation. Nucleic Acids Res. 40, 4691–4700 (2012).
Tornaletti, S., Park-Snyder, S. & Hanawalt, P.C. G4-forming sequences in the non-transcribed DNA strand pose blocks to T7 RNA polymerase and mammalian RNA polymerase II. J. Biol. Chem. 283, 12756–12762 (2008).
Belotserkovskii, B.P. et al. Mechanisms and implications of transcription blockage by guanine-rich DNA sequences. Proc. Natl. Acad. Sci. USA 107, 12816–12821 (2010).
Rodriguez, R. et al. Small molecule–induced DNA damage identifies alternative DNA structures in human genes. Nat. Chem. Biol. 8, 301–310 (2012).
Oksenych, V. & Coin, F. The long unwinding road: XPB and XPD helicases in damaged DNA opening. Cell Cycle 9, 90–96 (2010).
Egly, J.-M. & Coin, F. A history of TFIIH: two decades of molecular biology on a pivotal transcription/repair factor. DNA Repair (Amst.) 10, 714–721 (2011).
Fuss, J.O. & Tainer, J.A. XPB and XPD helicases in TFIIH orchestrate DNA duplex opening and damage verification to coordinate repair with transcription and cell cycle via CAK kinase. DNA Repair (Amst.) 10, 697–713 (2011).
Compe, E. & Egly, J.-M. TFIIH: when transcription met DNA repair. Nat. Rev. Mol. Cell Biol. 13, 343–354 (2012).
Fan, L. et al. XPD helicase structures and activities: insights into the cancer and aging phenotypes from XPD mutations. Cell 133, 789–800 (2008).
White, M.F. Structure, function and evolution of the XPD family of iron-sulfur-containing 5′→3′ DNA helicases. Biochem. Soc. Trans. 37, 547–551 (2009).
Hashimoto, S. & Egly, J.-M. Trichothiodystrophy view from the molecular basis of DNA repair/transcription factor TFIIH. Hum. Mol. Genet. 18, R224–R230 (2009).
Kamileri, I., Karakasilioti, I. & Garinis, G.A. Nucleotide excision repair: new tricks with old bricks. Trends Genet. 28, 566–573 (2012).
London, T.B.C. et al. FANCJ is a structure-specific DNA helicase associated with the maintenance of genomic G/C tracts. J. Biol. Chem. 283, 36132–36139 (2008).
Wu, Y., Shin-ya, K. & Brosh, R.M. FANCJ helicase defective in Fanconia anemia and breast cancer unwinds G-quadruplex DNA to defend genomic stability. Mol. Cell. Biol. 28, 4116–4128 (2008).
Sarkies, P., Reams, C., Simpson, L.J. & Sale, J.E. Epigenetic instability due to defective replication of structured DNA. Mol. Cell 40, 703–713 (2010).
Sarkies, P. et al. FANCJ coordinates two pathways that maintain epigenetic stability at G-quadruplex DNA. Nucleic Acids Res. 40, 1485–1498 (2012).
Uringa, E.-J., Youds, J.L., Lisaingo, K., Lansdorp, P.M. & Boulton, S.J. RTEL1: an essential helicase for telomere maintenance and the regulation of homologous recombination. Nucleic Acids Res. 39, 1647–1655 (2011).
Wu, Y., Sommers, J.A., Khan, I., de Winter, J.P. & Brosh, R.M. Biochemical characterization of Warsaw breakage syndrome helicase. J. Biol. Chem. 287, 1007–1021 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Besnard, E. et al. Unraveling cell type–specific and reprogrammable human replication origin signatures associated with G-quadruplex consensus motifs. Nat. Struct. Mol. Biol. 19, 837–844 (2012).
Guédin, A., Gros, J., Alberti, P. & Mergny, J.L. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 38, 7858–7868 (2010).
Dubaele, S. et al. Basal transcription defect discriminates between xeroderma pigmentosum and trichothiodystrophy in XPD patients. Mol. Cell 11, 1635–1646 (2003).
Liu, H. et al. Structure of the DNA repair helicase XPD. Cell 133, 801–812 (2008).
Rudolf, J., Rouillon, C., Schwarz-Linek, U. & White, M.F. The helicase XPD unwinds bubble structures and is not stalled by DNA lesions removed by the nucleotide excision repair pathway. Nucleic Acids Res. 38, 931–941 (2010).
Tirode, F., Busso, D., Coin, F. & Egly, J.M. Reconstitution of the transcription factor TFIIH: assignment of functions for the three enzymatic subunits, XPB, XPD, and cdk7. Mol. Cell 3, 87–95 (1999).
Fan, L. et al. Conserved XPB core structure and motifs for DNA unwinding: implications for pathway selection of transcription or excision repair. Mol. Cell 22, 27–37 (2006).
Lin, Y.C., Choi, W.S. & Gralla, J.D. TFIIH XPB mutants suggest a unified bacterial-like mechanism for promoter opening but not escape. Nat. Struct. Mol. Biol. 12, 603–607 (2005).
Coin, F., Oksenych, V. & Egly, J.-M. Distinct roles for the XPB/p52 and XPD/p44 subcomplexes of TFIIH in damaged DNA opening during nucleotide excision repair. Mol. Cell 26, 245–256 (2007).
Oksenych, V., de Jesus, B.B., Zhovmer, A., Egly, J.-M. & Coin, F. Molecular insights into the recruitment of TFIIH to sites of DNA damage. EMBO J. 28, 2971–2980 (2009).
