Abstract
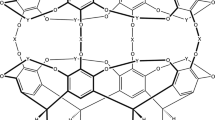
Small molecules that recognize protein surfaces are important tools for modifying protein interaction properties. Since the 1980s, several thousand studies concerning calixarenes and host–guest interactions have been published. Although there is growing interest in protein–calixarene interactions, only limited structural information has been available to date. We now report the crystal structure of a protein–calixarene complex. The water-soluble p-sulfonatocalix[4]arene is shown to bind the lysine-rich cytochrome c at three different sites. Binding curves obtained from NMR titrations reveal an interaction process that involves two or more binding sites. Together, the data indicate a dynamic complex in which the calixarene explores the surface of cytochrome c. In addition to providing valuable information on protein recognition, the data also indicate that the calixarene is a mediator of protein–protein interactions, with potential applications in generating assemblies and promoting crystallization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Zorn, J. A. & Wells, J. A. Turning enzymes on with small molecules. Nature Chem. Biol. 6, 179–188 (2010).
Zhou, H., Baldini, L., Hong, J., Wilson, A. J. & Hamilton, A. D. Pattern recognition of proteins based on an array of functionalized porphyrins. J. Am. Chem. Soc. 128, 2421–2425 (2006).
Larson, S. B., Day, J. S., Cudney, R. & McPherson, A. A novel strategy for the crystallization of proteins: X-ray diffraction validation. Acta Crystallogr. D 63, 310–318 (2007).
Ernst, J. T., Becerril, J., Park, H. S., Yin, H. & Hamilton, A. D. Design and application of an alpha-helix-mimetic scaffold based on an oligoamide-foldamer strategy: antagonism of the Bak BH3/Bcl-xL complex. Angew. Chem. Int. Ed. 42, 535–539 (2003).
Lin, Q., Park, H. S., Hamuro, Y., Lee, C. S. & Hamilton, A. D. Protein surface recognition by synthetic agents: design and structural requirements of a family of artificial receptors that bind to cytochrome c. Biopolymers 47, 285–297 (1998).
Crowley, P. B., Ganji, P. & Ibrahim, H. Protein surface recognition: structural characterisation of cytochrome c–porphyrin complexes. ChemBioChem 9, 1029–1033 (2008).
Renner, C., Piehler, J. & Schrader, T. Arginine- and lysine-specific polymers for protein recognition and immobilization. J. Am. Chem. Soc. 128, 620–628 (2006).
Ingerman, L. A., Cuellar, M. E. & Waters, M. L. A small molecule receptor that selectively recognizes trimethyl lysine in a histone peptide with native protein-like affinity. Chem. Commun. 46, 1839–1841 (2010).
Li, H. et al. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442, 91–95 (2006).
Coleman, A. W. et al. Novel layer structure of sodium calix[4]arenesulfonate complexes—a class of organic clay mimics. Angew. Chem. Int. Ed. 27, 1361–1362 (1988).
Selkti, M. et al. The first example of a substrate spanning the calix[4]arene bilayer: the solid state complex of p-sulfonatocalix[4]arene with L-lysine. Chem. Commun. 2, 161–162 (2000).
Atwood, J. L., Ness, T., Nichols, P. J. & Raston, C. L. Confinement of amino acids in tetra-p-sulfonated calix[4]arene bilayers. Cryst. Growth Des. 2, 171–176 (2002).
Arena, G. et al. Inclusion of naturally occurring amino acids in water soluble calix[4]arenes: a microcalorimetric and 1H NMR investigation supported by molecular modeling. Org. Biomol. Chem. 4, 243–249 (2006).
Gordo, S. et al. Stability and structural recovery of the tetramerization domain of p53–R337H mutant induced by a designed templating ligand. Proc. Natl Acad. Sci. USA 105, 16426–16431 (2008).
Martos, V. et al. Molecular recognition and self-assembly special feature: calix[4]arene-based conical-shaped ligands for voltage-dependent potassium channels. Proc. Natl Acad. Sci. USA 106, 10482–10486 (2009).