McLean, C.Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Paeschke, K., Capra, J.A. & Zakian, V.A. DNA replication through G-quadruplex motifs is promoted by the Saccharomyces cerevisiae Pif1 DNA helicase. Cell 145, 678–691 (2011).
Lagerwerf, S., Vrouwe, M.G., Overmeer, R.M., Fousteri, M.I. & Mullenders, L.H.F. DNA damage response and transcription. DNA Repair (Amst.) 10, 743–750 (2011).
Bauer, N.N., Chen, Y.-W., Samant, R.S., Shevde, L.A. & Fodstad, O. Rac1 activity regulates proliferation of aggressive metastatic melanoma. Exp. Cell Res. 313, 3832–3839 (2007).
Hodis, E. et al. A landscape of driver mutations in melanoma. Cell 150, 251–263 (2012).
Krauthammer, M. et al. Exome sequencing identifies recurrent somatic RAC1 mutations in melanoma. Nat. Genet. 44, 1006–1014 (2012).
Phan, A.T., Kuryavyi, V. & Patel, D.J. DNA architecture: from G to Z. Curr. Opin. Struct. Biol. 16, 288–298 (2006).
Biffi, G., Tannahill, D., McCafferty, J. & Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 5, 182–186 (2013).
Lam, E.Y.N., Beraldi, D., Tannahill, D. & Balasubramanian, S. G-quadruplex structures are stable and detectable in human genomic DNA. Nat. Commun. 4, 1796–1798 (2013).
Gray, L.T., Fong, K.K., Pavelitz, T. & Weiner, A.M. Tethering of the conserved piggyBac transposase fusion protein CSB-PGBD3 to chromosomal AP-1 proteins regulates expression of nearby genes in humans. PLoS Genet. 8, e1002972 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Kent, W.J., Zweig, A.S., Barber, G., Hinrichs, A.S. & Karolchik, D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics 26, 2204–2207 (2010).
Stajich, J.E. et al. The Bioperl toolkit: Perl modules for the life sciences. Genome Res. 12, 1611–1618 (2002).
Meyer, L.R. et al. The UCSC Genome Browser database: extensions and updates 2013. Nucleic Acids Res. 41, D64–D69 (2013).
Shin, H., Liu, T., Manrai, A.K. & Liu, X.S. CEAS: cis-regulatory element annotation system. Bioinformatics 25, 2605–2606 (2009).
Huppert, J.L. & Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 33, 2908–2916 (2005).
Decorsière, A., Cayrel, A., Vagner, S. & Millevoi, S. Essential role for the interaction between hnRNP H/F and a G quadruplex in maintaining p53 pre-mRNA 3′-end processing and function during DNA damage. Genes Dev. 25, 220–225 (2011).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Rudolf, J., Makrantoni, V., Ingledew, W.J., Stark, M.J.R. & White, M.F. The DNA repair helicases XPD and FancJ have essential iron-sulfur domains. Mol. Cell 23, 801–808 (2006).
Sun, H., Karow, J.K., Hickson, I.D. & Maizels, N. The Bloom's syndrome helicase unwinds G4 DNA. J. Biol. Chem. 273, 27587–27592 (1998).
Acknowledgements
Research reported in this manuscript was supported by US National Institutes of Health–National Cancer Institute grant P01 CA077852. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We thank our Program Project colleagues for active discussions, M. White (St. Andrews University, Scotland) for the XPD expression construct; J. Tainer (Scripps Research Institute) for the XPB expression construct; A. Bailey for help with sequencing library preparation and C. Lee and J. Shendure for sequencing of ChIP-Seq libraries on an Illumina Hi-Seq 2000 (University of Washington).
Author information
Authors and Affiliations
Contributions
L.T.G. performed ChIP-Seq and expression array experiments and analyzed those results. A.C.V. and J.E. performed biochemical studies of XPB and XPD. L.T.G., A.C.V., J.E. and N.M. conceived the study and designed experiments. L.T.G., A.C.V., J.E. and N.M. interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–4 and Supplementary Tables 1–3. (PDF 5952 kb)
Supplementary Data Set 1
Gene expression data for HT1080 cell lines. (XLSX 766 kb)
Supplementary Data Set 2
Genes with enrichment of XPB or XPD within 1 kb of TSS. (XLSX 264 kb)
Supplementary Data Set 3
Selected online GREAT results for XPB and XPD peaks containing motifs (XLSX 56 kb)
Rights and permissions
About this article
Cite this article
Gray, L., Vallur, A., Eddy, J. et al. G quadruplexes are genomewide targets of transcriptional helicases XPB and XPD. Nat Chem Biol 10, 313–318 (2014). https://doi.org/10.1038/nchembio.1475
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1475
This article is cited by
-
In vivo dynamics and regulation of DNA G-quadruplex structures in mammals
Cell & Bioscience (2023)
-
Profiling human pathogenic repeat expansion regions by synergistic and multi-level impacts on molecular connections
Human Genetics (2023)
-
TET deficiency perturbs mature B cell homeostasis and promotes oncogenesis associated with accumulation of G-quadruplex and R-loop structures
Nature Immunology (2022)
-
G4-quadruplex-binding proteins: review and insights into selectivity
Biophysical Reviews (2022)
-
Parallel G-quadruplexes recruit the HSV-1 transcription factor ICP4 to promote viral transcription in herpes virus-infected human cells
Communications Biology (2021)