Danylyuk, O. & Suwinska, K. Solid-state interactions of calixarenes with biorelevant molecules. Chem. Commun. 39, 5799–5813 (2009).
Beshara, C. S., Jones, C. E., Daze, K. D., Lilgert, B. J. & Hof, F. A simple calixarene recognizes post-translationally methylated lysine. ChemBioChem 11, 63–66 (2010).
Perret, F. & Coleman, A. W. Biochemistry of anionic calix[n]arenes. Chem. Commun. 47, 7303–7319 (2011).
Louie, G. V. & Brayer, G. D. High-resolution refinement of yeast iso-1-cytochrome c and comparisons with other eukaryotic cytochromes c. J. Mol. Biol. 214, 527–555 (1990).
Beissenhirtz, M. K. et al. Electroactive cytochrome c multilayers within a polyelectrolyte assembly. Angew. Chem. Int. Ed. 43, 4357–4360 (2004).
Zadmard, R. & Schrader, T. Nanomolar protein sensing with embedded receptor molecules. J. Am. Chem. Soc. 127, 904–915 (2005).
Lange, C. & Hunte, C. Crystal structure of the yeast cytochrome bc1 complex with its bound substrate cytochrome c. Proc. Natl Acad. Sci. USA 99, 2800–2805 (2002).
Crowley, P. B. & Carrondo, M. A. The architecture of the binding site in redox protein complexes: implications for fast dissociation. Proteins 55, 603–612 (2004).
Volkov, A. N., Bashir, Q., Worrall, J. A. R., Ullmann, G. M. & Ubbink, M. Shifting the equilibrium between the encounter state and the specific form of a protein complex by interfacial point mutations. J. Am. Chem. Soc. 132, 11487–11495 (2010).
Crowley, P. B., Chow, E. & Papkovskaia, T. Protein interactions in the Escherichia coli cytosol: an impediment to in-cell NMR spectroscopy. ChemBioChem 12, 1043–1048 (2011).
Assfalg, M., Bertini, I., Del Conte, R., Giachetti, A. & Turano, P. Cytochrome c and organic molecules: solution structure of the p-aminophenol adduct. Biochemistry 46, 6232–6238 (2007).
Patriarca, A. et al. ATP acts as a regulatory effector in modulating structural transitions of cytochrome c: implications for apoptotic activity. Biochemistry 48, 3279–3287 (2009).
Hill, M. M., Adrain, C., Duriez, P. J., Creagh, E. M. & Martin, S. J. Analysis of the composition, assembly kinetics and activity of native Apaf-1 apoptosomes. EMBO J. 23, 2134–2145 (2004).
Gallivan, J. P. & Dougherty, D. Cation–π interactions in structural biology. Proc. Natl Acad. Sci. USA 96, 9459–9464 (1999).
Fucke, K. et al. The structure of water in p-sulfonatocalix[4]arene. Chemistry 17, 10259–10271 (2011).
Hendsch, Z. S. & Tidor, B. Do salt bridges stabilize proteins? A continuum electrostatic analysis. Protein Sci. 3, 211–226 (1994).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Søndergaard, C. R., Garrett, A. E., Carstensen, T., Pollastri, G. & Nielsen, J. E. Structural artifacts in protein–ligand X-ray structures: implications for the development of docking scoring functions. J. Med. Chem. 52, 5673–5684 (2009).
Janin, J., Bahadur, R. P. & Chakrabarti, P. Protein–protein interaction and quaternary structure. Q. Rev. Biophys. 41, 133–180 (2008).
Weimann, D. P., Winkler, H. D., Falenski, J. A., Koksch, B. & Schalley, C. A. Highly dynamic motion of crown ethers along oligolysine peptide chains. Nature Chem. 1, 573–577 (2009).
Chinai, J. M. et al. Molecular recognition of insulin by a synthetic receptor. J. Am. Chem. Soc. 133, 8810–8813 (2011).
Oshima, T., Goto, M. & Furusaki, S. Complex formation of cytochrome c with a calixarene carboxylic acid derivative: a novel solubilization method for biomolecules in organic media. Biomacromolecules 3, 438–444 (2002).
Paul, D. et al. Chemical activation of cytochrome c proteins via crown ether complexation: cold-active synzymes for enantiomer-selective sulfoxide oxidation in methanol. J. Am. Chem. Soc. 125, 11478–11479 (2003).
Oshima, T., Higuchi, H., Ohto, K., Inoue, K. & Goto, M. Selective extraction and recovery of cytochrome c by liquid–liquid extraction using a calix[6]arene carboxylic acid derivative. Langmuir 21, 7280–7284 (2005).
Morgan, H. P. et al. An improved strategy for the crystallization of Leishmania mexicana pyruvate kinase. Acta Crystallogr. F 66, 215–218 (2010).
Walter, T. S. et al. Lysine methylation as a routine rescue strategy for protein crystallization. Structure 14, 1617–1622 (2006).
Derewenda, Z. S. & Vekilov, P. G. Entropy and surface engineering in protein crystallization. Acta Crystallogr. D 62, 116–124 (2006).
Shinkai, S., Araki, K., Tsubaki, T., Arimura, T. & Manabe, O. New syntheses of calixarene-p-sulphonates and p-nitrocalixarenes. J. Chem. Soc. Perkin Trans. 1, 2297–2299 (1987).
Volkov, A. N., Vanwetswinkel, S., Van de Water, K. & van Nuland, N. A. J. Redox-dependent conformational changes in eukaryotic cytochromes revealed by paramagnetic NMR spectroscopy. J. Biomol. NMR 52, 245–256 (2012).
Leslie, A. G. The integration of macromolecular diffraction data. Acta. Crystallogr. D 62, 48–57 (2006).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006).
Collaborative Computational Project Number 4. The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nature Protoc. 3, 1171–1179 (2008).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta. Crystallogr. D 66, 12–21 (2010).
Acknowledgements
This research was supported by NUI Galway (college scholarship to R.E.M., Millennium Fund grants to N.P. and P.B.C.) and Science Foundation Ireland (grants 07/IN.1/B975 to A.R.K. and 07/RFP/BICF236 to P.B.C.). The authors acknowledge the European Synchrotron Radiation Facility for beam time allocation, and the staff of beamline BM14 for assistance with data collection. The authors thank A. N. Volkov (Vrije Universiteit Brussel) for revised NMR assignments of cytochrome c and J. Donlon (NUI Galway) and S. S. Wijmenga (Radboud Universiteit Nijmegen) for helpful discussions.
Author information
Authors and Affiliations
Contributions
P.B.C. conceived and designed the experiments. R.E.M., H.F., A.R.K. and P.B.C. performed the experiments. R.E.M. and P.B.C. analysed the data. N.P.P. contributed the calixarene. P.B.C. wrote the paper. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 392 kb)
Supplementary Movie 1
Supplementary Movie 1 (WMV 1466 kb)
Rights and permissions
About this article
Cite this article
McGovern, R., Fernandes, H., Khan, A. et al. Protein camouflage in cytochrome c–calixarene complexes. Nature Chem 4, 527–533 (2012). https://doi.org/10.1038/nchem.1342
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.1342
This article is cited by
-
A molecular dynamics study of the complexation of tryptophan, phenylalanine and tyrosine amino acids with cucurbit[7]uril
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2022)
-
Specific inhibition of the Survivin–CRM1 interaction by peptide-modified molecular tweezers
Nature Communications (2021)
-
Host-guest complexation of cucurbit[8]uril with two enantiomers
Scientific Reports (2017)
-
Synthesis of coumarin-appended cyclophanes and evaluation of their complexation with myoglobin
Journal of Inclusion Phenomena and Macrocyclic Chemistry (2015)
-
A Stimuli-Responsive Nanopore Based on a Photoresponsive Host-Guest System
Scientific Reports (2013)